Template:Kyoto/Notebook/Sep 14
From 2013.igem.org
(Difference between revisions)
(→Liquid Culture) |
|||
| Line 42: | Line 42: | ||
===Liquid Culture=== | ===Liquid Culture=== | ||
| - | |||
<div class="experiment"> | <div class="experiment"> | ||
| - | <span class="author"> | + | <span class="author">Hirano</span> |
{| class="wikitable" | {| class="wikitable" | ||
!Sample||medium | !Sample||medium | ||
|- | |- | ||
| - | | | + | |9/11 Plac(BBa-R0011)-(3)||Plusgrow medium(+Amp) |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
|} | |} | ||
</div> | </div> | ||
Revision as of 10:30, 25 September 2013
Contents |
Sep 14
Restriction Enzyme Digestion
| 9/14 pSB4K5S | EcoRI | XbaI | SpeI | PstI | BufferB | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|---|
| 2cuts | 10µL | 1µL | 0µL | 1µL | 0µL | 3µL | 0µL | 3µL | 12µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 0µL | 0µL | 1µL | 0µL | 1µL | 7.5µL | 10µL |
| 9/13 Plac | XbaI | PstI | BufferH | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 1 cut | 9µL | 0µL | 1µL | 3µL | 3µL | 14µL | 30µL |
| NC | 1µL | 0µL | 0µL | 1µL | 1µL | 7µL | 10µL |
| 9/14 PKaiBC | EcoRI | XbaI | SpeI | PstI | BufferB | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cuts | 11µL | 1µL | 0µL | 1µL | 0µL | 3µL | 3µL | 11µL | 30µL |
| NC | 4µL | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 4µL | 10µL |
