Team:Exeter/Results
From 2013.igem.org
AdamThomas (Talk | contribs) |
|||
| (89 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
<div class="row"> | <div class="row"> | ||
<div class="span" style="text-align:justify"> | <div class="span" style="text-align:justify"> | ||
| - | + | == Results == | |
=== Green Light Module === __NOTOC__ | === Green Light Module === __NOTOC__ | ||
| - | We have had marked success with our Green Light Module. Not only is the magenta pigment bright and evenly synthesised, but we have also managed to show magenta synthesis is inhibited by exposure to white light. We are currently working on showing that green light <i>specifically</i> switches off magenta production. | + | We have had marked success with our Green Light Module. Not only is the magenta pigment bright and evenly synthesised, but we have also managed to show magenta synthesis is inhibited by exposure to white light. This was shown by setting up an experiment which involved growing spread plate cultures of ''E.coli'', containing our green light sensor in two separate light conditions. The first condition was total darkness; this was inside a standard static incubator. The second condition was exposure to constant white light through the use of a light box. Both of these cultures were grown overnight (16 hours) and then analysed for magenta pigmentation. As shown in the picture below we found that the ''E.coli'' grown in total darkness had expressed magenta pigment (eforRed), whereas in white light this expression had been repressed. These results were in agreement with what we had expected, however they do not conclusively determine the green light module to be exclusively sensitive to green light. We are currently working on showing that green light <i>specifically</i> switches off magenta production (see 'planned "post-freeze" work'). |
<center><TABLE cellpadding="20"> | <center><TABLE cellpadding="20"> | ||
<TR> | <TR> | ||
| - | <TD> [[File:Centrifuged_universal_magenta.jpg|right||200px]] <center>A | + | <TD> [[File:Centrifuged_universal_magenta.jpg|right||200px]] <center>A universal vial showing <br> our vibrant magenta pigment</center> |
<TD> [[File:Green_light_module_plates.jpg|right||400px]] <center>Plates grown in the dark <br> clearly show magenta pigment</center> | <TD> [[File:Green_light_module_plates.jpg|right||400px]] <center>Plates grown in the dark <br> clearly show magenta pigment</center> | ||
<TD> [[File:Green_light_module_universals.jpg|right||435px]] <center>Comparison of liquid cultures <br> grown in dark vs. light</center> | <TD> [[File:Green_light_module_universals.jpg|right||435px]] <center>Comparison of liquid cultures <br> grown in dark vs. light</center> | ||
| Line 17: | Line 17: | ||
</TABLE></center> | </TABLE></center> | ||
| + | [[File:Green Module Graph.png|left||400px|]] | ||
| - | |||
| - | |||
| - | We | + | We chose some of the cultures that were showing the strongest colour development and remade liquid cultures of them. We then ran these through a [http://en.wikipedia.org/wiki/Ultraviolet%E2%80%93visible_spectroscopy UV-Vis Spectrophotometer] the next morning and across the day. Both cultures clearly showed an increase in absorption across the day; it is reasonably safe to assume that this is due to continued synthesis of magenta pigment. One hurdle that this data may bring up is that different cultures may have different levels of expression and pigment output. It is therefore important that further data is gathered on this area, and full characterisation of a pathway and strain is made before an accurate colour photograph is possible. On this note it will also be important to have expression levels equal to those for the red light and blue light modules; we do not want one colour being expressed far faster than the others, as this will swamp the image and not produce an accurate coliroid. Despite integrating His-tags into the design of this pathway, due to time constraints we were not able to nickel column extract the proteins to check for their production. |
| + | === Red Light Module === | ||
| + | |||
| + | <TABLE cellpadding="10" align='right'> | ||
| + | <TR> | ||
| + | <TD> [[File:Thankyoudundee.jpg|centre||250px]] | ||
| + | <TR> | ||
| + | <TD> [[File:Cyan plate.jpg|centre||250px]] | ||
| + | </TR> | ||
| + | </TABLE> | ||
| + | |||
| + | We have successfully attached the OmpR specific promoter (''ompC'') to the cyan pigment coding region, and are now working on transforming the plasmid into EnvZ deficient cells (kindly provided to us by the [https://2013.igem.org/Team:Dundee Dundee] iGEM 2013 Team). | ||
| + | This is due to OmpR and ''ompC'' being endogenously expressed in most ''E. coli'' strains as part of their osmoregulation. EnvZ is the transmembrane protein which phosphorylates OmpR (to become OmpR-P) when osmolarity in the cell changes; its phosphorylation triggers the production of outer membrane porins. In our project, we only want OmpR to be phosphorylated when the cells are exposed to green or blue light, and for phosphorylation to halt when exposed to red light. Obviously, if OmpR-P is being formed outside of these rules, we will be getting pigment production (or lack thereof) when we have not allowed it, which will ruin the images we are trying to develop. Hence, we will express the Red Light Module in EnvZ deficient cells. The plasmid for the Red Light Module contains coding for EnvZ, but also for the second half of the transmembrane protein which allows our system to be light responsive; Cph1. EnvZ and Cph1 work together to form Cph8. | ||
| + | We came across multiple issues with the assembly of this pathway. These include: misunderstanding a part description causing cells to output the wrong colour pigment, the well in the 2012 Kit Plate which we expected to be Cph8 actually coding for a human oestrogen receptor, transformation issues with home made competent cells and ligation inefficiencies brought about by attempting to ligate very small parts like [http://parts.igem.org/Part:BBa_B0034?title=Part:BBa_B0034 BBa_B0034] to longer sequences. | ||
| + | Our work on the red light pathway uncovered that the coding sequence for [http://parts.igem.org/Part:BBa_K322124 BBa_K322124] when taken from the 2012 Kit Plates is not as stated on the Registry. This part should have been an RBS and the coding region for Cph8, however sequencing revealed that it is actually the coding sequence for a human oestrogen receptor. | ||
| + | We have managed to get the cyan pigment [http://parts.igem.org/wiki/index.php?title=Part:BBa_K592011 BBa_K592011] working after some difficulties with our digestions/ligations. The pigment takes a while to develop, around 36 hours compared to magenta's overnight development, but the colour is reasonably strong and is produced evenly by the cells. However, more time has been invested in the cyan pigment which is responsive to red light (see "Planned Post-Freeze Work"). | ||
=== Red Light Module output Brick with ''BglII'' and ''BamHI'' restriction sites === | === Red Light Module output Brick with ''BglII'' and ''BamHI'' restriction sites === | ||
| Line 43: | Line 57: | ||
