Team:Biwako Nagahama/Project
From 2013.igem.org
Yoshiharu994 (Talk | contribs) (→About curdlan) |
(→E.coli-ink) |
||
| (16 intermediate revisions not shown) | |||
| Line 20: | Line 20: | ||
<p>First, I need to study Paper.</p> | <p>First, I need to study Paper.</p> | ||
<br> | <br> | ||
| - | <p> | + | <p>Cellulose is the main ingredients of plant fiber whereas Paper is made by Cellulose and Hemicellulose, Lignin. Cellulose functions as scaffold, Hemicellulose functions as adhesion, Lignin functions as Space-filling.</p> |
<br> | <br> | ||
<p>Cellulose and Hemicellulose can be produced from synthetic biology. So, the issue regarding paper can surely be solved.</p> | <p>Cellulose and Hemicellulose can be produced from synthetic biology. So, the issue regarding paper can surely be solved.</p> | ||
| Line 122: | Line 122: | ||
<p>(http://www.botanty.utexas.edu/facstaff/facpages/mbroun/ongres/icheeze.htm)</p> | <p>(http://www.botanty.utexas.edu/facstaff/facpages/mbroun/ongres/icheeze.htm)</p> | ||
| - | === <h2> | + | === <h2>Curdlan's introduction</h2> === |
<p>Agrobacterim makes carbohydrate chain. For exampe it makes curdlan on culture midium.We'll explain curdlan on this page.</p> | <p>Agrobacterim makes carbohydrate chain. For exampe it makes curdlan on culture midium.We'll explain curdlan on this page.</p> | ||
| Line 128: | Line 128: | ||
<P>About</p> | <P>About</p> | ||
<P>It is fermentative polysaccharide that microorganizm makes. It polymerizes two position;C₁and C₃on D-glucose. Its binding position is not straight. So it is cycloid structure.</p> | <P>It is fermentative polysaccharide that microorganizm makes. It polymerizes two position;C₁and C₃on D-glucose. Its binding position is not straight. So it is cycloid structure.</p> | ||
| - | [[FIle:Biwako-nagahama_curdlan's-picture_yoshiharu-otaki1.png]] | + | [[FIle:Biwako-nagahama_curdlan's-picture_yoshiharu-otaki1.png|300px|]] |
<p>Property</p> | <p>Property</p> | ||
<p>It is insolubility in water. But it can become dispersing solution in cold water. And after high speed shuffle,the solution go to uniform dispersing solution. Moreover it become complete dispersing solution if it is mixxed by alkaline solution(avobe pH12). And last, dispersing solution become gel unnder the situation that it's heated by high temperature.</p> | <p>It is insolubility in water. But it can become dispersing solution in cold water. And after high speed shuffle,the solution go to uniform dispersing solution. Moreover it become complete dispersing solution if it is mixxed by alkaline solution(avobe pH12). And last, dispersing solution become gel unnder the situation that it's heated by high temperature.</p> | ||
| + | |||
<p>Gel</p> | <p>Gel</p> | ||
| Line 156: | Line 157: | ||
<p>Result</p> | <p>Result</p> | ||
<p>The Gel shown in figure below was made</p> | <p>The Gel shown in figure below was made</p> | ||
| - | [[ | + | [[FIle:Biwako-nagahama_curdlan's-picture_yoshiharu-otaki2.png|200px|]] |
| + | [[FIle:Biwako-nagahama_curdlan's-picture_yoshiharu-otaki3.png|200px|]] | ||
| + | [[FIle:Biwako-nagahama_curdlan's-picture_yoshiharu-otaki4.png|200px|]] | ||
| + | <p>Left 1st day Middle&Right 2nd day</p> | ||
| + | |||
| + | <p>1st days gel has elasticity and softly but 2nd days gel was almost dry. </p> | ||
=== <h2>Binary Vector</h2> === | === <h2>Binary Vector</h2> === | ||
| Line 208: | Line 214: | ||
<p>The nucleotide sequence of myoglobin of sperm whale was already published, and We have synthesis company to synthesis the DNA fragment encoding the myoglobin. </p> | <p>The nucleotide sequence of myoglobin of sperm whale was already published, and We have synthesis company to synthesis the DNA fragment encoding the myoglobin. </p> | ||
<p>Fig1 shows this DNA fragment sequence. Fig1 is published on paper.</p> | <p>Fig1 shows this DNA fragment sequence. Fig1 is published on paper.</p> | ||
| - | [[File:Biwako-Nagahama_sequence_ueda3.png]] | + | [[File:Biwako-Nagahama_sequence_ueda3.png|700px]] |
<p>Fig1.DNA fragment encoding myoglobin.</p> | <p>Fig1.DNA fragment encoding myoglobin.</p> | ||
<p>DNA synthesis company send us the DNA fragment with pTAKN-2. pTAKN-2 is plasmid that .Thus,We get DNA fragment with pTAKN-2.</p> | <p>DNA synthesis company send us the DNA fragment with pTAKN-2. pTAKN-2 is plasmid that .Thus,We get DNA fragment with pTAKN-2.</p> | ||
| Line 215: | Line 221: | ||
<p></p> | <p></p> | ||
