|
- Small Phage
- March-April
- May-June
- July-August
- September-October
|
8.26 Mutagen Concentration Test - Eighth Protocol
I) Purpose
- To mutate T7 phage for selection of different capsid sizes.
II) Expected Outcome
- A decrease in phage viability with increasing mutagen concentration.
- A few random mutations that produce smaller and larger phage which can then be isolated.
III) Reagents Used
- 5-bromodeoxyuridine stock in freezer labeled T7 (10mg/mL)
- Uracil solution (2.5mg/mL)
- Adenine solution (5mg/mL)
IV) Procedure
1) Propagation of T7 bacteria phage (8.22-8.25)
- Phage concentration of 7.19 T7 stock decreased during its storage in the fridge. Thus, we needed to propagate T7 in preparation for mutagenesis.
- Propagation procedure: 4mL of LB and 1mL of E coli B liquid culture overnight were added to 2 separate test tubes. To the test tube labeled +phage, 10uL of 7.19 T7 stock was added. Both test tubes were incubated on shaker at 37 Celsius for approximately 2 full days.
- After propagation, phage solution was aliquoted (1mL increments) into eppendorf tubes and centrifuged at 3000rpm for 5 minutes. The supernatant was removed and 100uL of chloroform was added to each.
- 1:10 dilution series was performed to generate -1 through -7 sample. Spot test followed when we spotted 5uL of each sample unto one plate overlaid with 0.5mL of E coli B overnight and 5mL of x1 top agar.
- Titers were performed for the -6 and -7 samples: 0.5 mL of E coli B liquid culture overnight was infected with 20 uL of each phage sample for approximately 20 minutes before 5mL of x1 top agar was added and content plated.
2) Applying the mutagen (8.26)
- Label 4 test tubes C, 0, 500, and 850. Add 10mL of LB and 200ul of E coli B overnight into each test tube. Incubate on the shaker at 37 Celsius for approximately an hour before 50uL of adenine solution was added to each test tube.
- Remove all the test tubes off the shaker after another hour of incubation. Add an additional 40ul of adenine and 80ul of uracil to each test tube. Also add the corresponding amount in ul of 5-bromodeoxyuridine, a mutagen, to each test tube. (Ex: Add 500ul of mutagen to tube labeled 500)
- Place all the tubes on the shaker at 37 Celsius.
- After 20 minutes, take only the tube labeled C from the shaker. Take 1mL from this tube and pipette it into a cuvette labeled C. Using 1mL of LB in a cuvette, blank the spectrophotometer at 600 OD. Then measure the absorbance of the cuvette labeled C. Measurement was repeated for verification.
- Remove the other three tubes after 30 minutes of incubation and add 150ul of T7 phage from the "8.24 T7 stock" to each tube. (There should be a 1:10 phage to bacteria concentration. The calculations are as follows: The spectrophotometer reading was 0.354A, which indicates there are 1.77E8 bacteria/ml. Because there are 10ml in each tube, there is roughly 1.77E9 bacteria per tube. This means that we need 1.77E8 phage added to each tube. Because the 8.24 stock is concentrated at 1.6E7 phage/20uL, 150ul of phage will provide 1.2E8 phage, slightly fewer than the intended 1.77E8 to allow for error in estimation.)
- Incubate all three tubes on the shaker at 37 Celsius for 80 minutes.
- Remove all the tubes and add 1mL of chloroform to each. Gently shake each tube and centrifuge it at 4000rpm for 10 minutes at 7 Celsius. Remove the supernatant from each tube with a pipette and place it in a new tube with the same label. Be careful not to get the chloroform or bacteria when you remove the supernatant.
- CsCl gradient was ran right after completion of mutagenesis.
3) Spot test to determine phage concentration (8.26)
- Dilution series -1 through -6 were performed all three samples (0, 500, and 850). These dilutions samples were then used in spot tests to estimate phage titer.
- Specifically, three plates were prepared for spot tests using E. coli B by adding 0.5mL overnight into a tube, mixing it with 5mL of x1 top agar, and then plating the content. 5ul of each dilution sample was spotted onto corresponding plate.
4) CsCl gradient (8.26)
- Phage Purification Team was able to finalize their procedures for performing CsCl gradient on T7 phage. For specifics please refer to ADD LINK
- CsCl separation was performed for both 0 and 500 sample. The finished gradient was aliquoted into 20 eppendorf tubes each, each containing 1-1.5 mL solution. We will need to determine phage characteristics in each aliquot before proceeding with dialysis and TEM.