Our output brick for the Red Light Module includes the ''ompC'' promoter which is specific to the protein [http://en.wikipedia.org/wiki/Porin_response:_osmoregulation_in_E.coli OmpR]. This is followed by an RBS and a restriction site for <i>BglII</i> and the coding region for a [http://parts.igem.org/Part:BBa_E0040 GFP]. A second restriction site for <i>BamHI</i> follows, then a series of STOP codons to act as a terminator. The whole sequence is flanked by the iGEM prefix and suffix allowing the BioBrick to be used in 3A Assembly. | Our output brick for the Red Light Module includes the ''ompC'' promoter which is specific to the protein [http://en.wikipedia.org/wiki/Porin_response:_osmoregulation_in_E.coli OmpR]. This is followed by an RBS and a restriction site for <i>BglII</i> and the coding region for a [http://parts.igem.org/Part:BBa_E0040 GFP]. A second restriction site for <i>BamHI</i> follows, then a series of STOP codons to act as a terminator. The whole sequence is flanked by the iGEM prefix and suffix allowing the BioBrick to be used in 3A Assembly. | ||
| - | This BioBrick was built for us by IDT in their gBlock format. We designed the | + | This BioBrick was built for us by IDT in their gBlock format. We designed the BioBrick to contain a central GFP that is split between the two gBlocks, thus acting as a marker for a successful Gibson Assembly. |
| - | As the image shows, the two halves of GFP have been ligated together to form a complete, working protein in the successful Gibson | + | As the image to the left shows, the two halves of GFP have been ligated together to form a complete, working protein in the successful Gibson Assembly of our output bricks. GFP is constitutively expressed within DH5α cells as endogenous ENVZ phosphorylates endogenous OmpR as a component of osmolarity sensing. Endogenous OmpR then binds to ''ompC''. Thus our output brick is working as expected. In order to test whether the output brick can be implemented into the Red light pathway and ENVZ- strain should be used to avoid false positives. |
| - | A restriction digest was performed on | + | A restriction digest was performed on four independent Gibson assembly products with ''BamHI'' and ''BglII'' to ascertain whether the restriction sites were implemented correctly into our brick. A gel electrophoresis of the digested parts showed two clear bands: plasmid DNA with ''ompC'' promoter and suffix at ~2300, and superfolded GFP coding sequence DNA at ~750. Both bands are in their expected position. |
In order to insert a new part, we suggest following a standard primer based mutagenesis protocol to change the restriction sites to ''BamHI'' and ''BglII''. Additionally when picking transformed colonies, screen for green fluorescence. These colonies' plasmids have ligated incorrectly. Colonies' plasmids that have no florescence should be miniprepped, digested and ran on a gel to check for the correct insert. | In order to insert a new part, we suggest following a standard primer based mutagenesis protocol to change the restriction sites to ''BamHI'' and ''BglII''. Additionally when picking transformed colonies, screen for green fluorescence. These colonies' plasmids have ligated incorrectly. Colonies' plasmids that have no florescence should be miniprepped, digested and ran on a gel to check for the correct insert. | ||
| - | |||
| Line 66: | Line 79: | ||
The epoxy resin was initially far more successful. It formed a thick, clear protective layer over the cultures; when it was set, we were even able to peel the plastic off the plates. However, the resin appeared to react very aggressively with the bacteria. The colours leaked into the agar, and the colonies did not stay intact when we peeled off the plastic. | The epoxy resin was initially far more successful. It formed a thick, clear protective layer over the cultures; when it was set, we were even able to peel the plastic off the plates. However, the resin appeared to react very aggressively with the bacteria. The colours leaked into the agar, and the colonies did not stay intact when we peeled off the plastic. | ||
| - | However, after we spoke to the members of the Exeter Camera Club (see our [https://2013.igem.org/Team:Exeter/Outreach Outreach] page) they offered some alternative techniques. We were recommended to try [http://en.wikipedia.org/wiki/Gum_arabic gum arabic], which is a natural gum derived from tree sap. It has been used extensively in [http://en.wikipedia.org/wiki/Lithography lithography], certain forms of historical [http://en.wikipedia.org/wiki/Gum_printing photography], watercolours and even the food industry. As it is a natural gum of polysaccharides and glycoproteins, it should react less aggressively than the epoxy resin but still form a protective layer. However, gum arabic is water-soluble, which may be an issue with the LB agar, which contains a high water percentage. Unfortunately, as we only spoke to the Camera Club the day before the Wiki-freeze, we will be unable to upload any results from experimenting with gum arabic to our site, but hopefully we will have some pictures to bring to Lyon. | + | However, after we spoke to the members of the Exeter Camera Club (see our [https://2013.igem.org/Team:Exeter/Outreach Outreach] page) they offered some alternative techniques. We were recommended to try [http://en.wikipedia.org/wiki/Gum_arabic gum arabic], which is a natural gum derived from tree sap. It has been used extensively in [http://en.wikipedia.org/wiki/Lithography lithography], certain forms of historical [http://en.wikipedia.org/wiki/Gum_printing photography], watercolours and even the food industry. As it is a natural gum of polysaccharides and glycoproteins, it should react less aggressively than the epoxy resin but still form a protective layer. However, gum arabic is water-soluble, which may be an issue with the LB agar, which contains a high water percentage. Unfortunately, as we only spoke to the Camera Club the day before the Wiki-freeze, we will be unable to upload any results from experimenting with gum arabic to our site, but hopefully we will have some pictures to bring to Lyon. |
| - | === Planned "Post-Freeze" Work == | + | == Construction Methods == |
| + | |||
| + | We have utilised three different construction methods in our project. Please find details of each below. | ||
| + | |||
| + | ====3A Assembly==== | ||
| + | |||
| + | This is the standard iGEM assembly technique for ligating BioBricks to one another. We initially found no problems with the technique, but as we moved onto more complicated experiments and ligations, we found that the technique wasn't working especially well for us. We consulted with a variety of PhD students, our supervisors and other scientists in the department, who offered a wealth of advice on the subject. | ||
| + | |||
| + | We tested a variety of variations on the standard procedure on the iGEM website; different concentrations of DNA (it was suggested that we had been using concentrations that were too low to show up well on our gels or give good results for the ligations), different types of enzymes and buffers (we had initially been using FastDigest enzymes, but also worked with standard enzymes from New England BioLabs) and switching from Purite water to RNAase-free water. | ||
| + | |||