| 9/14 RpaB | EcoRI | XbaI | SpeI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cuts | 13µL | 0µL | 1µL | 0µL | 1µL | 3µL | 3µL | 9µL | 30µL |
| NC | 2µL | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 5µL | 10µL |
| 8/17 RBS-GFP-DT | EcoRI | XbaI | SpeI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cut | 9µL | 0µL | 1µL | 0µL | 0µL | 3µL | 3µL | 14µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.5µL | 10µL |
- incubated 37°C 1hour
Liquid Culture
| Sample | medium |
|---|---|
| 9/11 Plac(BBa-R0011)-(3) | Plusgrow medium(+Amp) |
PCR
| genome DNA | KOD plus | 10x buffer | dNTP | MgSO4 | SasA_fwd primer | SasA_rev primer | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| °C | °C | °C | °C | -- |
| (◎Д◎) |
Restriction Enzyme Digestion
| 8/22 psB1C3-(2) | EcoRI | SpeI | XbaI | PstI | BufferB | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|---|
| 2 cuts | 8µL | 0.5µL | 0.5µL | 0µL | 0µL | 3µL | 0µL | 0.3µL | 17.7µL | 30µL |
| NC | 1µL | 0µL | 0µL | 0µL | 0µL | 1µL | 0µL | 0.1µL | 7.9µL | 10µL |
| 2 cuts | 8µL | 0µL | 0µL | 0.5µL | 0.5µL | 0µL | 3µL | 0.3µL | 17.7µL | 30µL |
| NC | 1µL | 0µL | 0µL | 0µL | 0µL | 0µL | 1µL | 0.1µL | 7.9µL | 10µL |