| - | <p> | + | <p>two samples(include smaⅠ,plasmid,10×buf) Incubated at 37℃ for 2h.add phenol chloroform mixture 50μℓ and Vortex. centrifugal at 4℃ with 13000rpm for 5min. pour supernatant into another tube. add 3M CH3COONa 5μℓ and 99.5% EtOH 125μℓ to supernatant and Vortex. centrifugal at 4℃ with 13000rpm for 10min. add mixture solution to pellet . mixture solution composition is BamHⅠ,plasmid,10×buf.</p> |
| - | + | ||
| - | + | ||
<p>This sample again incubated at 37℃ for 1h.add phenol chloroform mixture 50μℓ and Vortex. | <p>This sample again incubated at 37℃ for 1h.add phenol chloroform mixture 50μℓ and Vortex. | ||
| - | centrifugal at 4℃ with 13000rpm for 5min. pour supernatant into another tube. add 3M CH3COONa 5μℓ and 99.5% EtOH 125μℓ to supernatant and Vortex. centrifugal at 4℃ with 13000rpm for 10min. add 70% EtOH 500μℓ to pellet and up side down. Remove supernatant . dyr up at 65℃.add TEbuffer 10μℓ to pellet. Mix two samples and Ligation kit Ver2 . We use Ligation kit Ver2 on | + | centrifugal at 4℃ with 13000rpm for 5min. pour supernatant into another tube. add 3M CH3COONa 5μℓ and 99.5% EtOH 125μℓ to supernatant and Vortex. centrifugal at 4℃ with 13000rpm for 10min. add 70% EtOH 500μℓ to pellet and up side down. Remove supernatant . dyr up at 65℃.add TEbuffer 10μℓ to pellet. Mix two samples and Ligation kit Ver2. |
| + | We use Ligation kit Ver2 on Ligate reaction.</p> | ||
<p>Transformation (show protocol page) | <p>Transformation (show protocol page) | ||
Plating to LB-Amp plate. | Plating to LB-Amp plate. | ||
| Line 247: | Line 252: | ||
<p>Fig3. SDS/PAGE analysis(CBB stain one super)with 15%gel</p> | <p>Fig3. SDS/PAGE analysis(CBB stain one super)with 15%gel</p> | ||
<p>Lane1:JM109 with pUC18 Lane2:JM109 with pUC18/Mb</p> | <p>Lane1:JM109 with pUC18 Lane2:JM109 with pUC18/Mb</p> | ||
| - | <p>Next, We have to mutate this DNA fragment encoding myoglonbin because that includes EcoRⅠand XbaⅠsite. GAATTC that is EcoRⅠsite is changed to GAATTT | + | <p>Next, We have to mutate this DNA fragment encoding myoglonbin because that includes EcoRⅠand XbaⅠsite. GAATTC that is EcoRⅠsite is changed to GAATTT. As same TCTAGA that is XbaⅠsite is changed to TCCAGA.</p> |
<p>EcoRⅠand XbaⅠsite pointmutation protocol below</p> | <p>EcoRⅠand XbaⅠsite pointmutation protocol below</p> | ||
<p>First,We design the primer occurring pointmutation.Primer sequence below.</p> | <p>First,We design the primer occurring pointmutation.Primer sequence below.</p> | ||
| Line 265: | Line 270: | ||
<p>We use primer F2 and R3 to make middle fragment on PCR.</p> | <p>We use primer F2 and R3 to make middle fragment on PCR.</p> | ||
<p>We use thermal cycler on PCR. but,middle fragment theoretical value is about 60 base pair, it is very short. So this PCR don’t amplify DNA fragment we targeting but plasmid itself. PCR enzyme using by us is Prime star Max. Prime star Max is highly efficient enzyme. We replace Prime star Max with prime star hot start , but result is same. </p> | <p>We use thermal cycler on PCR. but,middle fragment theoretical value is about 60 base pair, it is very short. So this PCR don’t amplify DNA fragment we targeting but plasmid itself. PCR enzyme using by us is Prime star Max. Prime star Max is highly efficient enzyme. We replace Prime star Max with prime star hot start , but result is same. </p> | ||
| - | <p>Hereupon we hit upon an idea. We worked PCR by hand with water bath at 55℃, 70℃ and 98℃ because We think thermal cycler appear gradient temperature and this is a cause of failure | + | <p>Hereupon we hit upon an idea. We worked PCR by hand with water bath at 55℃, 70℃ and 98℃ because We think thermal cycler appear gradient temperature and this is a cause of failure. We dip sealed sample in water bath one by one. 98℃,55℃,70℃,98℃,55℃・・・・.</p> |
<p>Then,We success PCR that amplify 60 base pair DNA fragment.the PCR productis creaned up with exosap</p> | <p>Then,We success PCR that amplify 60 base pair DNA fragment.the PCR productis creaned up with exosap</p> | ||
<p></p> | <p></p> | ||
| Line 271: | Line 276: | ||
<p>mix the left end fragment and middle fragment and do overlap PCR. PCR products creaned up with exosap</p> | <p>mix the left end fragment and middle fragment and do overlap PCR. PCR products creaned up with exosap</p> | ||
<p>mix the constructed fragment and the right end fragment and do overlap PCR. PCR products creaned up with exosap</p> | <p>mix the constructed fragment and the right end fragment and do overlap PCR. PCR products creaned up with exosap</p> | ||
