Template:Kyoto/Notebook/Aug 28
From 2013.igem.org
(Difference between revisions)
(→Gel Extraction) |
(→Colony PCR) |
||
| Line 103: | Line 103: | ||
|5min||30s||30s||48s||30cycles | |5min||30s||30s||48s||30cycles | ||
|} | |} | ||
| - | [[File: | + | [[File:Igku Aug28 Colony PCR(N③pic).jpg]] |
</div> | </div> | ||
Revision as of 10:33, 24 September 2013
Contents |
Aug 28
Electrophoresis
Miniprep
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 8/27 Plux | 173.8 | 1.95 | 1.45 |
Ligation
| state | Vector | Inserter | ||
|---|---|---|---|---|
| experiment | 8/17 DT-1 | 3.0 | 8.0 | |
- Samples were evaporeted used evaporator into about 3 µL.
| sample | MilliQ | Ligation High | total |
|---|---|---|---|
| 3 | 4 | 3.5 | 10.5 |
- incubate one hour at 16 °C
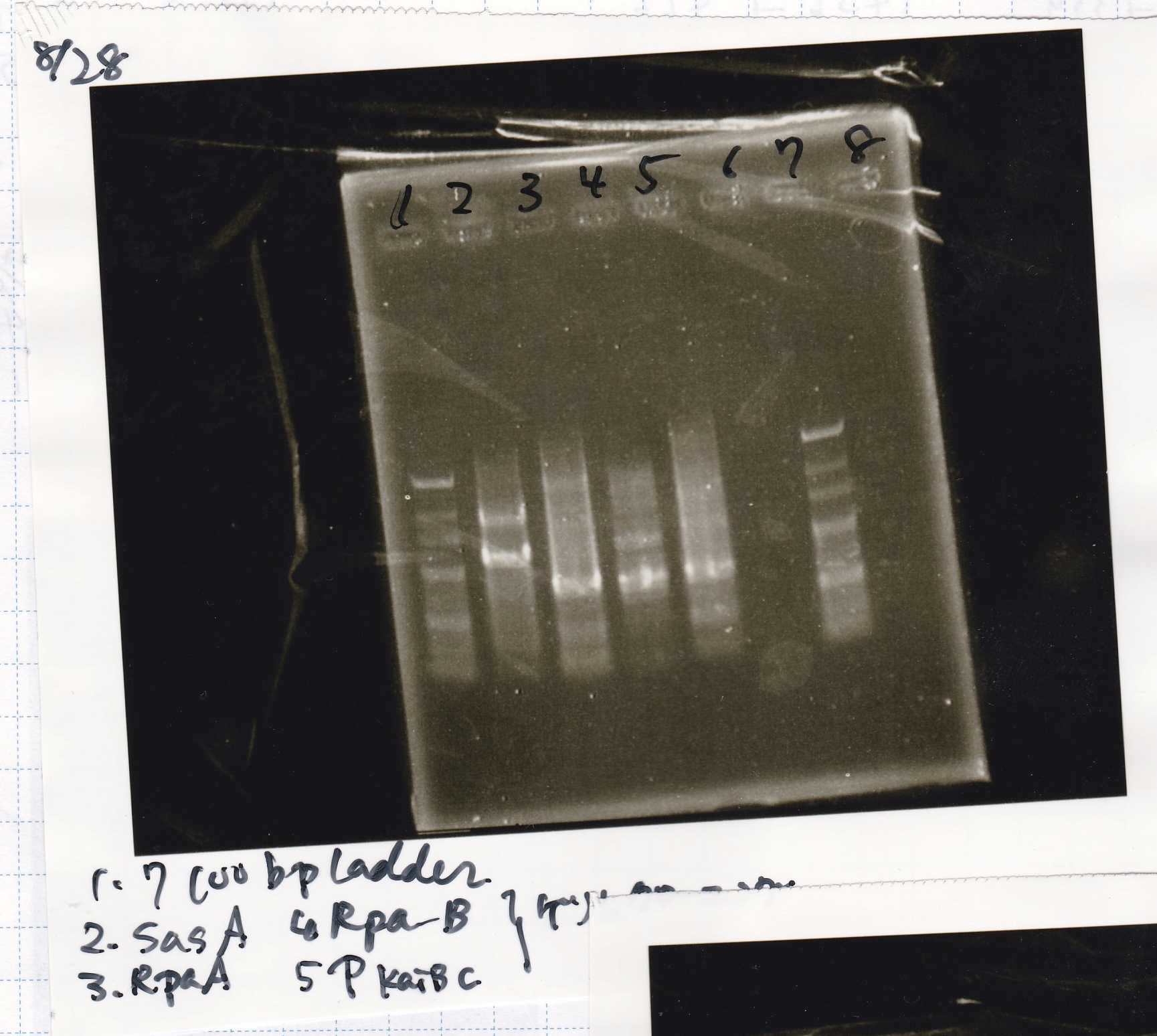
Gel Extraction
| Lane | DNA |
|---|---|
| 1 | 100bpladder |
| 3 | SasA(8/27 Genome PCR production) |
| 5 | RpaA(8/27 Genome PCR production) |
| 7 | RpaB(8/27 Genome PCR production) |
| 9 | PkaiBC(8/27 Genome PCR production) |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| RpaA | 15.4 | 1.88 | 0.03 |
| SasA | 21.6 | 1.66 | 0.79 |
| PkaiBC | 16.3 | 1.79 | 0.68 |
| RpaB | 26.7 | 1.77 | 0.82 |
Colony PCR
| Sample | base pair |
|---|---|
| 8/21 RBS-lysis2-(3) | 772 |
| 8/21 RBS-lysis2-(4) | 772 |
| 8/21 RBS-lysis2-(5) | 772 |
| 8/21 RBS-lysis2-(6) | 772 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 48s | 30cycles |
Restriction Enzyme Digestion
