Team:KU Leuven/Safety
From 2013.igem.org
Secret garden
Congratulations! You've found our secret garden! Follow the instructions below and win a great prize at the World jamboree!
- A video shows that two of our team members are having great fun at our favourite company. Do you know the name of the second member that appears in the video?
- For one of our models we had to do very extensive computations. To prevent our own computers from overheating and to keep the temperature in our iGEM room at a normal level, we used a supercomputer. Which centre maintains this supercomputer? (Dutch abbreviation)
- We organised a symposium with a debate, some seminars and 2 iGEM project presentations. An iGEM team came all the way from the Netherlands to present their project. What is the name of their city?
Now put all of these in this URL:https://2013.igem.org/Team:KU_Leuven/(firstname)(abbreviation)(city), (loose the brackets and put everything in lowercase) and follow the very last instruction to get your special jamboree prize!


Safety
It is of course very important to stay safe. Here we have gathered information on how we stayed safe in our lab, what the government rules and the rules of the KU Leuven are. Finally, we also elaborate on how we could keep our bacterium from spreading if it would eventually be commercialised and released into the environment. As well as which specific mechanisms have we implemented to keep our bacteria contained, with for instance, a 'kill switch'. We have also written an ethical evaluation of our system concerning certain safety issues.
Safety forms were approved on September 29, 2013 by the iGEM Safety Committee.
We have covered the safety aspect of our project in many ways. Within synthetic biology there should be three main risks that need to be covered: exposure of the laboratory workers to the bacteria and chemicals, effects of a possible escape of the bacteria and the unintentional disturbance of the ecosystem.


(Max Gibson)
In the past three months, we have added new branches into the pathways of E. coli. Our experiments were performed with non-pathogenic E. coli strains (BL21, DH5α, TOP10 and Rosetta™(DE3)pLysS), which belong to risk group 1 of the pathogenic organisms. Working with genetically modified organisms demands extra safety conditions; physical measures have been taken to separate the organism from the surrounding environment and vice versa.
Our project wants to protect plants and crops against aphid infestations and reduce the use of harmful insecticides. Our genetically modified bacteria (BanAphids) will produce (E)-β-farnesene (EBF) and methyl salicylate (MeS), which are naturally occurring substances. EBF repels aphids and MeS attracts ladybugs, one of the aphid's natural enemies. The low concentrations needed for these substances to be effective is not toxic for humans. No harmful effects are described for EBF at high concentrations either. Contrarily, pure MeS could cause harm when it is ingested due to its similarities with acetylsalicylic acid (aspirin) and can cause irritation after contact with eyes and skin. Nevertheless, it is typically used in small amounts as a flavouring agent in candy, or as fragrance oils due to its mint flavour and wintergreen fragrance. The two chemicals being produced by our BanAphids could cause allergic reactions (swelling, redness, ...), but serious reactions only occur in high concentrations and are very rare.
The effect of BanAphids on the environment is not trivial and thorough field experiments must definitely be conducted. The maximum production rate of a single bacterium needs to be determined so that this could be scaled up through a model. This way we would be able to estimate the concentration of EBF and MeS in the surroundings if a whole field would be occupied by BanAphids. We can then determine the necessary parameters in order to minimise ecological disruption according to these calculated estimates of surrounding EBF and MeS concentrations. We also thought about the interactions between aphids and ladybugs themselves. Aphids adapt readily when EBF is constantly present, therefore we included an oscillatory model. More information about this model can be found here.
Legislation in Belgium & Europe