| - | <p>Do PCR constructed fragment with Fw and Rv primer. </p> | + | <p>Do PCR of constructed fragment with Fw and Rv primer. </p> |
| - | <p>Thus We construct DNA fragment encoding myoglobin not including .</p> | + | <p>Thus We construct DNA fragment encoding myoglobin not including EcoRⅠ and XbaⅠ site .</p> |
<p>We read sequence of this DNA fragment to confirm pointmutation </p> | <p>We read sequence of this DNA fragment to confirm pointmutation </p> | ||
| + | <p>result of ncbi below</p> | ||
| + | [[File:Biwako-Nagahama_sequencing_ueda4.png]] | ||
<p></p> | <p></p> | ||
| + | <p>referance:High-level expression of sperm whale myoglobin in Escherichia coli | ||
| + | (gene synthesis/codon bias/protein engineering) | ||
| + | BARRY A. SPRINGER AND STEPHEN G. SLIGAR | ||
| + | Department of Biochemistry, University of Illinois, Urbana, IL 61801 | ||
| + | Communicated by Gregorio Weber, September 4, 1987</p> | ||
<p></p> | <p></p> | ||
| - | |||
| - | |||
<div class="spacer"></div> | <div class="spacer"></div> | ||
Latest revision as of 03:36, 28 September 2013
Contents |
AgRePaper&E.coli-ink
Cellulose is used as raw material for paper, so our team experimented various ways to increase the amount of cellulose produced by agrobacterium and using it to make papers. For this we developed the different parts to insert into the system of agrobacterium. Among them were the genes used for expression of the curdlan. Similarly, genetic parts in order to increase the expression of the cellulose, along with the agrobacterium type binary vector were also developed . We also worked on recycling of the produced paper by degrading the cellulose to D-Glucose using various enzymes. We worked for the preparation of the biological ink using the sperm whale's cells by genetically modification to increase amount of myoglobin. Then, we observed the change on the color of the product by altering the formation of myoglobin and the production amount of myoglobin with the insertion of T7 promoter to the cell system.
AgRePaper
Background
Today, Our Earth is confronted with global warming and depletion of ozone layer, extinction of organism along with energy crisis as few of many environmental issues.
We think deforestation as its major cause. Many great forests have fallen for production of paper, which we usually use in lots of day to day life for writing and Printing papers,newspaper, origami, wrappers etc… .
To solve this problem,we tried to make paper from resources except wood and raw materials that do not affect environment. So, we can save forests for future.
How to solve it?
First, I need to study Paper.
Cellulose is the main ingredients of plant fiber whereas Paper is made by Cellulose and Hemicellulose, Lignin. Cellulose functions as scaffold, Hemicellulose functions as adhesion, Lignin functions as Space-filling.
Cellulose and Hemicellulose can be produced from synthetic biology. So, the issue regarding paper can surely be solved.
Team Biwako Nagahama try to solve this issue for Agrobacterium tumefaciens C58.
Non-recombinant Agrobacterium tumefaciens synthesize cellulose and other glycan. Publication of Synthesized biofilm with Agrobacterium submitted. In this view, Agrobacterium synthesize cellulose. I suspect it's so.
But, Agrobacterium doesn’t make hemicellulose. So, Hemicellulose like Curdlan(beta-1,3-glycan) pick one out of synthesized glycan from Agrobacterium.
And, Agrobacterium have Cellulose synthase enzyme, including catabolic enzyme. We try to reuse glycan by this enzyme.
In this project, we try to make Cellulose and Curdlan with Agrobacterium. This Cellulose and Curdlan use to make Paper. And, this Paper reuse by catabolic enzyme. So, we approach to an environmental issue!!
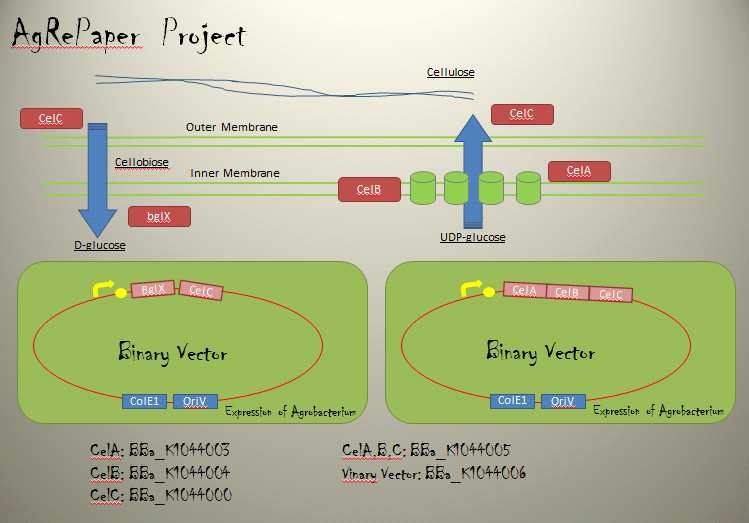
・Cellulose synthesis operon
Cellulose production requires genes encoded within the CelABCG and CelDE operons on the C58 linear chromosome.
CelA is a membrane-associated cellulose synthase (CS) enzyme, utilizing the precursor UDP-glucose.Homologues of CelB bind cyclic diguanosine monophosphate(c-di-GMP),an allosteric regulator of CS activity,and physically interact with CS in the membrane.CelC is a secreted protein similar to endoglucanases.Regulation of cel gene expression is not well understood.
The CS activity of A.tumefaciens is allosterically regulated by the intracellular signal molecule c-di-GMP, which strongly stimulates cellulose synthesis in cell extracts.