| 8/22 RBS-lysis1-(1) | EcoRI | SpeI | BufferB | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 9µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 16.7µL | 30µL |
| NC | 1µL | 0µL | 0µL | 1µL | 0.1µL | 7.9µL | 10µL |
| 8/17 RBS-GFP-DT-(2) | XbaI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 4µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 21.7µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 1µL | 0.1µL | 8.4µL | 10µL |
| 8/20 Pcon-RBS-GFP-DT-(1) | EcoRI | SpeI | BufferB | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 3µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 22.7µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1µL | 0.1µL | 8.6µL | 10µL |
| 8/20 Pcon-RBS-luxR-DT-(1) | EcoRI | XbaI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 2µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 23.7µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1µL | 0.1µL | 8.6µL | 10µL |
| 8/28 Plux | SpeI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 6µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 19.7µL | 30µL |
| NC | 1µL | 0µL | 0µL | 1µL | 0.1µL | 7.9µL | 10µL |
| 8/20 RBS-tetR-DT-(1) | XbaI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 5µL | 0.5µL | 0.5µL | 3µL | 0.3µL | 20.7µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 1µL | 0.1µL | 8.4µL | 10µL |
- incubated 37°C 1hour
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 8/22 RBS-lysis1-(1) | EcoRI | SpeI |
| 2 | 8/22 RBS-lysis1-(1) | -- | -- |