| + | We were lucky enough to be given several sets of new FastDigest enzymes, DNA ladders, protein purification and mini-prep kits from Thermo Scientific, which seemed to break our cycle and we suddenly started seeing successful gels and ligations. We did have limited success with the 3A Assembly technique, and would probably have had excellent results if we weren't under the time constraints of the summer. | ||
| + | |||
| + | ====Gibson Assembly (gBlock synthesised by IDT)==== | ||
| + | |||
| + | Working with IDT was an excellent decision for us; the gBlocks were simple to order, easy to assemble and worked well for us. We <i>did</i> invest a significant amount of time into check the sequences we were sending off, and correcting any problems with the sequences (hairpin bends, excessive or insufficient levels of G and C nucleotides, repeat sequences, etc), but this allowed us to feel confident in the sequences and encouraged us to look very closely at the best way to build our constructs. | ||
| + | |||
| + | ====DNA2.0 Gene Synthesis==== | ||
| + | |||
| + | Due to the time constraints posed by the iGEM project, and the size of our construct to be synthesised by DNA 2.0 being over 3kb we have not had any experience with the success of this assembly method. For future teams we recommend careful planning and serious consideration when using this method of construction. Although you are guaranteed a high quality construct, long production times can undesirable for short time-spanning projects, and costs (thus potential losses) are high. | ||
| + | |||
| + | == Modelling results == | ||
| + | |||
| + | Preliminary results suggest that the mix of colours in an individual e.coli is very unpredictable. The model was run multiple times with all sensors having the same activation rate. The mix of colours is very unstable with successive runs producing varying greatly. However the average for a cohort of cells is more stable. It is due to the relatively small number of DNA agents with which the RNAP interacts. This amplifies the effects of chance and causes the instability observed in the model. | ||
| + | |||
| + | [[Image:Multi-run.png|thumb|center|upright=5| This shows the pigment populations during 5 runs of a cell given the same initial conditions each time. It demonstrates the instability in the gene expression and also inequality in the magnitude of expression. The 'x' character lines represent the average of the 5 runs and are more stable as a results.]] | ||
| + | |||
| + | This will serve reduce the spacial control of tri-chromatic control system in E.Coli as a predictable average expression of a gene will require many individuals. For a biocamera this will reduce the pixel density and therefore the resolution. One solution to this potential problem could be to introduce many copies of the light light regulated genes. This could also serve to prevent one pathway dominating by having greater numbers of less prolific light regulated genes to compensate. | ||
| + | |||
| + | |||
| + | == Planned "Post-Freeze" Work == | ||
| + | |||
| + | The sequence coding for our blue light system was sent off to be synthesised by DNA 2.0, however despite the expected delivery date of 28/8/13, we are still awaiting its arrival. When delivered we plan to transform this into ''E.coli'' and experiment with its response to blue light. We would expect to see the inhibition of yellow pigment (amilGFP) when the light sensor is exposed to blue light. | ||
| + | |||
| + | In order to determine if our 3 light sensors are specifically responding to the correct light we plan to use acetate light filters. Using these filters will allow the ''E. coli'', containing the specific light sensors, to grow in the presence of monochromatic light (either blue, green or red). We would expect to see the production pigment when grown in the presence of either of the 2 colours that the light sensor does not respond to. However, when grown in the presence of light that the sensor is responsive to we would expect to see inhibited expression of the specific pigment. | ||
| + | |||
| + | As mentioned in the [https://2013.igem.org/Team:Exeter/Theory Theory] section of our wiki; the mechanism of our red light sensor involves the phosphorylation of OmpR by EnvZ, however, <i>E.coli</i> naturally expresses both OmpR and EnvZ. A direct result of this is false positives. An experimental method for overcoming these false positives would be to transform the plasmid containing the red light module into a strain of ''E.coli'' that is EnvZ deficient. The iGEM team from the [https://2013.igem.org/Team:Dundee University of Dundee] very kindly sent us a sample of an EnvZ- strain of ''E.coli'', thus giving us the resources to perform this experiment. | ||
=== Blue Light Module === | === Blue Light Module === | ||
| Line 75: | Line 123: | ||
[[Image:Better Diagram of Blue plasmid.png|center||800px|Image: 800 pixels]] | [[Image:Better Diagram of Blue plasmid.png|center||800px|Image: 800 pixels]] | ||
| - | === Red Light Module === | + | == Judging criteria == |
| + | |||
| + | Key: | ||
| + | |||
| + | [[file:Green_tick.png|15px]] = Completed | ||
| + | |||
| + | [[file:Yellow_tick2.png|15px]] = Will be completed at the Jamboree | ||
| + | |||
| + | [[file:Blue_arrow.png|15px]] = extra information about how the criteria was achieved | ||
| + | |||
| + | ===Bronze=== | ||
| + | |||
| + | [[file:Green_tick.png|15px]] Team registration. | ||
| + | |||
| + | [[file:Green_tick.png|15px]] Complete Judging form. | ||
| + | |||
| + | [[file:Green_tick.png|15px]] Team Wiki. | ||
| + | |||
| + | [[file:Yellow_tick2.png|17px]] Present a poster and a talk at the iGEM Jamboree. | ||
| + | |||
| + | [[file:Blue_arrow.png|15px]] This will take place on Saturday 12th October at 13:00 in Hall 2 at the European Jamboree. | ||
| + | |||
| + | [[file:Green_tick.png|15px]] Document at least one new standard BioBrick Part or Device used in your project/central to your project and submit this part to the iGEM Registry (submissions must adhere to the iGEM Registry guidelines). A new application of and outstanding documentation (quantitative data showing the Part’s/ Device’s function) of a previously existing BioBrick part in the “Experience” section of that BioBrick’s Registry entry also counts. Please note you must submit this new part to the iGEM Parts Registry. | ||
| + | |||
| + | [[file:Blue_arrow.png|15px]] The new standard BioBrick Parts used in our project were the [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 Green Light Module] and the Red Light Module output bricks, both [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 with] (contains GFP) and [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 without] (contains cyan pigment) <i>BglII</i> and <i>BamHI</i> cut sites, which were sent off to the iGEM Registry. | ||
| + | |||
| + | ===Silver=== | ||
| + | |||
| + | In addition to the Bronze Medal requirements, the following 4 goals must be achieved: | ||
| + | |||
| + | [[file:Green_tick.png|15px]] Experimentally validate that at least one new BioBrick Part or Device of your own design and construction works as expected. | ||
| + | |||
| + | [[file:Blue_arrow.png|15px]] The Green Light Module which we designed was proven to work as we grew it up in both light and dark conditions as seen in the 'Green Light Module Results page' above. Comparison of liquid cultures | ||