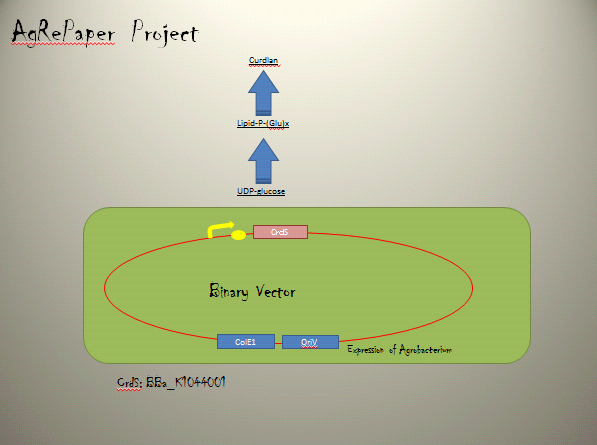
・Curdlan synthesis enzyme
Curdlan synthesis enzyme requires genes encoded within the CrdS on the C58 linear chromosome.
Providing genetic confirmation that crdS is essential for curdlan production.
CrdS was shown to be an integral membrane protein with seven transmembrane helices.
・CelC
Cellulase requires genes encoded within the CelC on the C58 linear chromosome.
CelC belong Glycoside Hydrolase 8(GH8). CelC work as Endo-glucanase gene.CelC is enzyme that catabolized Cellulose to Cellobiose.
Secretion of fibrils from Agro
For this year’s project our team aimed at making the paper by using Agro bacterium. So, we confirmed whether carbohydrate developed from Agrobacterium could be used as the raw material for making paper.
Materials
Agro bacterium tumefaciens C58
Agro bacterium tumefaciens C58 bacterial liquid
+LB medium 2 ml
+ampicillin 2 µl
+Agro bacterium tumefaciens C58
Congo red
LB medium
+1.0%(w/v) Triptone 2 g
+0.5%(w/v) yeast extract 1 g
+1.0%(w/v) NaCl 2 g
Experiment1
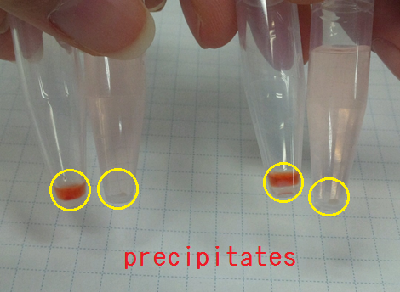
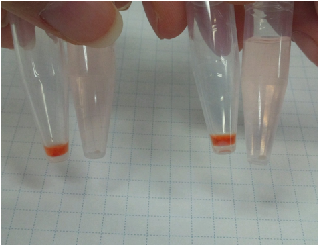
Sample 1 curdlan 15 µl +Congo red 5 µl +Agro bacterium precipitation
Sample 2 de-ionized water 15 µl +Congo red 5 µl +Agro bacterium precipitation
We centrifuged Sample 1 and 2 in 5 minutes with cooled centrifuges(4 ℃,13000 rpm).
As a result, Sample 1 had red precipitates. Sample 2 rationally melted.
So, we confirmed the presence of carbohydrate in the precipitates obtained from Agro bacterium.
Experiment2
Next, we found that carbohydrate could be produced when soy milk is added to the fluid bacteria
So, we experimented for the confirmation of the above findings.
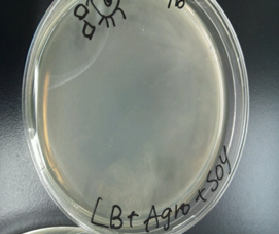
Sample 3 LB medium 7 ml +Agro bacterium+soy 40 µl
Sample 4 LB medium 7 ml +Agro bacterium
Sample 5 LB medium 7 ml +soy 40 µl
Sample 6 LB medium 7 ml
We cultivated Sample 3 ~ 6 at 28 ℃.for 12 hours
Then, Sample1 and 2 produced a white film like substance but Sample 3 and 4 did not.
We put the white films from Sample 1 and 2 into micro-tubes.
Sample3 Sample4
Sample 7 supernatant(Sample 1) 15 µl +Congo red 5 µl
Sample 8 supernatant(Sample 1) 15 µl +Congo red 5 µl +de-ionized water 1 ml
Sample 9 supernatant(Sample 2) 15 µl +Congo red 5 µl
Sample 10 supernatant(Sample 2) 15 µl +Congo red 5 µl +de-ionized water 1 ml
Sample 11 Congo red 5 µl +de-ionized water 1 ml
We centrifuged all the samples for 5 minutes at 4 ℃,13000 rpm.
As a result, Sample 7 ~ 10 gave red precipitates but Sample 11 did not.
So, we confirmed Sample 7 ~ 10 had carbohydrate.
Through these experiments, we confirmed that Agro bacterium can discharge carbohydrate without soy milk.
Reference
1, Laboratory maintenance of Agrobacterium.
(http://europepmc.org/articles/PMC3350319)
2, Coordination of Division and Development Influences Complex Multicellular Behavior in Agrobacterium tumefaciens (Jinwoo Kim.¤, Jason E. Heindl., Clay Fuqua)
3, Kirin Kyowa Foods Company (http://www.kirinkyowa-foods.co.jp/products/curdlan/)
4, Howard Hughes Molecular Biology Summer Research Program Poster, Austin, TX , August, 1995 Microcopy of Curdlan Structure
5, Iain M. Cheeseman and R.Malcom Brown,Jr, Department of Botanty , The University of Texas at Austin,TX,78713
(http://www.botanty.utexas.edu/facstaff/facpages/mbroun/ongres/icheeze.htm)
Curdlan's introduction
Agrobacterim makes carbohydrate chain. For exampe it makes curdlan on culture midium.We'll explain curdlan on this page.
