Team:Penn/AssayOverview
From 2013.igem.org
(Difference between revisions)
| Line 38: | Line 38: | ||
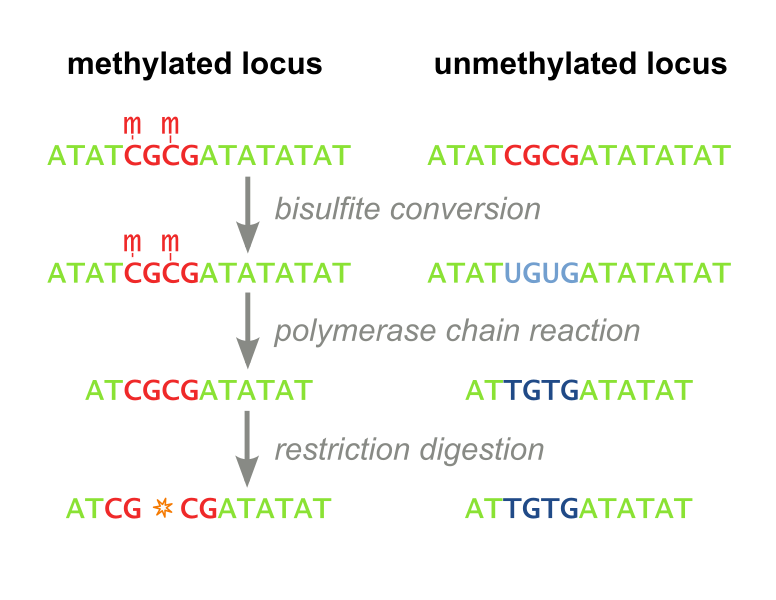
<i>Restriction Based.</i> The first, called Combined Bisulfite Restriction Analysis (COBRA), involves chemically converting unmethylated cytosines into uracils (a process called bisulfite conversion), while leaving methylated cytosines intact. Performing PCR that amplifies the region of interest leaves methylated cytosines intact and unmethylated cytosines converted to thymines (Figure 1). Samples are then digested using an enzyme that will only cut the unconverted (originally methylated) cytosines. The enzyme can no longer recognize unmethylated sites, as they are “TG” instead of “CG”. Designing primers for this process is not always feasible, even with the help of advanced algorithms, and the process needs to be optimized each time a new site is to be analyzed. Furthermore, the workflow takes several days, is expensive, and is not high throughput enough to accomodate screening libraries of candidate DNA-binding-domain-methyltransferase fusion proteins. It has recently fallen out of favor because it is difficult to interpret and does not consider all CpG sites, but only ones which fall within a restriction enzyme’s recognition sequence. (Xiong 1997 and Li 2002). Our methylation assay, MaGellin, is also restriction-based but is much simpler than COBRA because it does not require bisulfite conversion of the DNA. This eliminates most of the problems that made COBRA unwieldy. | <i>Restriction Based.</i> The first, called Combined Bisulfite Restriction Analysis (COBRA), involves chemically converting unmethylated cytosines into uracils (a process called bisulfite conversion), while leaving methylated cytosines intact. Performing PCR that amplifies the region of interest leaves methylated cytosines intact and unmethylated cytosines converted to thymines (Figure 1). Samples are then digested using an enzyme that will only cut the unconverted (originally methylated) cytosines. The enzyme can no longer recognize unmethylated sites, as they are “TG” instead of “CG”. Designing primers for this process is not always feasible, even with the help of advanced algorithms, and the process needs to be optimized each time a new site is to be analyzed. Furthermore, the workflow takes several days, is expensive, and is not high throughput enough to accomodate screening libraries of candidate DNA-binding-domain-methyltransferase fusion proteins. It has recently fallen out of favor because it is difficult to interpret and does not consider all CpG sites, but only ones which fall within a restriction enzyme’s recognition sequence. (Xiong 1997 and Li 2002). Our methylation assay, MaGellin, is also restriction-based but is much simpler than COBRA because it does not require bisulfite conversion of the DNA. This eliminates most of the problems that made COBRA unwieldy. | ||
</br> | </br> | ||
| - | <div style="margin-left:auto;margin-right:auto;text-align:center"><figure><img border="0" src="http://upload.wikimedia.org/wikipedia/commons/thumb/f/fe/Cobra_workflow.svg/776px-Cobra_workflow.svg.png" alt="COBRA Methylation Assay" width="400" ><figcaption><i> Figure 1: COBRA. This restriction based assay detects methylation at CpG sites that fall within restriction enzyme recognition sequences.</i></figcaption></figure></div> | + | <div style="margin-left:auto;margin-right:auto;text-align:center"><figure><img border="0" src="http://upload.wikimedia.org/wikipedia/commons/thumb/f/fe/Cobra_workflow.svg/776px-Cobra_workflow.svg.png" alt="COBRA Methylation Assay" width="400" > |
| + | </br> | ||
| + | <figure><img border="0" src="http://upload.wikimedia.org/wikipedia/commons/thumb/7/7f/Cobra_quantification.svg/776px-Cobra_quantification.svg.png" width="400" > | ||
| + | </figure> | ||
| + | |||
| + | <figcaption><i> Figure 1: COBRA. This restriction based assay detects methylation at CpG sites that fall within restriction enzyme recognition sequences.</i></figcaption></figure></div> | ||
| + | </br> | ||
</br> | </br> | ||
Revision as of 10:37, 27 October 2013
Assay Overview

 "
"