Team:Kyoto/projectRNA
From 2013.igem.org
(→Experiment) |
(→Reporter) |
||
| Line 62: | Line 62: | ||
我々は、RNAでできたレポーターとなりうる分子として、Spinachを挙げる。これはJeremy S. Paige, Karen Y. Wu, Samie R. Jaffrey, によって設計されたアプタマーの一種で、GFPを模倣している。SpinachはGFPの蛍光部位によく似た合成物であるDMHBIに特異的に結合するアプタマーから設計された。GFPのfluorophoreはdenatured GFPでは蛍光を示すことがなく、分子の奥に折りたたまれて初めて蛍光を発するようになる。DMHBIもこれと似た性質を持っており、単体ではほぼ蛍光を示すことはなく、GFPの構造の持つ機能を真似たSpinachの高次構造の奥に取り込まれて初めて蛍光するようになる。そのため、サンプルにDMHBIを加えた後に蛍光を確認すると、サンプル内にSpinachが存在するかどうかがわかる。もし存在すればSpinachはDFHBIと結合して蛍光を発するだろうし、存在しなければ蛍光は発しえない。Spinachを用いることで、RNAを直接イメージングできる他、安定なタンパク質では確認できない、大きく変化するRNAの発現量を正確に反映することが出来る。<br> | 我々は、RNAでできたレポーターとなりうる分子として、Spinachを挙げる。これはJeremy S. Paige, Karen Y. Wu, Samie R. Jaffrey, によって設計されたアプタマーの一種で、GFPを模倣している。SpinachはGFPの蛍光部位によく似た合成物であるDMHBIに特異的に結合するアプタマーから設計された。GFPのfluorophoreはdenatured GFPでは蛍光を示すことがなく、分子の奥に折りたたまれて初めて蛍光を発するようになる。DMHBIもこれと似た性質を持っており、単体ではほぼ蛍光を示すことはなく、GFPの構造の持つ機能を真似たSpinachの高次構造の奥に取り込まれて初めて蛍光するようになる。そのため、サンプルにDMHBIを加えた後に蛍光を確認すると、サンプル内にSpinachが存在するかどうかがわかる。もし存在すればSpinachはDFHBIと結合して蛍光を発するだろうし、存在しなければ蛍光は発しえない。Spinachを用いることで、RNAを直接イメージングできる他、安定なタンパク質では確認できない、大きく変化するRNAの発現量を正確に反映することが出来る。<br> | ||
| - | Spinach is an example of a reporter RNA aptamer,which emits the green fluorescence like GFP when it binds to a fluorophore | + | Spinach is an example of a reporter RNA aptamer,which emits the green fluorescence like GFP when it binds to a fluorophore (DFHBI), which is a derivative fluorophore of GFP. DFHBI doesn't emit fluorescence alone. That is to say, if fluorescence is observed after DFHBI is added into liquid culture, it is manifest that Spinach is expressed. If Spinach exists, it combines with DFHBI and DFHBI emits fluorescence. Hence, by using Spinach, it’s possible not only to image RNA directly, but also accurately reflect the transcription level which vary intensely, which can’t be confirmed via stable protein because RNA is degraded faster than . |
<br><br> | <br><br> | ||
| - | We suggest Spinach as a | + | We strongly suggest Spinach aptamer as a reporter of RNA. |
===Fusion=== | ===Fusion=== | ||
Revision as of 02:16, 28 September 2013
count down
Contents |
RNA Oscillator
Introduction
Motivation
Simulating cell-cell interaction model is too complicated to compute because there is a need to consider about not only intracellular condition but also more complex conditions such as positional relationship. Then we focused on intracellular condition, and considered what make this difference between dry work and wet work and make modeling and experiment closer. A study of synthetic biology shows an oscillation model which is confirmed in both dry and wet lab. Under this experiment the effect of cell division which seems to give biggest interference with oscillation cycle can be approximated into zero, therefore this circuit is robust enough. From this example, one of the solution to deal with difficulties in reconstructing dry model in wet lab is that constructing robust enough gene-circuit model to ignore the complexity by approximation. However, there are difficulties in choosing factors and under the limitation of remaining the robustness of the cycle. We worked on a consisting oscillation circuit which can be closely reproduced by computer simulation. Our goal is forming oscillation cycle in both wet and dry lab.