About
It is fermentative polysaccharide that microorganizm makes. It polymerizes two position;C₁and C₃on D-glucose. Its binding position is not straight. So it is cycloid structure.
File:Biwako-nagahama curdlan's-picture yoshiharu-otaki1.png
Property
It is insolubility in water. But it can become dispersing solution in cold water. And after high speed shuffle,the solution go to uniform dispersing solution. Moreover it become complete dispersing solution if it is mixxed by alkaline solution(avobe pH12). And last, dispersing solution become gel unnder the situation that it's heated by high temperature.
Gel
Dispersing solution of curdlan rizes viscosity if its temperature incleses to 50℃,and become gel if it is heated moreover. The strenth of gel is ploportional by concentration of curdlan and/or temperature of heating.
The lelation curdlan and our life
It is used as food additive. The object is improving quolity ,mainting water-holding capasity and improving stability of hot. It is used in nudle,processed food of meet and garnish food. So curdlan isn't distant in our environment. It is relating in our life very much.
Gelatification of the Curdllan
Objective:Gelatification of the curdlan by heating the crdlan &producting3%(w/v) curdlan gel
Procedure
↓10ml of de-ionized water in the beaker and cool the beaker using ice pack.(Curdlan is soluble to cool water)
↓Take 0.30g of curdlan and mux with the cooled de-ionized water using starrer for 20mins.(Occasional cooling of the beaker)
↓Heat the solution until vapors are seen &cool at room temp