| 8/21 tRMA-spinach(1) | XbaI | SpeI | BufferB | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 4µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 21.7µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 1µL | 0.1µL | 8.4µL | 10µL |
| 8/21 tetR aptamer12-1R(1) | EcoRI | SpeI | BufferB | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 3µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 22.7µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 0.1µL | 8.5µL | 10µL |
| 8/21 PT181 attenuator-(2) | EcoRI | SpeI | XbaI | PstI | BufferB | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|---|
| 2 cuts | 4µL | 0.5µL | 0.5µL | 0µL | 0µL | 3µL | 0µL | 0.3µL | 21.7µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 0µL | 0µL | 1µL | 0µL | 0.1µL | 8.4µL | 10µL |
| 2 cuts | 4µL | 0µL | 0µL | 0.5µL | 0.5µL | 0µL | 3µL | 0.3µL | 21.7µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 0µL | 0µL | 0µL | 1µL | 0.1µL | 8.4µL | 10µL |
| 8/21 PT181 antisense-(2) | EcoRI | SpeI | XbaI | PstI | BufferB | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|---|
| 2 cuts | 10µL | 0.5µL | 0.5µL | 0µL | 0µL | 3µL | 0µL | 0.3µL | 15.7µL | 30µL |
| NC | 1µL | 0µL | 0&;L | 0µL | 0µL | 1µL | 0µL | 0.1µL | 7.9µL | 10µL |
| 2 cuts | 10µL | 0µL | 0µL | 0.5µL | 0.5µL | 0µL | 3µL | 0.3µL | 15.7µL | 30µL |
| NC | 1µL | 0µL | 0µL | 0µL | 0µL | 0µL | 1µL | 0.1µL | 7.9µL | 10µL |
- incubated 37°C 1hour