Oscillation
その回路として、ncRNA-mRNAの相互作用を抑制機構に持ち、RNA アプタマーとtetR Proteinを促進機構に用いた以下の様な回路を提案する。というのも、分解や生産の速度がタンパク質よりも速いRNAならば、周期の形成について考えなければならない時間幅が狭くなり、細胞分裂や濃度変動のようなモデルに影響を与える分の変動幅を無視できると考えるからだ。アウトプット機構としては、Spinachを想定する。 We propose following circuit with ncRNA-mRNA interaction as repression mechanism and RNA aptamer-tetR protein interaction as activation mechanism
This circuit generates oscillation in the following way: This circuitBefore starting the oscillation, First, tet promoter(Ptet) is repressed by TetR at the downstream of constitutive promotor. Then, the oscillator is turned on by IPTG. IPTG induces a transcription of TetR aptamer at the downstream of Plac, Spinach, and pT181 antisense at the downstream of Ptet which are transcribed. Because tetRaptamer activates tet promotor, positive feedback occurs and more and more tetRaptamer, Spinach, and Antisense are accumulated. Then, this circuit gets fluorescence. After Antisense is accumulated to some extent, tetRaptamer, at the downstream of Attenuator region, is repressed. Then, because new tetRaptamer is not created, the amount of tetRaptamer decreases quickly. So, tet promotor is repressed by tetR protein and the amount of Antisense and Spinarch falls, too. Then, this circuit loses fluorescence. After the amount of Antisense decreases sufficiently, this circuit recovers first condition. Through this cycle, this circuit acts as an oscillator.
促進、抑制の各機構と、この回路のアウトプットとしてのSpinachの機能は以下に述べる。
Repressor
We took up non-coding RNA (ncRNA) complementarily binding mRNA as an example of functional RNA which repress transcription. ncRNA in pT181 plasmid (pT181 attenuator) controls the fate of transcriptional elongation in response to an input by complementary antisense RNA. Attenuator region, which lies in 5' untranslated region of a transcript, folds into two different RNA structure. By an interaction with complementary antisense RNA, attenuator region forms Rho-independent terminator and the transcription of the downstream is stopped. Without antisense RNA, attenuator region RNA folds into an alternative structure which allow transcription of the downstream (Novick, 1989). The uniqueness of this mechanism is that it is constructed with only RNA without other small molecules. Synthetic biologists variant of it by means of nucleotide substitution etc. (Takahashi et al, 2013). In this paper, many variants ofpT181 attenuator/antisense is constructed and the attenuation rate of each variants is different. We chose this mechanism in gene repression.
Activator
We took up TetR aptamer as an example of functional RNA which activates a transcription. TetR aptamer induces tetracycline promoter (Ptet) by binding to tetracycline repressor (TetR), which represses Ptet. When TetR aptamer binds to TetR, induces conformational change of TetR. As a result, TetR cannot come to bind to tetracycline operator (tetO). We ordered MBL=IDT gene synthesis of pT181 attenuator region DNA, antisense DNA and TetR aptamer with prefix and suffix. We transferred these parts to pSB1C3 and constructed device for antisense and attenuator assay (Fig. ).
Reporter
我々は、RNAでできたレポーターとなりうる分子として、Spinachを挙げる。これはJeremy S. Paige, Karen Y. Wu, Samie R. Jaffrey, によって設計されたアプタマーの一種で、GFPを模倣している。SpinachはGFPの蛍光部位によく似た合成物であるDMHBIに特異的に結合するアプタマーから設計された。GFPのfluorophoreはdenatured GFPでは蛍光を示すことがなく、分子の奥に折りたたまれて初めて蛍光を発するようになる。DMHBIもこれと似た性質を持っており、単体ではほぼ蛍光を示すことはなく、GFPの構造の持つ機能を真似たSpinachの高次構造の奥に取り込まれて初めて蛍光するようになる。そのため、サンプルにDMHBIを加えた後に蛍光を確認すると、サンプル内にSpinachが存在するかどうかがわかる。もし存在すればSpinachはDFHBIと結合して蛍光を発するだろうし、存在しなければ蛍光は発しえない。Spinachを用いることで、RNAを直接イメージングできる他、安定なタンパク質では確認できない、大きく変化するRNAの発現量を正確に反映することが出来る。
Spinach is an example of a reporter RNA aptamer,which emits the green fluorescence like GFP when it binds to a fluorophore (DFHBI), which is a derivative fluorophore of GFP. DFHBI doesn't emit fluorescence alone. That is to say, if fluorescence is observed after DFHBI is added into liquid culture, it is manifest that Spinach is expressed. If Spinach exists, it combines with DFHBI and DFHBI emits fluorescence. Hence, by using Spinach, it’s possible not only to image RNA directly, but also accurately reflect the transcription level which vary intensely, which can’t be confirmed via stable protein because RNA is degraded faster than .