↓Throw out the excess of water
↓@30℃ incubator for overnight for dehydrating
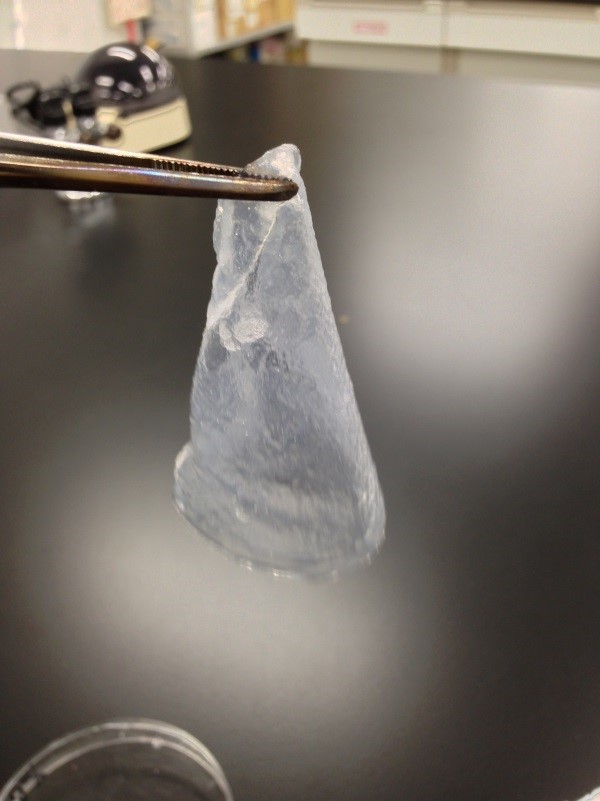
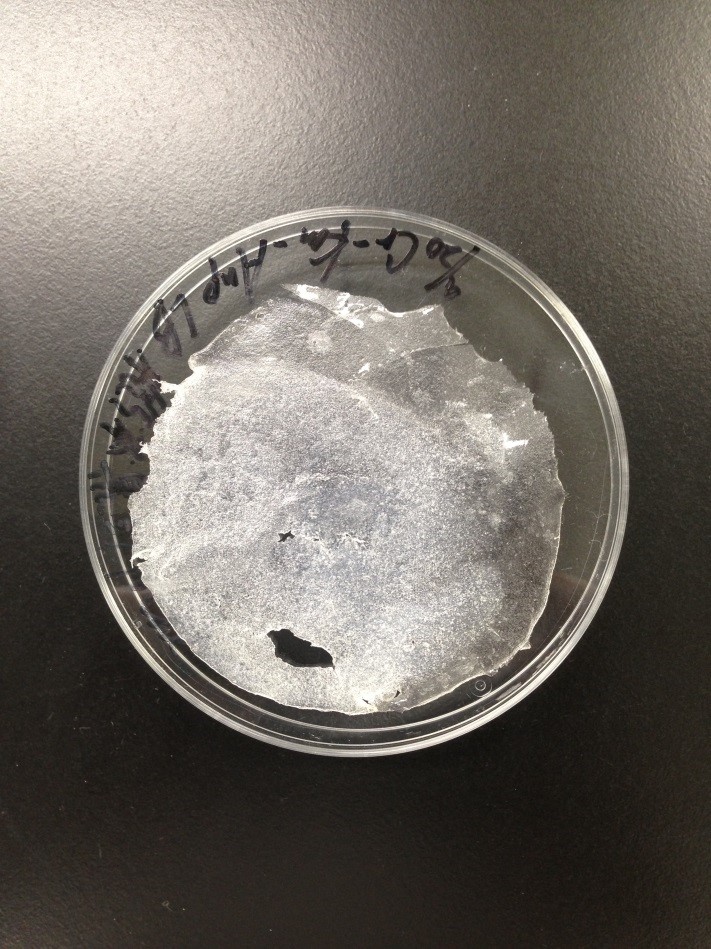
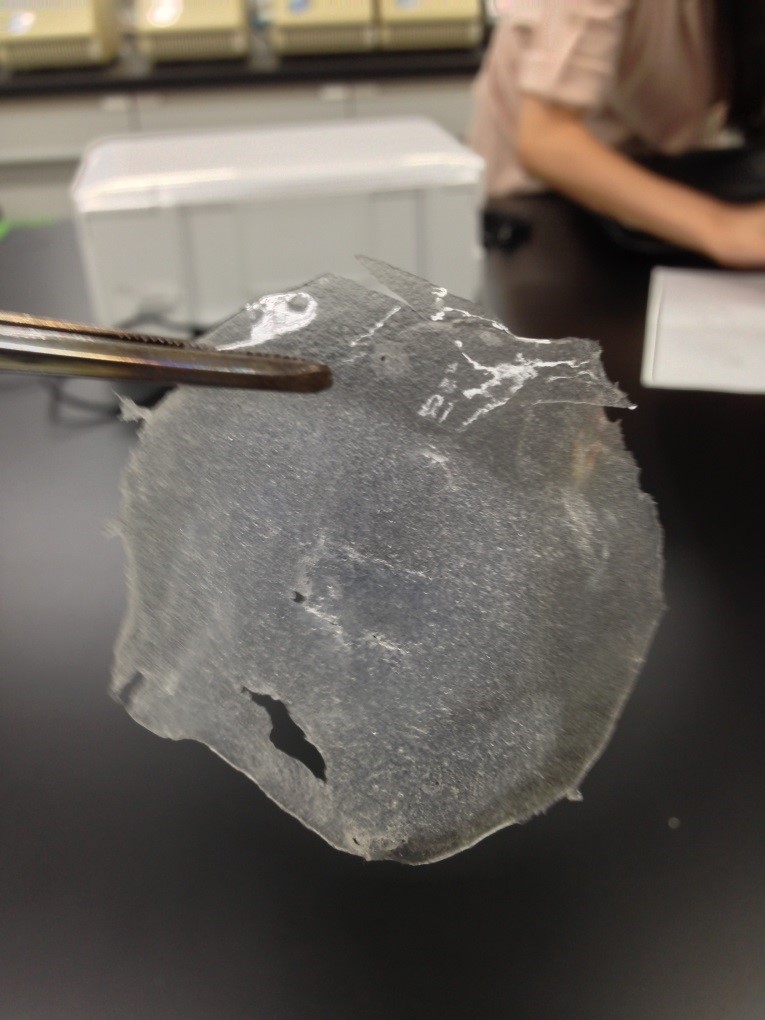
Result
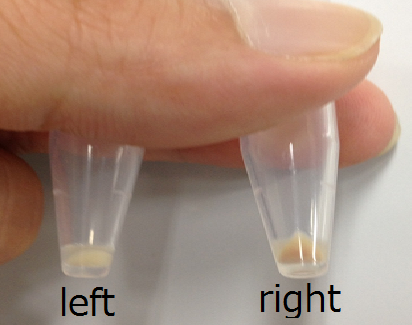
The Gel shown in figure below was made
Left 1st day Middle&Right 2nd day
1st days gel has elasticity and softly but 2nd days gel was almost dry.
Binary Vector
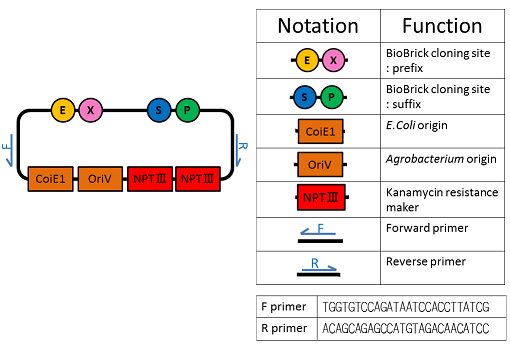
We named BBa_K1044006 "pBI107".This plasmid length is about 9,000 bp.
pBI107 is improved form of empty backbone vector obtained from pre-existing binary vector of E.coli & Agrobacterium. Selective with Kanamycin, there is possibility of reproduction by both E.coli and Agrobacterium.
pBI107 having both prefix and suffix, we can freely insert the cassette into the cloning site of the BioBrick which gives possibility of different expressions of BioBricks with Agrobacterium.

Also the possibility of use of OriV can just not be limited with Agrobacterium but also may broaden to be used with other organisms.
●Proof for the reproduction of E.coli’s pBI107 and selection (NPT III)
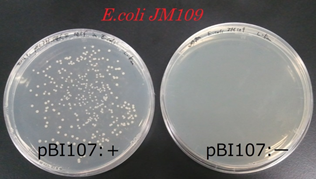
After cloning the E.coli JM109 with pBI107, we plated the contents to a LB-Agar plate with 50 µg/ml kanamycin and cultured it at 37℃ for overnight. Similar experiment was performed for the E.coli JM109 without cloning to pBI107. From the experiment we found that under 50 µg/ml kanamycin, E.coliJM109 with pBI107 cultured and developed colony but the E.coli without pBI107 did not develop any colony.
