Team:Biwako Nagahama/Material & Method
From 2013.igem.org
| Line 6: | Line 6: | ||
<div class="top-sentence"> | <div class="top-sentence"> | ||
= <h2>Material & Method</h2> = | = <h2>Material & Method</h2> = | ||
| - | <html> | + | == <html> |
<ul> | <ul> | ||
| - | + | <li><a href="https://2013.igem.org/Team:Biwako_Nagahama/general protocol"><h2>General protocol</h2></a></li> | |
| - | <ul> | + | <ul> |
| - | </html> | + | |
| + | |||
| + | </html> == | ||
== <h2>Genaral protocol</h2> == | == <h2>Genaral protocol</h2> == | ||
<h3>Distribution kit</h3> | <h3>Distribution kit</h3> | ||
Revision as of 17:54, 27 September 2013
Contents |
Material & Method
Genaral protocol
Distribution kit
↓With a pipette tip, punch a hole in the foil
↓Add 10μL of dH2O,and pipetting
↓Put 5min
↓Pipette 1uL of the resuspended DNA Transformation into your desired 100μL of competent cells
↓Hold on ice for 20min
↓Heat shock at 42℃ for 30sec
↓quickly
↓On ice for 2min
↓Add 900μL of SOCborth
↓Hold at 37℃ for 30min
↓Plating 100μL of DNA Transformation
↓Centrifuge for 1 min(13,000rpm)
↓Waste supernatant for 800μL, and pipetting
↓Plating all
↓Incubate at 37℃ (over night)
Phenol-chloroform extraction
Add Phenol-chloroform where is equivalent to Exo Star process sample
↓Centrifuge 4℃ 13,000rpm 5min
Only supernatant was taken
EtOH crystalization
↓Add 1μL 20mg/mL Glycogen
↓Mix
↓Add 1/10 volume 3M CH3COONa(pH5.2)
↓Add 2.5 times volume 99.5% EtOH
↓Vortex
↓Centrifuge 4℃ 13,000rpm 20min
↓Waste supernatant
↓Add 500μL 70% EtOH
↓Mix
↓Centrifuge for 10s in a table-top microcentrifuge
↓Waste supernatant
↓65℃ Dry up
↓Add 11μL TE buffer
CelC
Agro Notebook
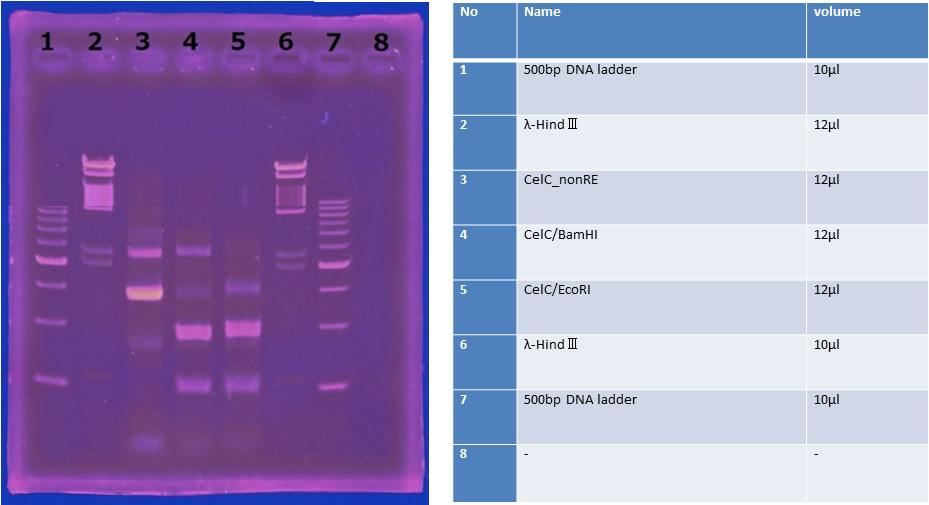
5/31 Cloning of CelC and Restriction Enzyme
By Koki Tsutsumi
CelC gene had produced clone from Agrobacterium tumefaciens C58, but I confirmed whether it’s true or not. CelC gene has restriction enzyme sites,EcoRI and BamHI.
 "
"