| + | grown in dark vs. light also showed that the magenta pigment grew up in the dark but not in the light. | ||
| + | |||
| + | [[file:Blue_arrow.png|15px]] Our output brick containing GFP and the <i>BglII</i> and <i>BamHI</i> cut sites has also given promising results, producing a strong green fluorescence and showing the expected bands when cut with <i>BglII</i> and <i>BamHI</i>. | ||
| + | |||
| + | [[file:Green_tick.png|15px]] Document the characterization of this part in the “Main Page” section of that Part’s/Device’s Registry entry. | ||
| + | |||
| + | [[file:Blue_arrow.png|15px]] [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 BBa_K1173500] | ||
| + | |||
| + | [[file:Blue_arrow.png|15px]] [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 BBa_K1173501] | ||
| - | [[ | + | [[file:Blue_arrow.png|15px]] [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 BBa_K1173502] |
| - | + | [[file:Green_tick.png|15px]] Submit this new part to the iGEM Parts Registry (submissions must adhere to the iGEM Registry guidelines). | |
| - | + | [[file:Blue_arrow.png|15px]] [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 BBa_K1173500] | |
| - | + | [[file:Blue_arrow.png|15px]] [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 BBa_K1173501] | |
| - | + | [[file:Blue_arrow.png|15px]] [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 BBa_K1173502] | |
| + | [[file:Green_tick.png|15px]] Your project may have implications for the environment, security, safety and ethics and/or ownership and sharing. Describe one or more ways in which these or other broader implications have been taken into consideration in the design and execution of your project. | ||
| - | = | + | [[file:Blue_arrow.png|15px]] We have devoted plenty of thought into the ethics and environmental issues within our project; from how we undertake the project ourselves, to how our BioCamera/bacteria could be safely used by other iGEM teams or potential users. As can be seen on our Safety Attributions page, we have implemented a variety of rules and habits in the lab to prevent unethical or unsafe use of our bacteria. As with most strains of bacteria used in R&D laboratories, the strains of <i>E, coli</i> that we utilised (DH5α and XL1 competent cells) are not designed to exist outside of the lab environment; they are grown in nutrient rich media and would be unlikely to proliferate if removed from the lab. However, if people develop their own images using our bacteria, we need to offer a technique where they can “fix” the image to preserve their picture but prevent release of the bacteria. We experimented with trapping the colonies under layers of varnish or epoxy resin with mixed [https://2013.igem.org/Team:Exeter/Results#.22Fixing.22_our_images_using_pouring_plastics results]. We have also thought about how the “camera” could be built; allowing images to develop in a completely sealed environment, much like a traditional pinhole camera. For the other potential applications of our Light Modules and output bricks, we would consider these to only be undertaken in laboratory settings, as manipulating our output bricks would require equipment and techniques (enzymes, thermocyclers, gel electrophoresis equipment, competent cells and mini-prep kits, for example) usually only available in a laboratory. |
| + | |||
| + | [[file:Blue_arrow.png|15px]]When it comes to ownership and sharing, our project was a result of inspiring work from other iGEM teams, and a desire to build on existing work within the iGEM competition. The bright, reliable pigments developed by the 2011 [https://2011.igem.org/Team:Uppsala-Sweden Uppsala] and the fantastic work on [https://2004.igem.org/austin.cgi red], [http://parts.igem.org/wiki/index.php?title=Part:BBa_K592004 blue] and [https://2009.igem.org/Team:Tokyo-Nokogen/Project/Light-receptor green] light sensors by various other teams encouraged us to combine the two aspects and attempt to build a full colour bacterial “camera”. Every part we have used has been linked to its Registry page in our [https://2013.igem.org/Team:Exeter/Notebook Notebook]. | ||
| - | == | + | ===Gold=== |
| + | In addition to the Bronze and Silver Medal requirements, any one or more of the following: | ||
| - | + | * Improve the function of an existing BioBrick Part or Device (created by another team or your own institution in a previous year), enter this information in the Registry (in the “Experience” section of that BioBrick’s Registry entry), create a new registry page for the improved part, and submit this part to the iGEM Parts Registry (submissions must adhere to the iGEM Registry guidelines). The growth of the Registry depends on having a broad base of reliable parts. This is why the improvement of an existing part is just as important as the creation and documentation of a new part. An "improvement" is anything that improves the functionality and ease-of-use of a part, so that it is more likely to be used by the community. For instance: strengthening the expression of a part by mutating the DNA sequence; modifying one or a few parts in construct (Device) so that it performs its intended job better; improving a cloning or expression vector that can be easily used by the entire community; and of course, troubleshooting and fixing a part reported to be non-functional. Data from an experimental comparison between the original and improved part/ device is strongly recommended. | |
| + | * Help any registered iGEM team from another school or institution by, for example, characterizing a part, debugging a construct, or modelling or simulating their system. | ||
| - | + | * Your project may have implications for the environment, security, safety and ethics and/or ownership and sharing. Describe a novel approach that your team has used to help you and others consider these aspects of the design and outcomes of synthetic biology efforts. Please justify its novelty and how this approach might be adapted and scaled for others to use. | |
</div> | </div> | ||
Latest revision as of 03:41, 5 October 2013
Results
Green Light Module
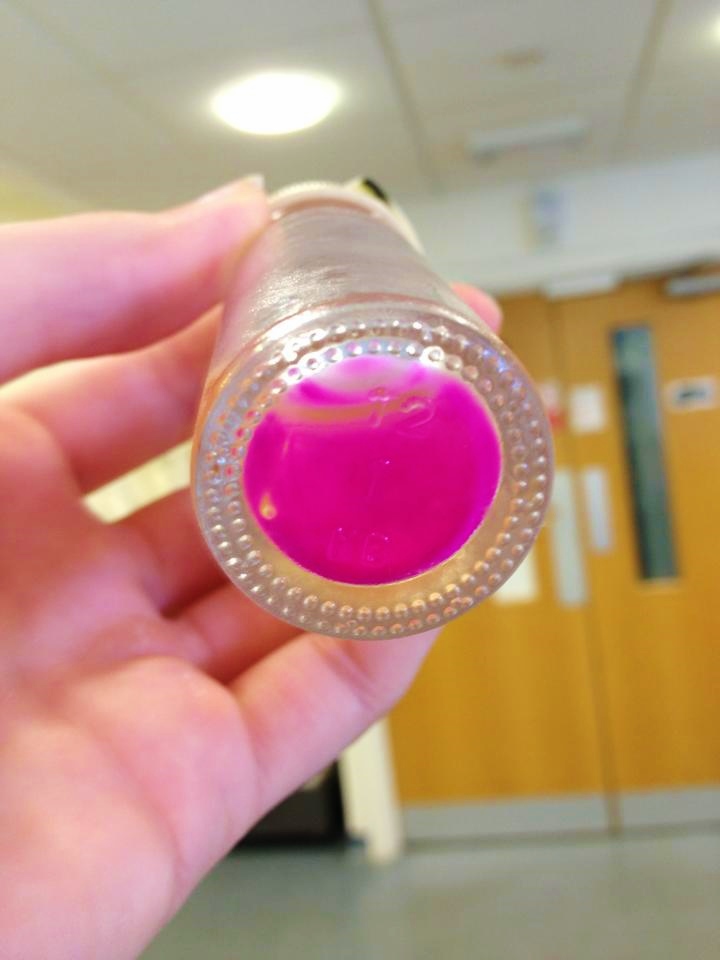
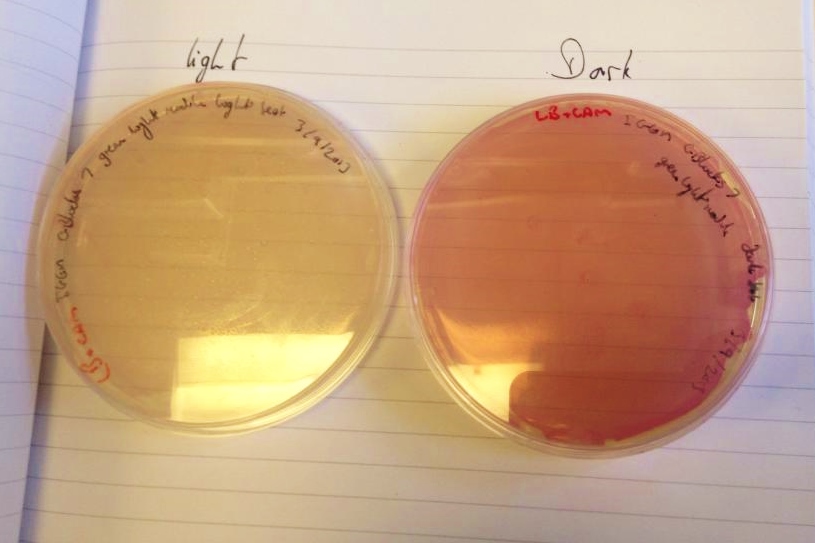
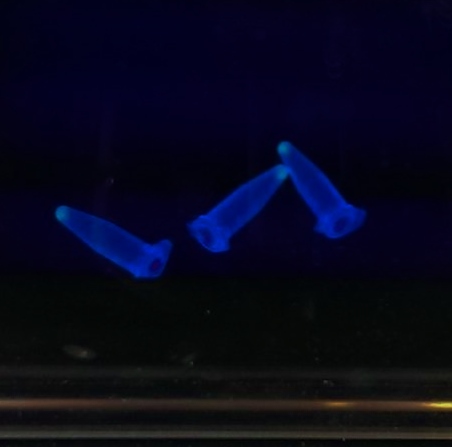
We have had marked success with our Green Light Module. Not only is the magenta pigment bright and evenly synthesised, but we have also managed to show magenta synthesis is inhibited by exposure to white light. This was shown by setting up an experiment which involved growing spread plate cultures of E.coli, containing our green light sensor in two separate light conditions. The first condition was total darkness; this was inside a standard static incubator. The second condition was exposure to constant white light through the use of a light box. Both of these cultures were grown overnight (16 hours) and then analysed for magenta pigmentation. As shown in the picture below we found that the E.coli grown in total darkness had expressed magenta pigment (eforRed), whereas in white light this expression had been repressed. These results were in agreement with what we had expected, however they do not conclusively determine the green light module to be exclusively sensitive to green light. We are currently working on showing that green light specifically switches off magenta production (see 'planned "post-freeze" work').