From above result it was confirmed that E.coli JM109 cloned with pBI107 can reproduce under the influence of kanamycin. Hence, the functionality of ColE & NPTⅢwith regard to E.coli JM109 was proved.
●Proof for the reproduction of Agrobacterium’s pBI107 and selection (NPT III)
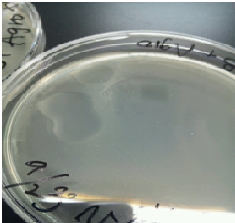
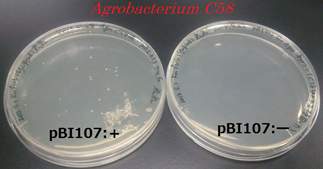
After cloning Agrobacterium C58 with pBI107, it was plated to LB-Agar plate with 50 µg/ml kanamycin and cultured for 2 days at 30℃. Similar, experiment was carried out for the Agrobacterium C58 without cloning it. Colonies were confirmed for the cloned Agrobacetrium C58 under 50 µg/ml kanamycin but no colonies were found for the normal Agrobacterium C58.
From above result it was confirmed that Agrobacterium C58 cloned with pBI107 can reproduce under the influence of kanamycin. Hence, the functionality of OriV & NPTⅢwith regard to Agrobacterium C58 was proved.
●Proof for the prefix and suffix of pBI107
We determined the prefix and suffix of the BioBrick cloning site and inserted RFP device(BBa_J04450) into pBI107and cloned E.coli JM109.
Then it was cultured in LB-Agar plate with 50 µg/ml kanamycin and IPTG at 37℃. After 6 hours of culturing, colonies were detected and after 24 hours of culturing the colonies with inserted RFP device were confirmed by the colonies which were shining red.
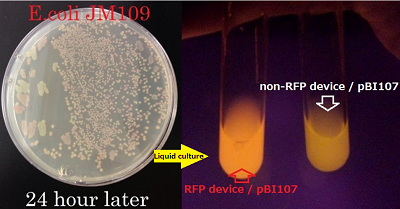
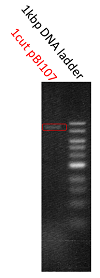
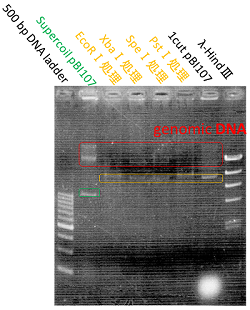
The shining red colonies were extracted and treated with restriction enzyme used for extraction of prefix and suffix. Then the length of vector(pBI107) & insert(RFP device) were confirmed through electrophoresis.
As expected the actual length of pBI107 was 9000bp whereas the extracted RFP device’s length was around 1100bp.
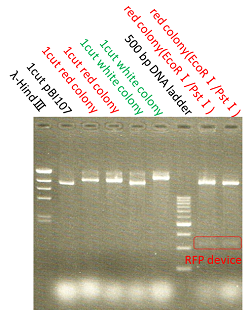
From the above result, we found out the existence of prefix and suffix as the iGEM standard in pBI107 and hence BioBrick was expressed with E.coli JM109.
E.coli-ink
Another one of our project is E.coli ink.
We are planning to use Escherichia coli expressing the sperm whale myoglobin as ink. We are also trying to change the color of transgenic Escherichia coli by changing the growth and medium conditions.
Why do we use myoglobin as ink ?
Recently, fluorescent pigment is very famous and popular. It is nice to look at.
But, Many fluorescent pigment is only colouring when it’s exposed to ultraviolet rays.
We want to make pigment colouring under natural light. Thus,We decided to use myoglobin as ink.
Finally, We will draw my advisor’ face on Agrepaper with this E.coli ink !
The nucleotide sequence of myoglobin of sperm whale was already published, and We have synthesis company to synthesis the DNA fragment encoding the myoglobin.
Fig1 shows this DNA fragment sequence. Fig1 is published on paper.
Fig1.DNA fragment encoding myoglobin.
DNA synthesis company send us the DNA fragment with pTAKN-2. pTAKN-2 is plasmid that .Thus,We get DNA fragment with pTAKN-2.
Next,We insert to pUC18 to confirm the function of this DNA fragment.
We did double digestion pUC18 and pTAKN-2 with BamHⅠand SmaⅠ.This experiment protocol shows below.
two samples(include smaⅠ,plasmid,10×buf) Incubated at 37℃ for 2h.add phenol chloroform mixture 50μℓ and Vortex. centrifugal at 4℃ with 13000rpm for 5min. pour supernatant into another tube. add 3M CH3COONa 5μℓ and 99.5% EtOH 125μℓ to supernatant and Vortex. centrifugal at 4℃ with 13000rpm for 10min. add mixture solution to pellet . mixture solution composition is BamHⅠ,plasmid,10×buf.
This sample again incubated at 37℃ for 1h.add phenol chloroform mixture 50μℓ and Vortex. centrifugal at 4℃ with 13000rpm for 5min. pour supernatant into another tube. add 3M CH3COONa 5μℓ and 99.5% EtOH 125μℓ to supernatant and Vortex. centrifugal at 4℃ with 13000rpm for 10min. add 70% EtOH 500μℓ to pellet and up side down. Remove supernatant . dyr up at 65℃.add TEbuffer 10μℓ to pellet. Mix two samples and Ligation kit Ver2. We use Ligation kit Ver2 on Ligate reaction.