5) Characterizing post-CsCl phage (8.27-9.2)
- Spot test was performed to estimate phage concentration within each aliquot. Specifically, 1:100 dilution series were performed for each aliquot to generate a -2 and -4 dilution sample for each. All dilution samples were spotted (5uL) onto plates overlaid with 0.5mL of E coli B overnight and 5mL of x1 agar.
- Nearly all aliquot formed plaques during the previous spot test at -4. Thus further dilutions (to -5 and -6) and spot test was performed to more accurately estimate phage concentration. Spot test procedures are similar to that used previously.
- 12 extra spot tests were performed to verify results where plaque formation was faint or unclear.
- Results from concentration spot test were superimposed on the deduced gradient. Reconstructed CsCl gradient is drawn below in the Results section.
V) Results
1) T7 propagation
- Based on the results from the titer experiment, there are roughly 1.6E7 phage/20uL
2) Applying the mutagen
- The OD reading was 0.354A, which indicates there are 1.77E8 bacteria/ml. Because there are 10ml in each tube, there is roughly 1.77E9 bacteria per tube.
3) Spot test to determine phage concentration
- All three samples (0, 500, 850) went down to -6. No obvious death detected.
5) Characterizing post-CsCl phage
- Specific descriptions of results are recorded with the procedures.
- Spot test to estimate phage titer in each aliquot -2/-4 (left) and -4/-5/-6 (right).
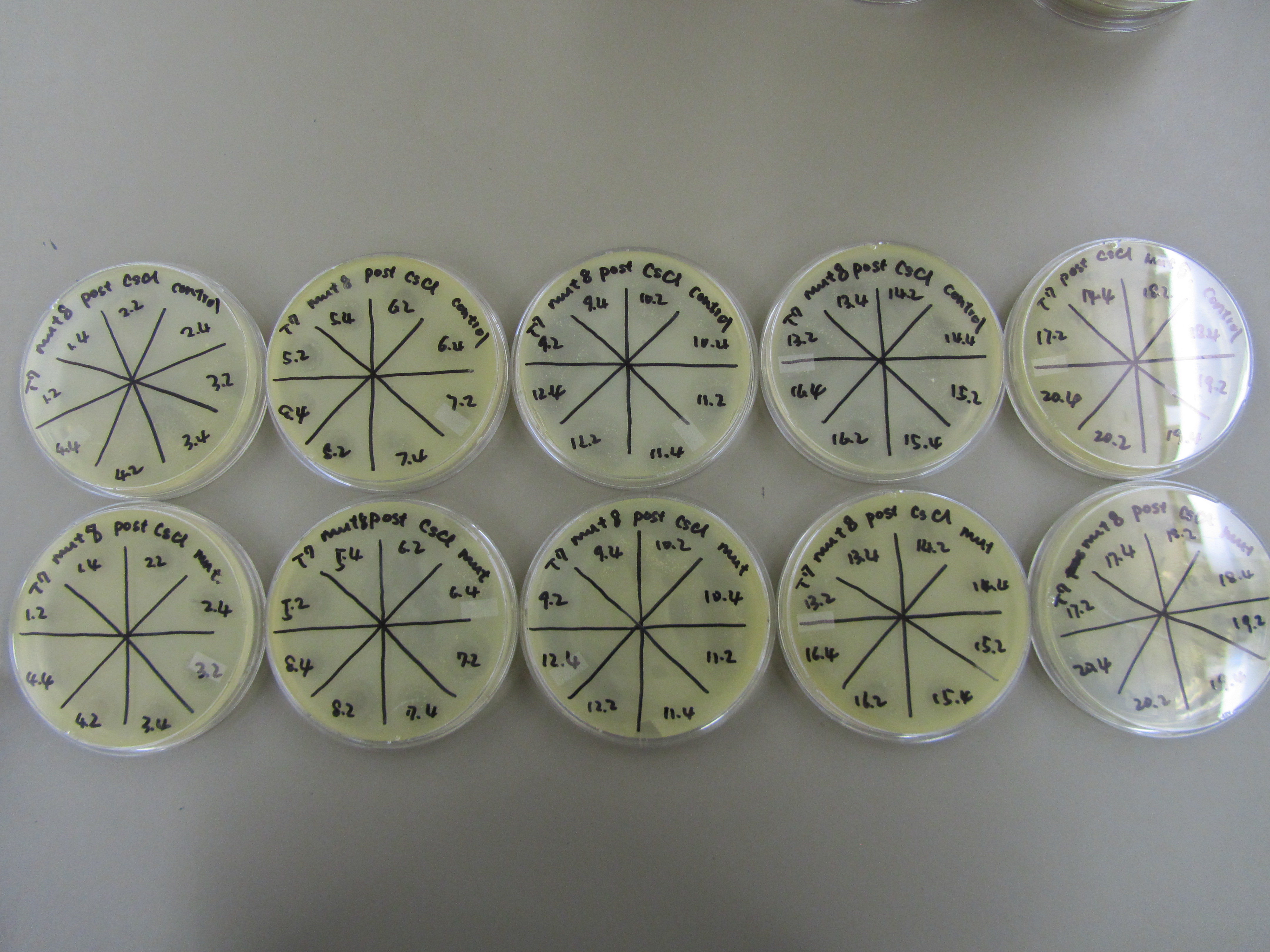
 
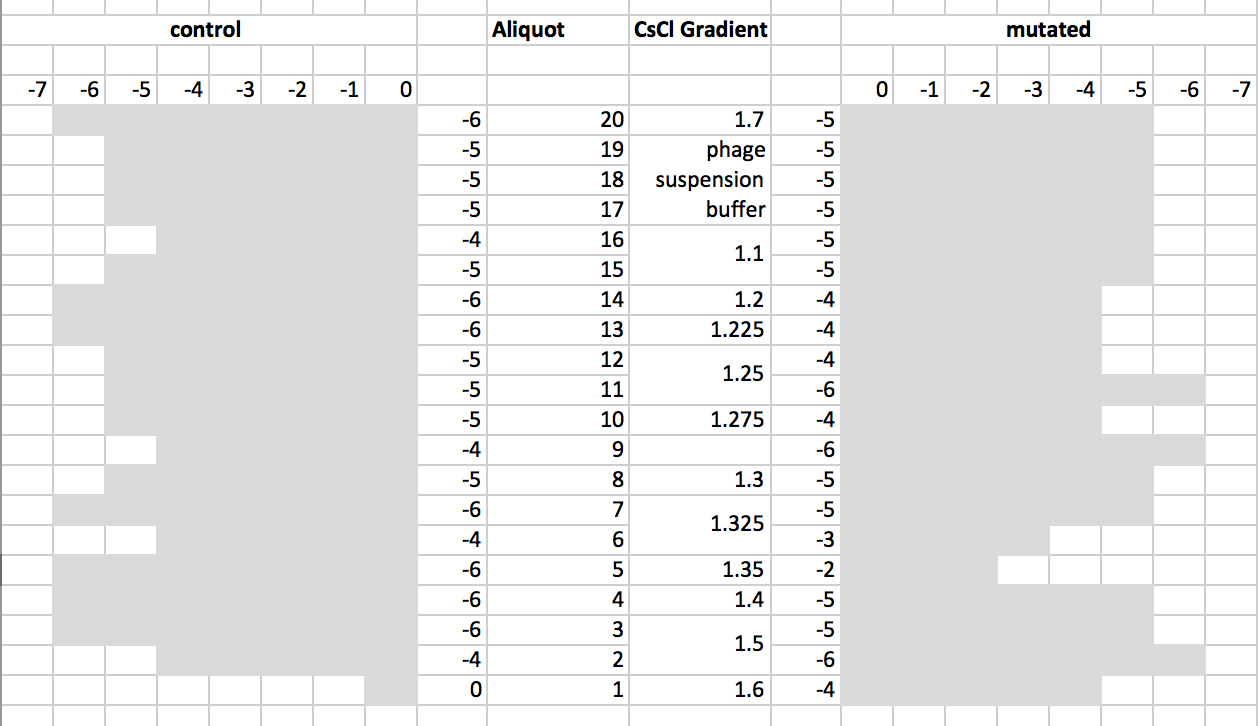
- Deduced gradient with respective estimated titer.
VI) Conclusion
- This round of mutagenesis was successful in that we were able to get a high titer of phage in each aliquot (with an estimated average of 10E7 pfu/mL). However, because phages are spread out throughout the gradient and did not band, we cannot identify which ones are wild type and which ones are mutant. After discussion with Dr. Grose, we decided to hold off our next round of mutagenesis until the Phage Purification Team can get the wild type phage to band. In the meantime, we'll complete our modeling experiments.
|


 "
"