We strongly suggest Spinach aptamer as a reporter of RNA.
Fusion
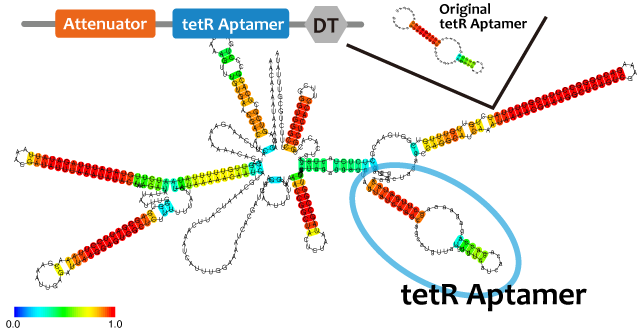
これらを使って遺伝子回路を組み立てるとき、複数のmoduleを同じ機能要素に組みこまなければならないときも十分あり得る。例えば転写抑制の様子をレポートするとき、異なる因子で促進と抑制を行うような系を作るときである。このとき、複数のモジュールを連結したことによる相互作用や立体構造の問題により機能が阻害される可能性がある。タンパク質ではその問題を予測するのは難しいが、RNAであれば配列情報から比較的簡単に二次構造を予測することができ、これらの問題を回避出来る。われわれは、機能を確認したtetR aptamer, Antisense-Attenuator RNA, をそれぞれつなぎあわせ、二次構造を予測し、実際に働いていることを確認した。tetRタンパク質存在下でtetR aptamerとAttenuator antisense RNAを組み合わせたRNAがPtetプロモーター下流のGFPの転写量を増加させるかを確認した。
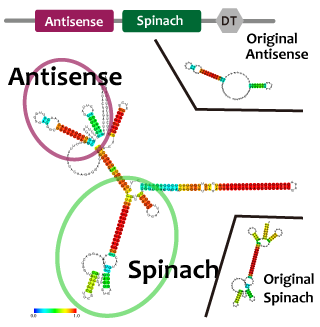
並びにAttenuator antisense RNAとSpinarchを連結したRNAを発現させ、Attenuator Region下流のGFP遺伝子の発現量が減少していることとSpinarchがDFHBI存在下で蛍光するかどうかを確認した。
Intending to check the process of transcriptional repression system and the system which promotes and represses processes of transcription by using different factors, we have to join some modules into a single RNA strand.
When we combine plural modules, the function of the modules may be inhibited by interactions and steric structures between each other. in the case of RNA,it is easier to predict and avoid the steric problems than that of proteins because we can predict the secondary structure of RNA from its primary structure. We combined tetR aptamer and Antisense-Attenuator RNA, whose functions are confirmed, and predicted secondary structures, as a result it actually worked. We also observed tetR aptamer-Attenuator antisense fusion RNA increased expression level of downstream GFP of tet promoter in the presence of tetR proteins.
-----const-------
experimental group
a. Pcon-atte-tetRaptamer-DT Ptet-GFP-DT Pcon-tetR-DT
positive control
b. Pcon-tetRaptamer-DT Ptet-GFP-DT Pcon-tetR-DT
-Fusionする前とのtetRaptamerの働きの比較
negative control
c. Ptet-GFP-DT Pcon-tetR-DT
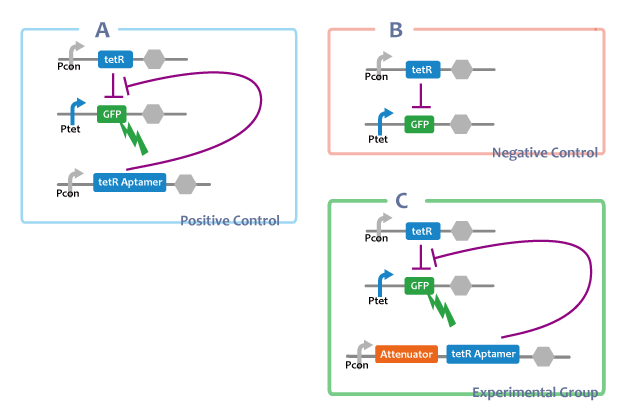
Positive Control
A. Pcon-tetRaptamer Ptet-GFP Pcon-tetR
Through this E.coli, we can confirm that separated tetRaptamer restricts the function of tetR protein.