Transformation (show protocol page) Plating to LB-Amp plate.
Next day.
The Colony is on plate. We inoculate this colony into 2μℓ Amp in 2ml 2YT medium in culture tube
Next day
We did Miniprep of this sample.
Theoretical value is but this plasmid is supercoil structure. Therefore, Theoretical value is .
We confirm bund size is theoretical value with agarose electrophoresis.
We again inoculate this colony into 3μℓ 50ng/μℓ Amp in 30μℓ 100mM IPTG in 3ml 2YT medium in culture tube .
Next day, centrifugal this bacterial cell medium at 4℃ with 13000rpm for 1min.
Fig2 shows part’s function!
Fig2.myoglobin is changed it’s color in Ecoli
left:pellet of JM109 with pUC18. right:pellet of JM109 with pUC18/Mb
Next,We use SDS/PAGE to confirm E.coli cloned pUC18/Mb produce myoglobin.
SDS/PAGE sample preparation protocol below.
Centrifugal Bacterial cells 200μℓ at 4℃ with 13000rpm for 1min. remove supernatant. Add 1×sample buffer to pellet and vortex. dip this sample into water bath at 98℃(not heat block). Let it sit on ice 30sec. Centrifugal the sample at 4℃ with 13000rpm for 10min.We use this sample 10μℓ for SDS/PAGE.
Fig3 shows Contative date of part’s function !
Fig3. SDS/PAGE analysis(CBB stain one super)with 15%gel
Lane1:JM109 with pUC18 Lane2:JM109 with pUC18/Mb
Next, We have to mutate this DNA fragment encoding myoglonbin because that includes EcoRⅠand XbaⅠsite. GAATTC that is EcoRⅠsite is changed to GAATTT. As same TCTAGA that is XbaⅠsite is changed to TCCAGA.
EcoRⅠand XbaⅠsite pointmutation protocol below
First,We design the primer occurring pointmutation.Primer sequence below.
F1:T T C T A G A G A G G A G A A C A A C A A C A
R1:ACTAGT A TTATTA TCA T T A A C C C T G G T
F2:C T G G A A T T T A T C T C T G A A G C G
R2:C G C T T C A G A G A T A A A T T C C A G
F3:C T G C A T TC C A G A C A T C C A G G T A A C
R3:G T T A C C T G G A T G T C T G G A A T G C A G
Fw:GTTTCTTCGAATTCGCGGCCGCTTCTAGATG
Rv:GTTTCTTCCTGCAGCGGCCGCTACTAGTATTATTATC
We use primer F1 and R2 to make the left end fragment on PCR. PCR productis creaned up with exosap
We use primer F2 and R3 to make middle fragment on PCR.
We use thermal cycler on PCR. but,middle fragment theoretical value is about 60 base pair, it is very short. So this PCR don’t amplify DNA fragment we targeting but plasmid itself. PCR enzyme using by us is Prime star Max. Prime star Max is highly efficient enzyme. We replace Prime star Max with prime star hot start , but result is same.
Hereupon we hit upon an idea. We worked PCR by hand with water bath at 55℃, 70℃ and 98℃ because We think thermal cycler appear gradient temperature and this is a cause of failure. We dip sealed sample in water bath one by one. 98℃,55℃,70℃,98℃,55℃・・・・.
Then,We success PCR that amplify 60 base pair DNA fragment.the PCR productis creaned up with exosap
We use primer F3 and R1 to make the right end fragment . PCR products creaned up with exosap
mix the left end fragment and middle fragment and do overlap PCR. PCR products creaned up with exosap
mix the constructed fragment and the right end fragment and do overlap PCR. PCR products creaned up with exosap
Do PCR of constructed fragment with Fw and Rv primer.
Thus We construct DNA fragment encoding myoglobin not including EcoRⅠ and XbaⅠ site .
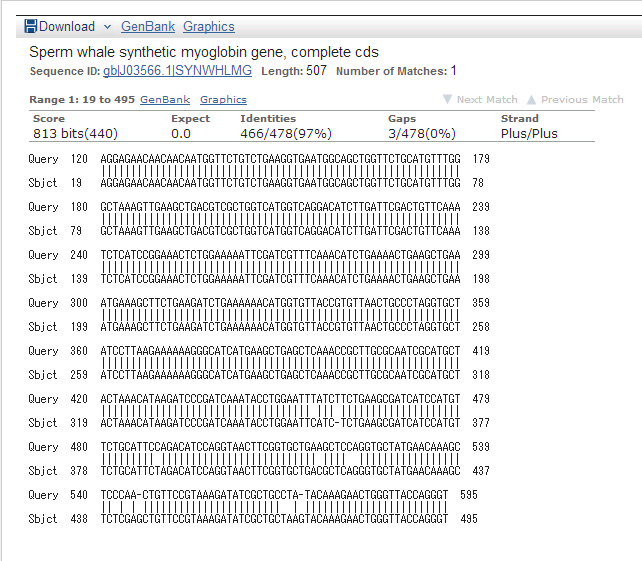
We read sequence of this DNA fragment to confirm pointmutation
result of ncbi below
referance:High-level expression of sperm whale myoglobin in Escherichia coli (gene synthesis/codon bias/protein engineering) BARRY A. SPRINGER AND STEPHEN G. SLIGAR Department of Biochemistry, University of Illinois, Urbana, IL 61801 Communicated by Gregorio Weber, September 4, 1987
 "
"