(JRC - European Commission)
KU Leuven "rules"
The KU Leuven has its own biosafety rules in addition to the national biosafety regulations, where Marianne Schoukens acts as the central contact point for Health, Safety and Environment (HSE) issues. All members of our team have received training in Safe Microbiological Techniques as well as general and specific lab safety required by the KU Leuven. The safety training consisted of a summary of the important biosafety issues and the composition of the lab. At the beginning of our project, all members of the iGEM team have received an update from the KU Leuven's HSE department. Via liaison Iris Govaerts, we followed a small seminar and filled in an orientation checklist concerning health and safety training prior to the start of project laboratory work.
The project we ultimately chose was the result of a whole series of brainstorm sessions. In these brainstorm sessions we discussed the feasibility of our ideas with our instructor (Dr. Ir. Ingmar Claes) and our coordinators (Prof. Dr. Johan Robben and Dr. Veerle De Wever). We also invited different PhD students, professors, ex-iGEMers and people from the industrial sector to our brainstorm sessions. They have a lot more knowledge about the specific topics and could help us greatly with our decisions. Once the project was in progress, we also contacted a lot of people with questions about the possibilities. For instance we talked to the company Perseus (Partner in Regulatory and Safety Services) to discuss the possible hurdles we would face and which safety aspects we need to incorporate. All these people had another view on our project and helped us taking into account different issues about the safety of our project: the molecules that our bacteria will produce, release into the environment, etc ...
If our project would be used outside the laboratory, many parameters would need to be determined. As stated earlier, the pheromones produced by the BanAphids are not harmful for our health unless available in high concentrations. So if the BanAphids would be spread across acres of land, the concentration of pheromones could attain a high level. The ceiling of this production would need to be determined to see whether it could be a harmful concentration for our health. Insects however do not need as high of a concentration as humans do to develop a reaction. Therefore the disruption of the ecosystem would need to be considered as well. As with most products released in the environment, its bioaccumulation needs to be at a minimum.

(Norcal)
In all our experiments, we used non-pathogenic E. coli strains. These strains are not harmful for humans, except when they are inhaled or swallowed. The chemicals that are produced by our genetically modified E. coli strain are natural molecules and are not harmful for the environment. An accidental release of our bacteria into the environment could have unexpected effects on the ecological balance. As suggested by Moe-Behrens et al., 2013, containment through engineered auxotrophy so that BanAphids are unable to synthesise an essential compound required for their survival, is the most reliable method currently for biocontainment. We also believe that providing a bacterium with an advantage eg. antibiotic resistance, provides bacteria with the opportunity to mutate into even stronger bacteria. We therefore propose the use of a tryptophan auxotrophic bacterium.
In the second model we have designed, the bacterium would be free in the environment and thus not contained in an enclosed container with medium. Therefore, an auxotrophic mechanism would not be applicable. Here we turn to a ‘kill switch’ or induced lethality mechanism, which is less reliable because of spontaneous mutations that could inactivate or disable this ‘kill switch’. Different kill switches are already being used in the iGEM competition and we find it very important that the work of previous iGEM teams do not go to waste which is why we decided to look into several 'kill switches' developed by previous iGEM teams.
Genetic safeguard strategies
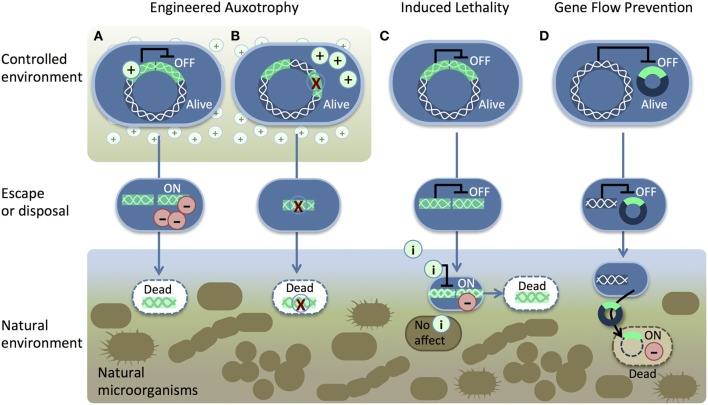
Recombinant DNA (green) is introduced into the host chromosome. Two pathways for engineered auxotrophy (A and B) kill synthetic organisms once they lose access to a supplement (+) in a controlled environment. The supplement either (A) suppresses a toxic gene product (−) or (B) provides nutrition to compensate for a genetic deletion. The induced lethality system (C) produces a toxic gene product (−) in response to an inducer (i) such as IPTG, sucrose, arabinose, or heat (Moe-Behrens et al., 2013).
Auxotrophic mechanism
We, the KU Leuven iGEM team of 2013, propose the use of an auxotrophic mechanism as our main genetic safeguard, when we use our 'sticker system'.
The basic mechanism of the tryptophan auxotrophic bacteria is based on the fact that they are unable to synthesise the essential compound, tryptophan, required for their survival, which means we must provide tryptophan in their medium. They therefore rapidly die once they escape from the controlled environment where the compound is supplied.
Already in 1987, Molin and colleagues designed a DNA cassette that functions as a conditional suicide system in any healthy bacterial strain. In the absence of an artificially supplied growth supplement, the cassette produces a toxic protein that disrupts the bacterial cell's membrane and kills the cells. A stochastic activation of this toxic component could also be used to kill a predetermined fraction of cells per unit of time. This helps to tune the level of lethality so that an optimal level of bioproduction is achieved (Moe-Behrens et al., 2013).
This is a good candidate for the biocontainment of BanAphids in the 'sticker' (Design of Groningen iGEM 2012) or 'sticker system'. This plastic sticker is composed of a plastic film with pores that are too small for the bacteria, but through which our volatiles (EBF and MeS) can escape. However, we need a different mechanisms to contain the BanAphids of our 'spray' or 'honeydew' system.