Negative Control
B. Ptet-GFP Pcon-tetR
This E.coli shows that tetR protein represses the expression of genes at the downstream of tet promotor.
Experimental Group
C. Pcon-atte-tetRaptamer Ptet-GFP Pcon-tetR
------const-------
We used centroid fold (URL) and mfold (URL) to predict the secondary structure of RNA(a). As the picture shown below, the structure of tetR aptamer is not affected by attenuator stem loop. This suggests that the efficiency of tetR aptamer is not affected by the existence of attenuator stem loop.
Centroid fold, mfoldのfig-tetR aptamer only----tetR aptamer-antisense
tetRが常時発現されている状態では、AttenuatorとtetRaptamerを連結したRNAを転写する大腸菌(figC)のGFP発現量は、tetRaptamerを転写しない大腸菌(figB)よりも多く、ほかのRNAと連結していないtetRaptamerを転写する大腸菌(figA)と比較して{ほぼ同等 or 小さい}であることから、AttenuatorとtetRaptamerを連結すると、tetRaptamerは{全く干渉せずに機能する or 効果は下がるが機能する}ことがわかる。
If tetR is expressed, E.coli in which the united RNA(figC) was introduced expresses more GFP than E.coli which didn’t have the tetRaptamer sequence(figB). By comparing E.coli expressing independent tetRaptamer(figA) and E.coli expressing the united RNA(figC), it is convinced that tetRaptamer next door to Attenuator { works as well as independent one / works more weakly than a independent one. However, it certainly works.}
Experiment
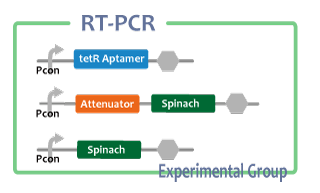
After we construct functional RNA generator, we checked whether transcription of the RNA parts. To confirm this, we performed RT-PCR.
samples are following:
Negative control
- Spinach-DT
- Pcon-tetR aptamer-DT
- Pcon-antisense-spinach-DT
- Pcon-spinach-DT
- Pcon-RBS-GFP-DT
We also check whether fusion RNA designed by us functions or not considering secondary structure with Centroid Fold[5]
Result
RT-PCR
We performed RT-PCR to confirm transcription of tetR aptamer, antisense-spinach, spinach, and GFP(GFP generator). File:ElectrophoresisRT
Structure Prediction
Conclusion
We confirmed transcription of tetR aptamer, antisense-spinach, spinach, and GFP by using RT-PCR method.
We predicted second structure of fusion RNA: atenuator-tetRaptamer and antisence-spinach with centroid fold. It seems to be expected structure and to function as expected.
We got ready for construction oscilator circuit in wet lab.
Future work
To solve simultaneous differential equations meaning oscilation model numerically, we will try to found exact values of some constants. For example, to determine binding constant between tetR and tetR aptamer, we will try to build up assay method and to get quantitative data.
After that, we will substitute the values for oscilation model and try to solve simulate. And we will continue assaying of our parts.
Then, after finishing construction of gene circuits that makes oscilation, we assay the oscilation circuit in wet lab. Our plans for the construction and assay are shown in this page
Finaly, we compair results of wet lab and dry lab and discuss a point in common/difference between the results.
Parts List
<groupparts>iGEM013 Kyoto</groupparts>
Reference
[1][http://www.nature.com/nature/journal/v456/n7221/abs/nature07389.html Jesse Stricker et al.(2008)"A fast, robust and tunable synthetic gene oscillator" Nature 456, 516-519]
[2][http://www.ncbi.nlm.nih.gov/pubmed/23761434 Melissa K. Takahashi and Julius B. Lucks.(2013)"A modular strategy for engineering orthogonal chimeric RNA transcription regulators"Nucleic Acids Research 41(15),7577-88]
[3][http://www.ncbi.nlm.nih.gov/pubmed/19246008 Anke Hunsicker et al.(2009)"An RNA aptamer that induces transcription"Chem Biol,16(2),173-180]
[4][http://www.sciencemag.org/content/333/6042/642.abstract Jeremy S. Paige et al.(2011)"RNA Mimics of Green Fluorescent Protein"Science Vol. 333 no. 6042 pp. 642-646]
[5][http://www.ncrna.org/ Functional RNA Project provided by Computational Biology Research Center (CBRC)]
 "
"