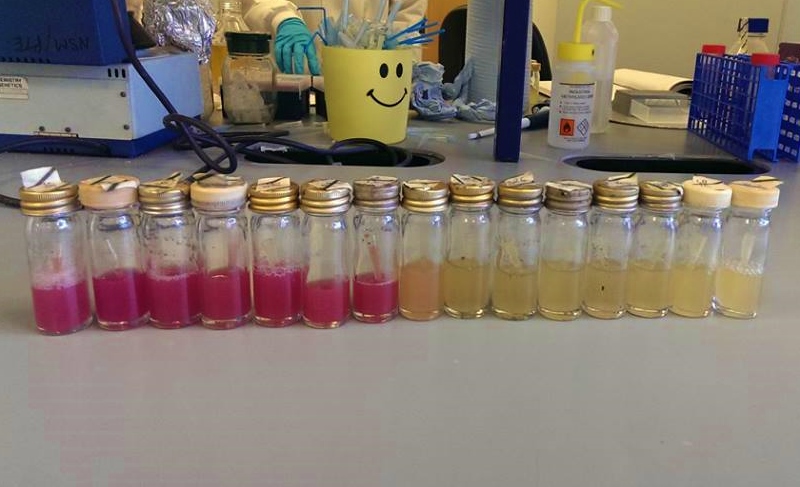
| our vibrant magenta pigment | clearly show magenta pigment | grown in dark vs. light |
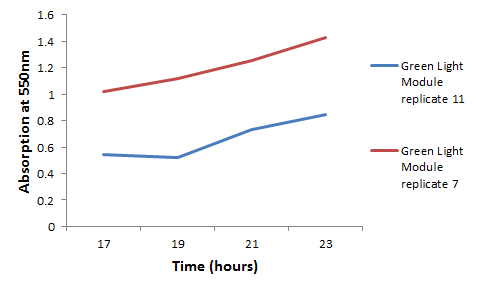
We chose some of the cultures that were showing the strongest colour development and remade liquid cultures of them. We then ran these through a [http://en.wikipedia.org/wiki/Ultraviolet%E2%80%93visible_spectroscopy UV-Vis Spectrophotometer] the next morning and across the day. Both cultures clearly showed an increase in absorption across the day; it is reasonably safe to assume that this is due to continued synthesis of magenta pigment. One hurdle that this data may bring up is that different cultures may have different levels of expression and pigment output. It is therefore important that further data is gathered on this area, and full characterisation of a pathway and strain is made before an accurate colour photograph is possible. On this note it will also be important to have expression levels equal to those for the red light and blue light modules; we do not want one colour being expressed far faster than the others, as this will swamp the image and not produce an accurate coliroid. Despite integrating His-tags into the design of this pathway, due to time constraints we were not able to nickel column extract the proteins to check for their production.
Red Light Module
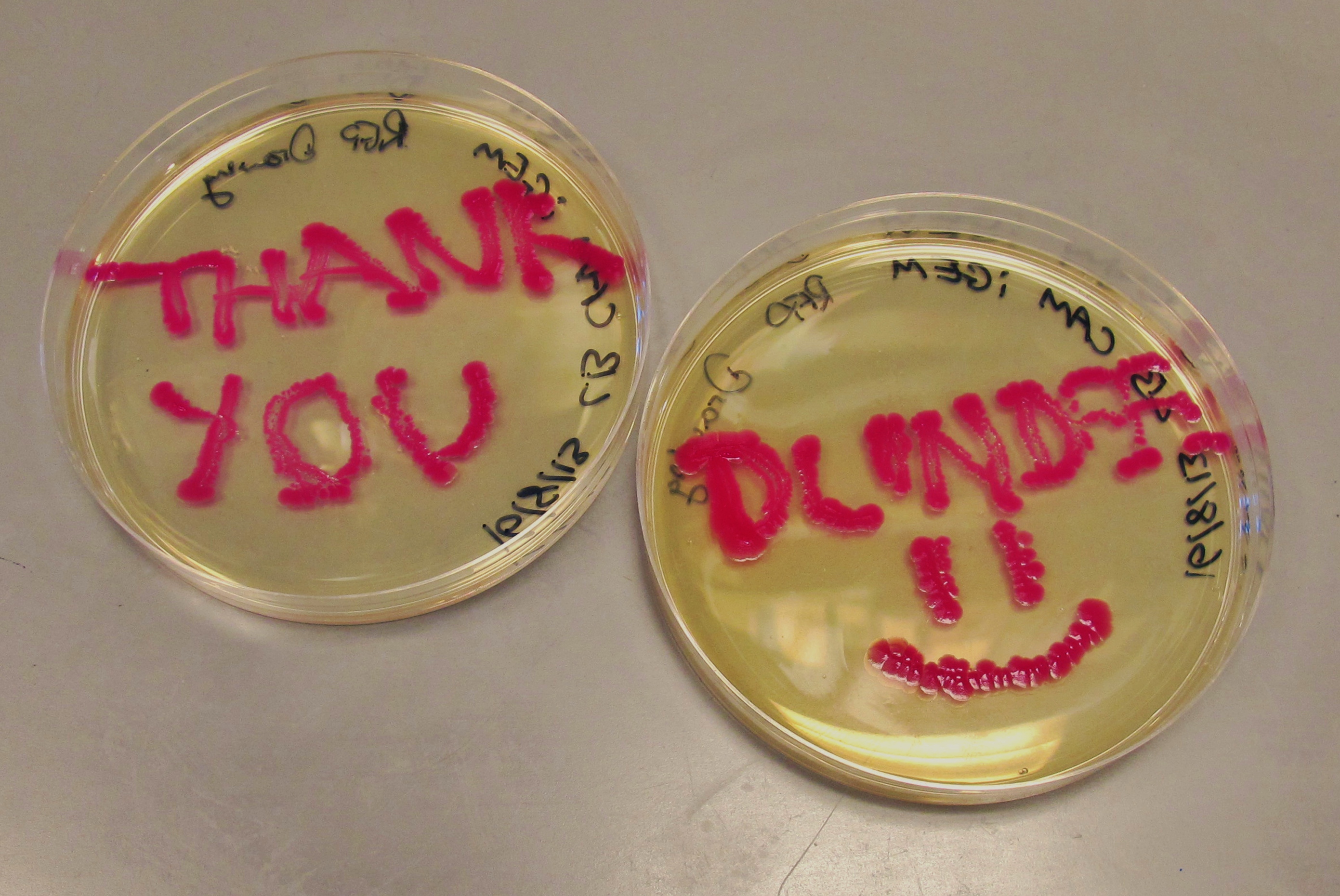
We have successfully attached the OmpR specific promoter (ompC) to the cyan pigment coding region, and are now working on transforming the plasmid into EnvZ deficient cells (kindly provided to us by the Dundee iGEM 2013 Team).
This is due to OmpR and ompC being endogenously expressed in most E. coli strains as part of their osmoregulation. EnvZ is the transmembrane protein which phosphorylates OmpR (to become OmpR-P) when osmolarity in the cell changes; its phosphorylation triggers the production of outer membrane porins. In our project, we only want OmpR to be phosphorylated when the cells are exposed to green or blue light, and for phosphorylation to halt when exposed to red light. Obviously, if OmpR-P is being formed outside of these rules, we will be getting pigment production (or lack thereof) when we have not allowed it, which will ruin the images we are trying to develop. Hence, we will express the Red Light Module in EnvZ deficient cells. The plasmid for the Red Light Module contains coding for EnvZ, but also for the second half of the transmembrane protein which allows our system to be light responsive; Cph1. EnvZ and Cph1 work together to form Cph8.
We came across multiple issues with the assembly of this pathway. These include: misunderstanding a part description causing cells to output the wrong colour pigment, the well in the 2012 Kit Plate which we expected to be Cph8 actually coding for a human oestrogen receptor, transformation issues with home made competent cells and ligation inefficiencies brought about by attempting to ligate very small parts like [http://parts.igem.org/Part:BBa_B0034?title=Part:BBa_B0034 BBa_B0034] to longer sequences.
Our work on the red light pathway uncovered that the coding sequence for [http://parts.igem.org/Part:BBa_K322124 BBa_K322124] when taken from the 2012 Kit Plates is not as stated on the Registry. This part should have been an RBS and the coding region for Cph8, however sequencing revealed that it is actually the coding sequence for a human oestrogen receptor.