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 8/30 pSB1C3 (EcoRI & SpeI) | 6.6 | 8/30 tRNA-Spinach (EcoRI & SpeI) | 12.5 | 3.5 |
| experiment | 8/30 pSB1C3 (EcoRI & SpeI) | 6.6 | 8/30 pT181 antisense (EcoRI & SpeI) | 12.5 | 3.5 |
| experiment | 8/30 pSB1C3 (XbaI & Pst) | 12.5 | 8/30 pT181 attenuator (XbaI & PstI) | 25 | 3.5 |
| experiment | 8/30 pSB1C3 (XbaI & Pst) | 12.5 | 8/30 pT181 antisense (XbaI & PstI) | 25 | 3.5 |
| experiment | 8/20 DT (EcoRI & XbaI) | 6.6 | 8/30 pT181 attenuator(2) (EcoRI & SpeI) | 12.5 | 3.5 |
| experiment | 8/20 DT (EcoRI & XbaI) | 6.6 | 8/30 pT181 antisense (EcoRI & SpeI) | 12.5 | 3.5 |
| experiment | 8/27 DT (EcoRI & XbaI) | 3.0 | 8/30 tRNA-Spinach (EcoRI & SpeI) | 12.5 | 3.5 |
| experiment | 8/20 DT (EcoRI & XbaI) | 6.6 | 8/30 RBS-lysis2 (EcoRI & SpeI) | 12.5 | 3.5 |
| experiment | 8/22 J23100 (SpeI & PstI) | -- | RBS-tetR-DT (XbaI & PstI) | -- | 3.5 |
| experiment | 8/22 Plac (SpeI & PstI) | 8.0 | 8/18 RBS-lacZα-DT (XbaI & PstI) | 10 | 3.5 |
Samples were evaporeted used evaporator into about 7 µL
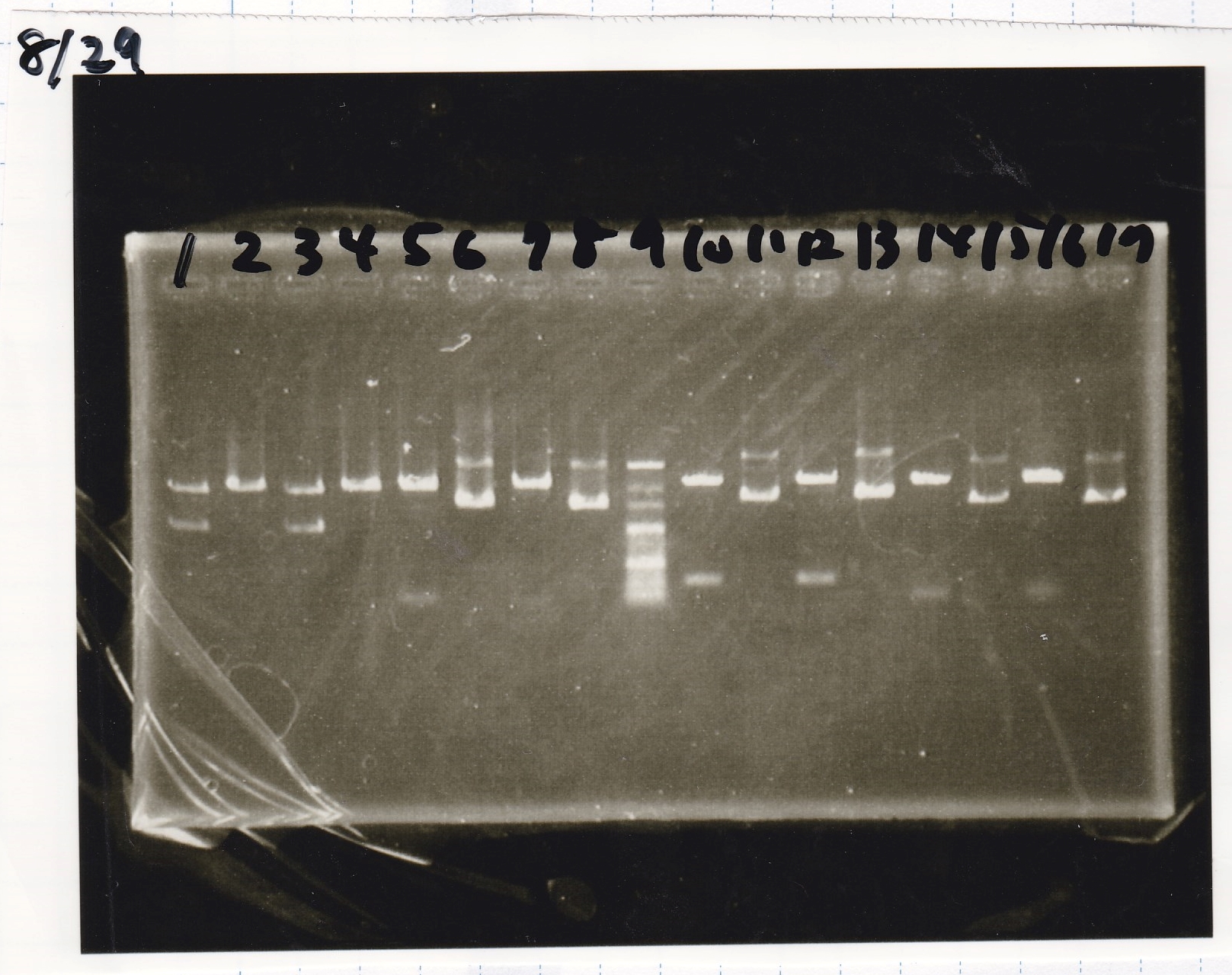
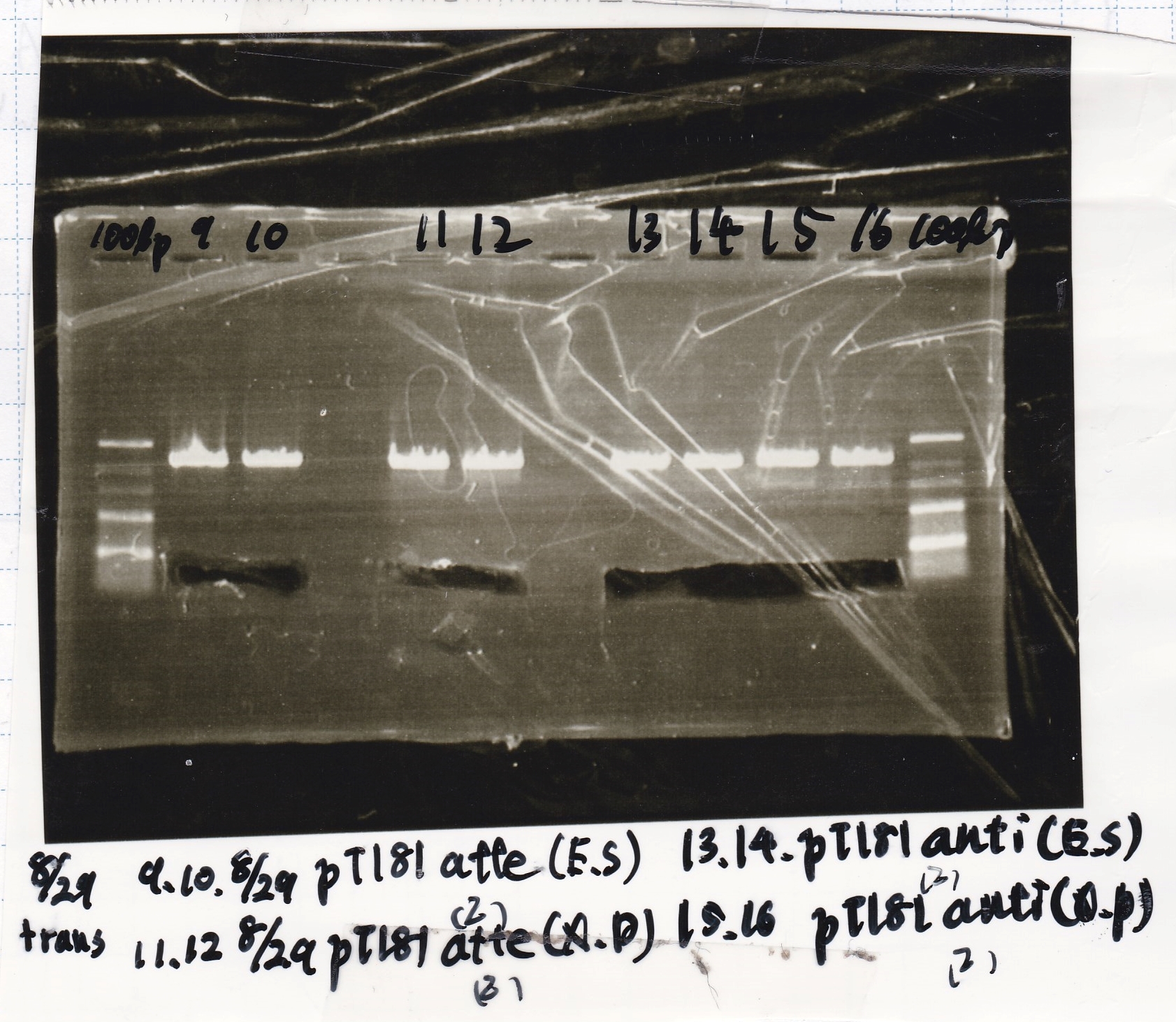
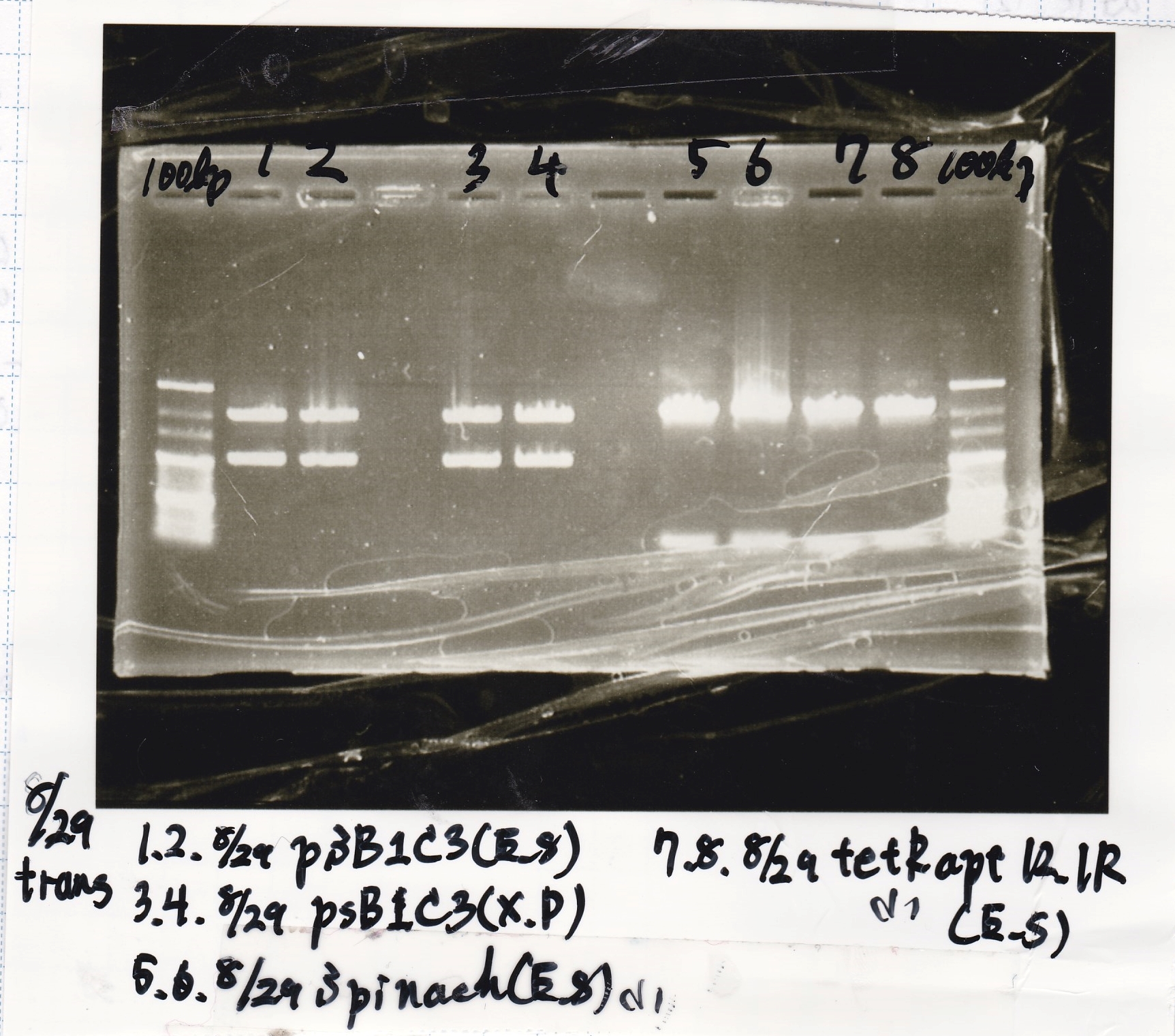
Colony PCR
| Sample | base pair |
|---|---|
| 8/29 Pcon-RBS-GFP-DT-Pcon-RBS-luxR-DT(8/20 LB Amp) (1)~(4) | 2143 |
| 8/29 Plux-RBS-GFP-DT(8/27 LB-CP) (1)~(2) | 1227 |