| 3 | 8/17 RBS-GFP-DT-(2) | XbaI | PstI |
| 4 | 8/17 RBS-GFP-DT-(2) | -- | -- |
| 5 | 8/20 Pcon-RBS-GFP-DT-(1) | EcoRI | XbaI |
| 6 | 8/20 Pcon-RBS-GFP-DT-(1) | -- | -- |
| 7 | 100bp ladder | -- | -- |
| 8 | 8/20 Pcon-RBS-luxR-DT-(1) | EcoRI | XbaI |
| 9 | 8/20 Pcon-RBS-luxR-DT-(1) | -- | -- |
| 10 | 8/28 Plux | SpeI | PstI |
| 11 | 8/28 Plux | -- | -- |
| 12 | 8/20 RBS-tetR-DT | XbaI | PstI |
| 13 | 8/20 RBS-tetR-DT | -- | -- |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2 | 8/20 RBS-lysis1-(1) | EcoRI & SpeI |
| 3 | ||
| 4 | -- | -- |
| 5 | 8/28 RBS-GFP-DT-(2) | XbaI & PstI |
| 6 |
File:Igku xxbeforexx.xxx
File:Igku xxafterxx.xxx
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2 | 8/28 Plux | SpeI & PstI |
| 3 | ||
| 4 | -- | -- |
| 5 | 8/20 RBS-tetR-DT-(1) | XbaI & PstI |
| 6 |
File:Igku xxbeforexx.xxx
File:Igku xxafterxx.xxx
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-RBS-GFP-DT (EcoRI&SpeI) | 7.1 | 1.88 | 0.07 |
| Pcon-RBS-luxR-DT (EcoRI&XbaI) | 15.3 | 1.93 | 0.70 |
| Plux (SpeI&PstI) | 4.5 | 1.79 | 0.31 |
| RBS-GFP-DT (XbaI&PstI) | 6.0 | 2.16 | 0.03 |
| RBS-tetR-DT (XbaI&PstI) | 7.6 | 1.91 | 0.38 |
Colony PCR
| Sample | base pair |
|---|---|
| 8/27 pT181 attenuator-DT-1 | 738 |
| 8/27 pT181 attenuator-DT-2 | 738 |
| 8/27 pT181 attenuator-DT-3 | 738 |
| 8/27 TetR-aptamer 12_1R-DT-1 | 521 |
| 8/27 TetR-aptamer 12_1R-DT-2 | 521 |
| 8/27 pT181 antisense-DT-1 | 542 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 30s | 30cycles |
File:Igku Aug28electrophoresis3?.png