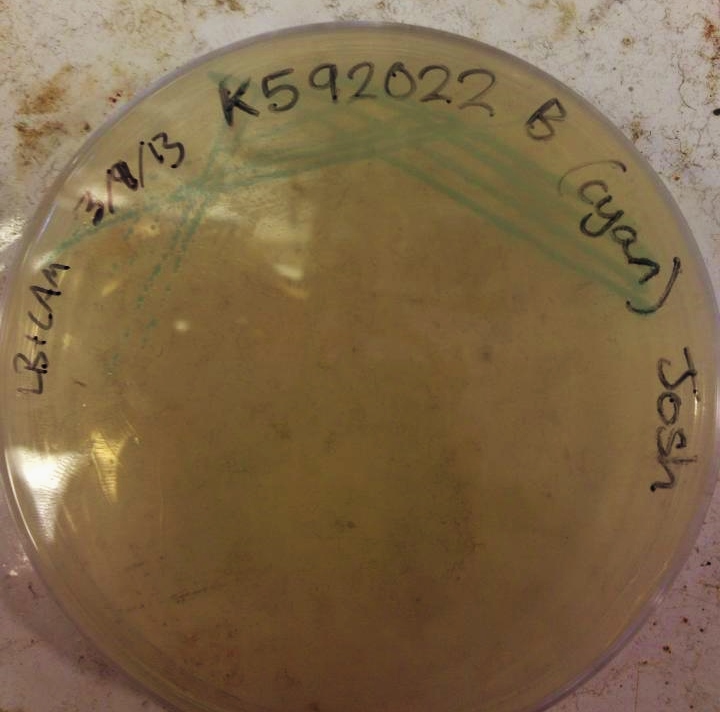
We have managed to get the cyan pigment [http://parts.igem.org/wiki/index.php?title=Part:BBa_K592011 BBa_K592011] working after some difficulties with our digestions/ligations. The pigment takes a while to develop, around 36 hours compared to magenta's overnight development, but the colour is reasonably strong and is produced evenly by the cells. However, more time has been invested in the cyan pigment which is responsive to red light (see "Planned Post-Freeze Work").
Red Light Module output Brick with BglII and BamHI restriction sites
| assembly parts showing superfolder GFP |
| output bricks showing band at around 950bp |
Our output brick for the Red Light Module includes the ompC promoter which is specific to the protein [http://en.wikipedia.org/wiki/Porin_response:_osmoregulation_in_E.coli OmpR]. This is followed by an RBS and a restriction site for BglII and the coding region for a [http://parts.igem.org/Part:BBa_E0040 GFP]. A second restriction site for BamHI follows, then a series of STOP codons to act as a terminator. The whole sequence is flanked by the iGEM prefix and suffix allowing the BioBrick to be used in 3A Assembly.
This BioBrick was built for us by IDT in their gBlock format. We designed the BioBrick to contain a central GFP that is split between the two gBlocks, thus acting as a marker for a successful Gibson Assembly.
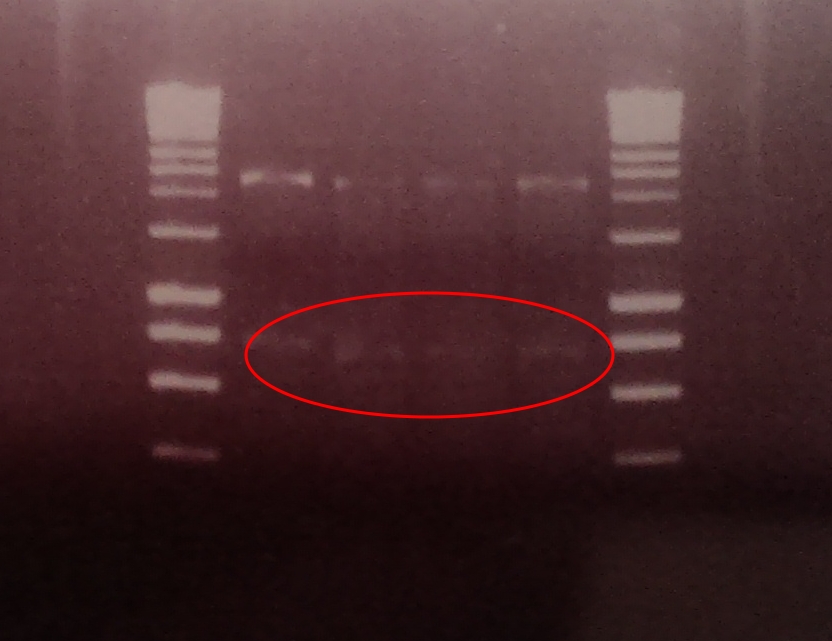
As the image to the left shows, the two halves of GFP have been ligated together to form a complete, working protein in the successful Gibson Assembly of our output bricks. GFP is constitutively expressed within DH5α cells as endogenous ENVZ phosphorylates endogenous OmpR as a component of osmolarity sensing. Endogenous OmpR then binds to ompC. Thus our output brick is working as expected. In order to test whether the output brick can be implemented into the Red light pathway and ENVZ- strain should be used to avoid false positives.
A restriction digest was performed on four independent Gibson assembly products with BamHI and BglII to ascertain whether the restriction sites were implemented correctly into our brick. A gel electrophoresis of the digested parts showed two clear bands: plasmid DNA with ompC promoter and suffix at ~2300, and superfolded GFP coding sequence DNA at ~750. Both bands are in their expected position.
In order to insert a new part, we suggest following a standard primer based mutagenesis protocol to change the restriction sites to BamHI and BglII. Additionally when picking transformed colonies, screen for green fluorescence. These colonies' plasmids have ligated incorrectly. Colonies' plasmids that have no florescence should be miniprepped, digested and ran on a gel to check for the correct insert.
"Fixing" our images using pouring plastics
When working with modified organisms such as our colourful E. coli, you must always take into account that their exposure to the environment must be minimised. Once our bacterial photographs have "developed", we want to be able to preserve the image but prevent release of the bacteria into the environment. Considering that, once the image had been made, we can't reverse the process or use the bacteria to generate another picture, we can utilise techniques to fix the image with no concerns about keeping the E. coli alive.
Our initial idea was to insert a "kill-switch" into the bacteria to switch of the image development at a chosen moment, then seal the plates they were grown on to prevent any unwanted release. However, this seemed overly complicated considering that all we needed to do was kill the bacteria then seal them into a contained environment. We looked instead at trapping the cultures between layers of plastic.
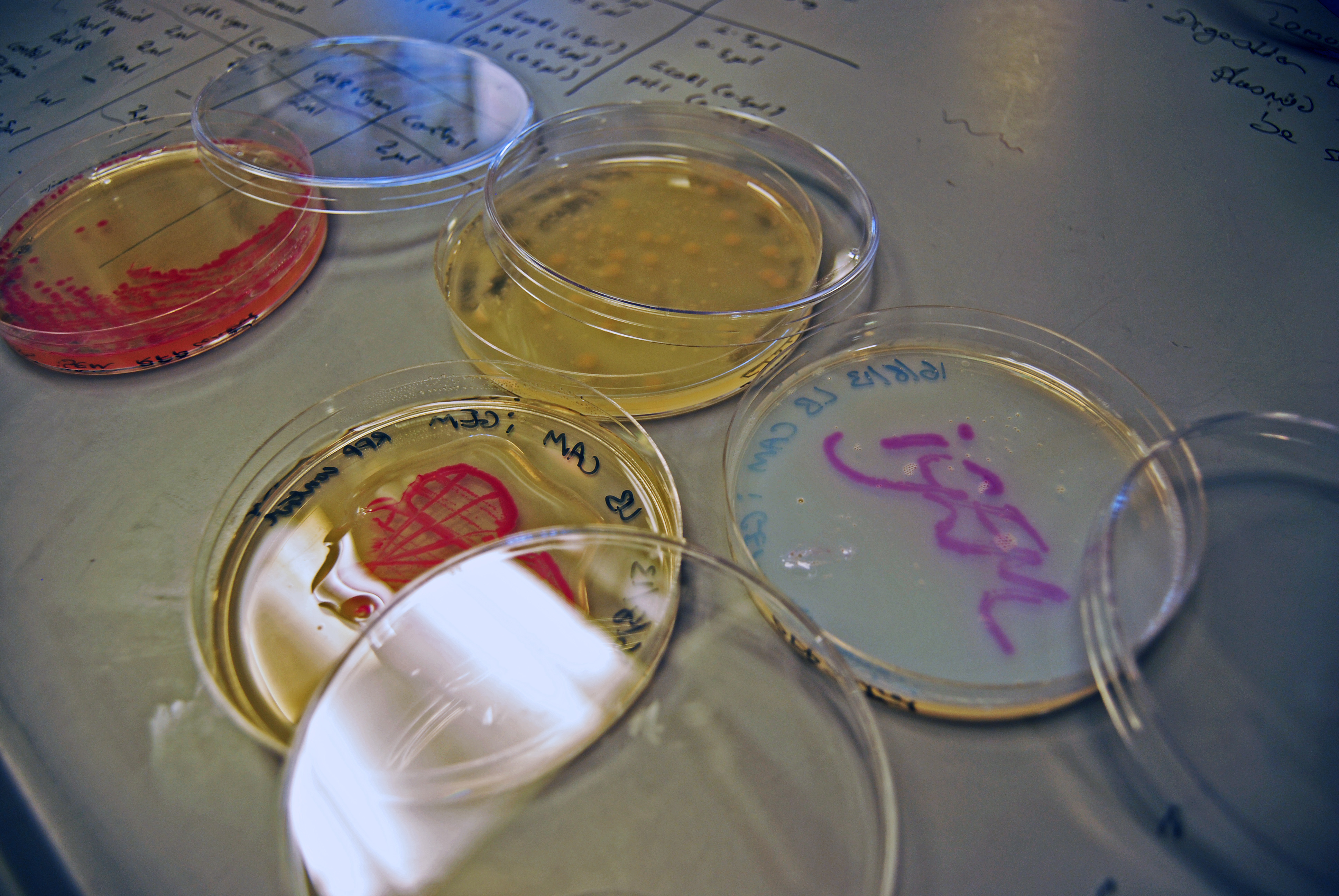
Our first test were using cultures of RFP, which exhibits a rich pink colour when grown overnight on LB agar plates overnight. We looked at using a clear, water based varnish (which is usually used for preserving craft works) and a thick, viscous epoxy resin.