Kill Switch
The basic mechanism of a kill switch is that the engineered organisms survive until an inducer signal (e.g. IPTG) is added, and this will activate a cascade mechanism that will kill the cells. This induced lethality could, for instance, be used to clean up synthetic microbe spills without harming other cells in the environment (Moe-Behrens et al., 2013).
Furthermore, as suggested by Balan et al. (2005), the use of a nuclease enzyme suicide system disfavours horizontal gene transfer compared to the above suggested suicide system that targets the cell membrane. If a bacterial cell is lysed, that bacterium might be dead but other bacteria in the environment are not dead (yet) and a suicide system that targets the disruption of the cell membrane does not guarantee the degradation of DNA. The lysed bacterium's DNA is now free and available to be taken up by neighbouring bacteria. For this reason we have chosen a 'kill switch' that activates a nuclease system.
Inducible kill systems are not new to iGEM, there are already several constructs available in the database, such as the inducible BamHI system contributed by Berkeley in 2007 (BBa_I716462, tested by Lethbridge in 2011). Here, a BamHI gene was placed downstream of an arabinose-inducible promotor.
Unfortunately, spontaneous genetic mutations in this lethal BamHI gene is a central cause of safeguard failure. Spontaneous genetic mutations can deactivate lethal gene expression causing failure of biocontainment. As these engineered cells propagate in the laboratory, or in large bioreactors, disfunctional genetic safeguards can gradually accumulate in the population. If this accumulation is high enough, then the synthetic organisms might survive in the environment after disposal or accidental release (Moe-Behrens et al., 2013).
Next, a lack of tight control is also a big problem within these systems. Leaky systems cannot be controlled strictly. Therefore, the Calgary team of 2012 added a riboswitch, which is a small piece of mRNA that binds ligands and modifies translation of downstream genes; this way the kill switch can be more tightly controlled. For further details, we would gladly invite you to read the extensive literature study on the wiki of the Calgary 2012 Team.
Moe-Behrens, G. H. G., Davis, R., Haynes, K., Preparing synthetic biology for the world, Frontiers in Microbiology 2013, 4(5): 1-10.
Balan, A., Schenberg, A. C. G., A conditional suicide system for Saccaromyces cerevisiae relying on the intracellular production of the Serratia marcescens nuclease, Yeast 2005; 22; 203-212.
 "
"