| 8/29 Pbad/araC-RBS-RFP{BBa_I13516} (8/21 LB-CP) (1)~(2) | 2256 |
| 8/29 RBS-lysis3-DT-(1) | 1210 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 2min | 30cycle |
Liquid Culture
| Sample | medium |
|---|---|
| 8/29 Pcon-RBS-GFP-DT-Pcon-RBS-luxR-DT-(1) | Plusgrow medium(+Amp) |
| 8/29 Pcon-RBS-GFP-DT-Pcon-RBS-luxR-DT-(2) | Plusgrow medium(+Amp) |
| 8/29 Pcon-RBS-GFP-DT-Pcon-RBS-luxR-DT-(3) | Plusgrow medium(+Amp) |
| 8/29 Pcon-RBS-GFP-DT-Pcon-RBS-luxR-DT-(4) | Plusgrow medium(+Amp) |
| 8/29 Plux-RBS-GFP-DT-(1) | Plusgrow medium(+CP) |
| 8/29 Plux-RBS-GFP-DT-(2) | Plusgrow medium(+CP) |
| 8/29 Pbad/araC-RBS-RFP-DT-(1) | Plusgrow medium(+CP) |
| 8/29 Pbad-araC-RBS-RFP-DT-(2) | Plusgrow medium(+CP) |
| RBS-lysis3-DT-(1) | Plusgrow medium(+CP) |
Transformation
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| 8/27 pT181 attenuator-DT | 2µL | 20µL | 22µL | CP |
| 8/27 RBS-lysis3-DT | 2µL | 20µL | 22µL | CP |
| 8/28 RBS-lysis3-DT | 2µL | 20µL | 22µL | CP |
| 8/28 Pcon-RBS-GFP-DT-Pcon-RBS-LuxR-DT | 2µL | 20µL | 22µL | Amp |
| 8/28 RBS-lysis1-DT | 2µL | 20µL | 22µL | CP |
| 8/28 Plux-RBS-GFP-DT | 2µL | 20µL | 22µL | CP |
| 8/28 Spinach-DT | 2µL | 20µL | 22µL | CP |
PCR
| genome DNA | KOD plus | 10x buffer | dNTP | MgSO4 | SasA_fwd primer | SasA_rev primer | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| °C | °C | °C | °C | -- |
| (◎Д◎) |
 "
"