The clear varnish was not particularly successful. It seems like the reasonably high water content of the agar plates prevented the varnish from setting, and instead of being clear, the varnish was cloudy and difficult to see through (see images).
The epoxy resin was initially far more successful. It formed a thick, clear protective layer over the cultures; when it was set, we were even able to peel the plastic off the plates. However, the resin appeared to react very aggressively with the bacteria. The colours leaked into the agar, and the colonies did not stay intact when we peeled off the plastic.
However, after we spoke to the members of the Exeter Camera Club (see our Outreach page) they offered some alternative techniques. We were recommended to try [http://en.wikipedia.org/wiki/Gum_arabic gum arabic], which is a natural gum derived from tree sap. It has been used extensively in [http://en.wikipedia.org/wiki/Lithography lithography], certain forms of historical [http://en.wikipedia.org/wiki/Gum_printing photography], watercolours and even the food industry. As it is a natural gum of polysaccharides and glycoproteins, it should react less aggressively than the epoxy resin but still form a protective layer. However, gum arabic is water-soluble, which may be an issue with the LB agar, which contains a high water percentage. Unfortunately, as we only spoke to the Camera Club the day before the Wiki-freeze, we will be unable to upload any results from experimenting with gum arabic to our site, but hopefully we will have some pictures to bring to Lyon.
Construction Methods
We have utilised three different construction methods in our project. Please find details of each below.
3A Assembly
This is the standard iGEM assembly technique for ligating BioBricks to one another. We initially found no problems with the technique, but as we moved onto more complicated experiments and ligations, we found that the technique wasn't working especially well for us. We consulted with a variety of PhD students, our supervisors and other scientists in the department, who offered a wealth of advice on the subject.
We tested a variety of variations on the standard procedure on the iGEM website; different concentrations of DNA (it was suggested that we had been using concentrations that were too low to show up well on our gels or give good results for the ligations), different types of enzymes and buffers (we had initially been using FastDigest enzymes, but also worked with standard enzymes from New England BioLabs) and switching from Purite water to RNAase-free water.
We were lucky enough to be given several sets of new FastDigest enzymes, DNA ladders, protein purification and mini-prep kits from Thermo Scientific, which seemed to break our cycle and we suddenly started seeing successful gels and ligations. We did have limited success with the 3A Assembly technique, and would probably have had excellent results if we weren't under the time constraints of the summer.
Gibson Assembly (gBlock synthesised by IDT)
Working with IDT was an excellent decision for us; the gBlocks were simple to order, easy to assemble and worked well for us. We did invest a significant amount of time into check the sequences we were sending off, and correcting any problems with the sequences (hairpin bends, excessive or insufficient levels of G and C nucleotides, repeat sequences, etc), but this allowed us to feel confident in the sequences and encouraged us to look very closely at the best way to build our constructs.
DNA2.0 Gene Synthesis
Due to the time constraints posed by the iGEM project, and the size of our construct to be synthesised by DNA 2.0 being over 3kb we have not had any experience with the success of this assembly method. For future teams we recommend careful planning and serious consideration when using this method of construction. Although you are guaranteed a high quality construct, long production times can undesirable for short time-spanning projects, and costs (thus potential losses) are high.
Modelling results
Preliminary results suggest that the mix of colours in an individual e.coli is very unpredictable. The model was run multiple times with all sensors having the same activation rate. The mix of colours is very unstable with successive runs producing varying greatly. However the average for a cohort of cells is more stable. It is due to the relatively small number of DNA agents with which the RNAP interacts. This amplifies the effects of chance and causes the instability observed in the model.

This will serve reduce the spacial control of tri-chromatic control system in E.Coli as a predictable average expression of a gene will require many individuals. For a biocamera this will reduce the pixel density and therefore the resolution. One solution to this potential problem could be to introduce many copies of the light light regulated genes. This could also serve to prevent one pathway dominating by having greater numbers of less prolific light regulated genes to compensate.
Planned "Post-Freeze" Work
The sequence coding for our blue light system was sent off to be synthesised by DNA 2.0, however despite the expected delivery date of 28/8/13, we are still awaiting its arrival. When delivered we plan to transform this into E.coli and experiment with its response to blue light. We would expect to see the inhibition of yellow pigment (amilGFP) when the light sensor is exposed to blue light.
In order to determine if our 3 light sensors are specifically responding to the correct light we plan to use acetate light filters. Using these filters will allow the E. coli, containing the specific light sensors, to grow in the presence of monochromatic light (either blue, green or red). We would expect to see the production pigment when grown in the presence of either of the 2 colours that the light sensor does not respond to. However, when grown in the presence of light that the sensor is responsive to we would expect to see inhibited expression of the specific pigment.
As mentioned in the Theory section of our wiki; the mechanism of our red light sensor involves the phosphorylation of OmpR by EnvZ, however, E.coli naturally expresses both OmpR and EnvZ. A direct result of this is false positives. An experimental method for overcoming these false positives would be to transform the plasmid containing the red light module into a strain of E.coli that is EnvZ deficient. The iGEM team from the University of Dundee very kindly sent us a sample of an EnvZ- strain of E.coli, thus giving us the resources to perform this experiment.
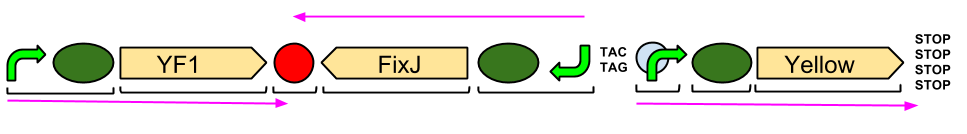
Blue Light Module
Our Blue Light Module, which is being built by DNA2.0, will unfortunately not arrive in the labs until after the Wiki-freeze. However, we will be bringing plenty of pictures of it working (hopefully!) to the Jamboree. Considering it has been constructed in a very similar manner to the Green Light Module, which is functioning in the lab, we are reasonably hopefully that we can have the Blue Light Module functioning in the lab soon after its arrival.