| Sample | base pair |
|---|---|
| 8/27 Pλ-luxI-1 | ??? |
| 8/27 Pλ-luxI | ??? |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 30min | 3cycles |
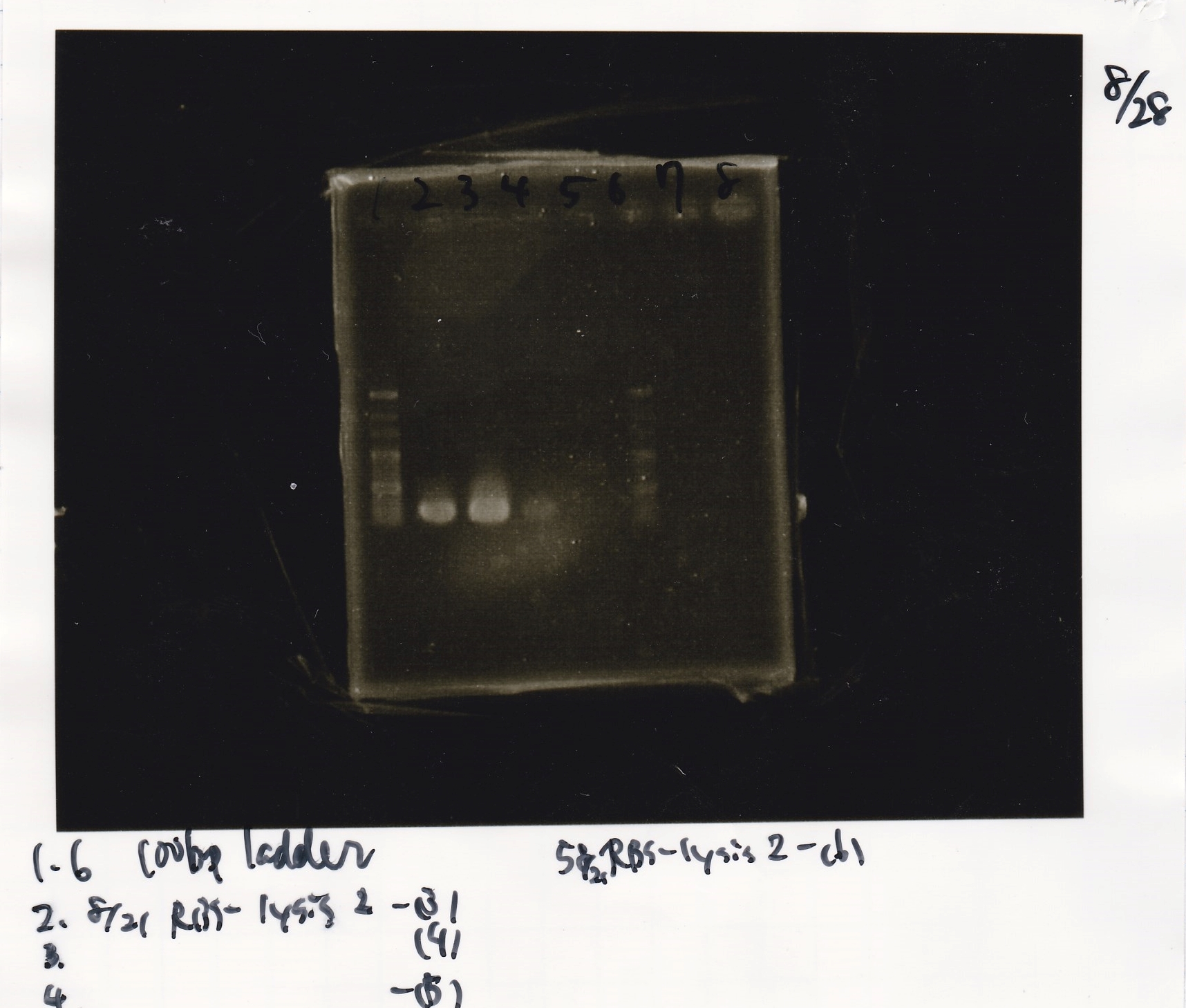
Colony PCR
No name
| Sample | base pair |
|---|---|
| 8/21 RBS-lysis2-9 | 772 |
| 8/21 RBS-lysis2-10 | 772 |
| 8/21 RBS-lysis2-11 | 772 |
| 8/21 RBS-lysis2-12 | 772 |
| 8/21 RBS-lysis2-13 | 772 |
| 8/21 RBS-lysis2-14 | 772 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 48s | 30cycles |
File:Igku Aug28electrophoresis4?.png
Liquid Culture
| Sample | medium |
|---|---|
| 8/16 Master Plate(CP)-6 DT | Plusgrow medium(+CP) |
| J23100 Master Plate-1 | Plusgrow medium(+Amp) |
| Plac Master Plate-1 | Plusgrow medium(+CP) |
Ligation
| state | Vector | Inserter | ||
|---|---|---|---|---|
| experiment | 8/27 DT (EcoRI & XbaI) | 2.9 | 8/28 RBS-lysis1 (EcoRI & SpeI) | 9.5 |
| experiment | 8/28 Plux (SpeI & PstI) | 2.3 | 8/28 RBS-GFP-DT (XbaI & PstI) | 18 |
| experiment | 8/28 Pcon-RBS-luxR-DT (EcoRI & XbaI) | 3.3 | 8/28 Pcon^RBS-GFP-DT-(1) (EcoRI & SpeI) | 28 |
| experiment | 8/27 DT (EcoRI & XbaI) | 2.7 | 8/24 Spinach (EcoRI & SpeI) | 2.4 |
- Samples were evaporeted used evaporator into about 1 µL.
| sample | MilliQ | Ligation High | total |
|---|---|---|---|
| 1 | 3 | 4 | 9 |
- incubate overnight at 4 °C
Liquid Culture
| Sample | medium |
|---|---|
| 8/26 Pλ-RBS-luxI-DT-1 | Plusgrow medium (+Amp) |
| 8/26 Pλ-RBS-luxI-DT-2 | Plusgrow medium (+Amp) |
- incubate 37°C 10hour
 "
"