Judging criteria
Key:
![]() = Will be completed at the Jamboree
= Will be completed at the Jamboree
![]() = extra information about how the criteria was achieved
= extra information about how the criteria was achieved
Bronze
Team registration.
Complete Judging form.
Team Wiki.
Present a poster and a talk at the iGEM Jamboree.
![]() This will take place on Saturday 12th October at 13:00 in Hall 2 at the European Jamboree.
This will take place on Saturday 12th October at 13:00 in Hall 2 at the European Jamboree.
Document at least one new standard BioBrick Part or Device used in your project/central to your project and submit this part to the iGEM Registry (submissions must adhere to the iGEM Registry guidelines). A new application of and outstanding documentation (quantitative data showing the Part’s/ Device’s function) of a previously existing BioBrick part in the “Experience” section of that BioBrick’s Registry entry also counts. Please note you must submit this new part to the iGEM Parts Registry.
![]() The new standard BioBrick Parts used in our project were the [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 Green Light Module] and the Red Light Module output bricks, both [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 with] (contains GFP) and [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 without] (contains cyan pigment) BglII and BamHI cut sites, which were sent off to the iGEM Registry.
The new standard BioBrick Parts used in our project were the [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 Green Light Module] and the Red Light Module output bricks, both [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 with] (contains GFP) and [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 without] (contains cyan pigment) BglII and BamHI cut sites, which were sent off to the iGEM Registry.
Silver
In addition to the Bronze Medal requirements, the following 4 goals must be achieved:
Experimentally validate that at least one new BioBrick Part or Device of your own design and construction works as expected.
![]() The Green Light Module which we designed was proven to work as we grew it up in both light and dark conditions as seen in the 'Green Light Module Results page' above. Comparison of liquid cultures
grown in dark vs. light also showed that the magenta pigment grew up in the dark but not in the light.
The Green Light Module which we designed was proven to work as we grew it up in both light and dark conditions as seen in the 'Green Light Module Results page' above. Comparison of liquid cultures
grown in dark vs. light also showed that the magenta pigment grew up in the dark but not in the light.
![]() Our output brick containing GFP and the BglII and BamHI cut sites has also given promising results, producing a strong green fluorescence and showing the expected bands when cut with BglII and BamHI.
Our output brick containing GFP and the BglII and BamHI cut sites has also given promising results, producing a strong green fluorescence and showing the expected bands when cut with BglII and BamHI.
Document the characterization of this part in the “Main Page” section of that Part’s/Device’s Registry entry.
![]() [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 BBa_K1173500]
[http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 BBa_K1173500]
![]() [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 BBa_K1173501]
[http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 BBa_K1173501]
![]() [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 BBa_K1173502]
[http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 BBa_K1173502]
Submit this new part to the iGEM Parts Registry (submissions must adhere to the iGEM Registry guidelines).
![]() [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 BBa_K1173500]
[http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173500 BBa_K1173500]
![]() [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 BBa_K1173501]
[http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173501 BBa_K1173501]
![]() [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 BBa_K1173502]
[http://parts.igem.org/wiki/index.php?title=Part:BBa_K1173502 BBa_K1173502]
Your project may have implications for the environment, security, safety and ethics and/or ownership and sharing. Describe one or more ways in which these or other broader implications have been taken into consideration in the design and execution of your project.
![]() We have devoted plenty of thought into the ethics and environmental issues within our project; from how we undertake the project ourselves, to how our BioCamera/bacteria could be safely used by other iGEM teams or potential users. As can be seen on our Safety Attributions page, we have implemented a variety of rules and habits in the lab to prevent unethical or unsafe use of our bacteria. As with most strains of bacteria used in R&D laboratories, the strains of E, coli that we utilised (DH5α and XL1 competent cells) are not designed to exist outside of the lab environment; they are grown in nutrient rich media and would be unlikely to proliferate if removed from the lab. However, if people develop their own images using our bacteria, we need to offer a technique where they can “fix” the image to preserve their picture but prevent release of the bacteria. We experimented with trapping the colonies under layers of varnish or epoxy resin with mixed results. We have also thought about how the “camera” could be built; allowing images to develop in a completely sealed environment, much like a traditional pinhole camera. For the other potential applications of our Light Modules and output bricks, we would consider these to only be undertaken in laboratory settings, as manipulating our output bricks would require equipment and techniques (enzymes, thermocyclers, gel electrophoresis equipment, competent cells and mini-prep kits, for example) usually only available in a laboratory.
We have devoted plenty of thought into the ethics and environmental issues within our project; from how we undertake the project ourselves, to how our BioCamera/bacteria could be safely used by other iGEM teams or potential users. As can be seen on our Safety Attributions page, we have implemented a variety of rules and habits in the lab to prevent unethical or unsafe use of our bacteria. As with most strains of bacteria used in R&D laboratories, the strains of E, coli that we utilised (DH5α and XL1 competent cells) are not designed to exist outside of the lab environment; they are grown in nutrient rich media and would be unlikely to proliferate if removed from the lab. However, if people develop their own images using our bacteria, we need to offer a technique where they can “fix” the image to preserve their picture but prevent release of the bacteria. We experimented with trapping the colonies under layers of varnish or epoxy resin with mixed results. We have also thought about how the “camera” could be built; allowing images to develop in a completely sealed environment, much like a traditional pinhole camera. For the other potential applications of our Light Modules and output bricks, we would consider these to only be undertaken in laboratory settings, as manipulating our output bricks would require equipment and techniques (enzymes, thermocyclers, gel electrophoresis equipment, competent cells and mini-prep kits, for example) usually only available in a laboratory.
![]() When it comes to ownership and sharing, our project was a result of inspiring work from other iGEM teams, and a desire to build on existing work within the iGEM competition. The bright, reliable pigments developed by the 2011 Uppsala and the fantastic work on red, [http://parts.igem.org/wiki/index.php?title=Part:BBa_K592004 blue] and green light sensors by various other teams encouraged us to combine the two aspects and attempt to build a full colour bacterial “camera”. Every part we have used has been linked to its Registry page in our Notebook.
When it comes to ownership and sharing, our project was a result of inspiring work from other iGEM teams, and a desire to build on existing work within the iGEM competition. The bright, reliable pigments developed by the 2011 Uppsala and the fantastic work on red, [http://parts.igem.org/wiki/index.php?title=Part:BBa_K592004 blue] and green light sensors by various other teams encouraged us to combine the two aspects and attempt to build a full colour bacterial “camera”. Every part we have used has been linked to its Registry page in our Notebook.
Gold
In addition to the Bronze and Silver Medal requirements, any one or more of the following:
- Improve the function of an existing BioBrick Part or Device (created by another team or your own institution in a previous year), enter this information in the Registry (in the “Experience” section of that BioBrick’s Registry entry), create a new registry page for the improved part, and submit this part to the iGEM Parts Registry (submissions must adhere to the iGEM Registry guidelines). The growth of the Registry depends on having a broad base of reliable parts. This is why the improvement of an existing part is just as important as the creation and documentation of a new part. An "improvement" is anything that improves the functionality and ease-of-use of a part, so that it is more likely to be used by the community. For instance: strengthening the expression of a part by mutating the DNA sequence; modifying one or a few parts in construct (Device) so that it performs its intended job better; improving a cloning or expression vector that can be easily used by the entire community; and of course, troubleshooting and fixing a part reported to be non-functional. Data from an experimental comparison between the original and improved part/ device is strongly recommended.
- Help any registered iGEM team from another school or institution by, for example, characterizing a part, debugging a construct, or modelling or simulating their system.
- Your project may have implications for the environment, security, safety and ethics and/or ownership and sharing. Describe a novel approach that your team has used to help you and others consider these aspects of the design and outcomes of synthetic biology efforts. Please justify its novelty and how this approach might be adapted and scaled for others to use.
 "
"