Template:Kyoto/Notebook/Sep 13
From 2013.igem.org
(Difference between revisions)
(→Restriction Enzyme Digestion) |
(→PCR) |
||
| (2 intermediate revisions not shown) | |||
| Line 235: | Line 235: | ||
|1cut(SpeI)||23.5||0µL||0.5µL||0µL||0µL||3µL||3µL||0µL||30µL | |1cut(SpeI)||23.5||0µL||0.5µL||0µL||0µL||3µL||3µL||0µL||30µL | ||
|- | |- | ||
| - | |NC||02.3||0µL | + | |NC||02.3||0µL||0µL||0µL||0µL||1µL||1µL||5.7µL||10µL |
|} | |} | ||
| Line 332: | Line 332: | ||
!genome DNA||Big Dye||MilliQ||5x buffer||temp(400mg)||SasA_fwd primer||SasA_rev primer||total | !genome DNA||Big Dye||MilliQ||5x buffer||temp(400mg)||SasA_fwd primer||SasA_rev primer||total | ||
|- | |- | ||
| - | |8/20 J23100-RBS-lacZα-DT||0.5||6.35||1.75||1.4||0.5||10.5 | + | |8/20 J23100-RBS-lacZα-DT||0.5||6.35||1.75||1.4||0.5||0||10.5 |
|} | |} | ||
{| class="wikitable" | {| class="wikitable" | ||
| Line 347: | Line 347: | ||
!genome DNA||Big Dye||MilliQ||5x buffer||temp(400mg)||SasA_fwd primer||SasA_rev primer||total | !genome DNA||Big Dye||MilliQ||5x buffer||temp(400mg)||SasA_fwd primer||SasA_rev primer||total | ||
|- | |- | ||
| - | |9/11 RBS-lysis1-DT||0.5||6.05||1.75||1.7||0.5||10.5 | + | |9/11 RBS-lysis1-DT||0.5||6.05||1.75||1.7||0.5||0||10.5 |
|} | |} | ||
{| class="wikitable" | {| class="wikitable" | ||
| Line 390: | Line 390: | ||
===PCR=== | ===PCR=== | ||
<div class="experiment"> | <div class="experiment"> | ||
| - | <span class="author"> | + | <span class="author">Yoshida</span> |
{| class="wikitable" | {| class="wikitable" | ||
| - | !genome DNA||KOD plus||10x buffer||dNTP||MgSO4|| | + | !genome DNA||KOD plus||10x buffer||dNTP||MgSO4||Prefix-RBS-RpaB_fwd primer||RpaB-GG4-suffix_rev primer||MilliQ||total |
|- | |- | ||
|RpaB||0.5||2.5||2.5||1.5||0.75||0.75||16.5||25 | |RpaB||0.5||2.5||2.5||1.5||0.75||0.75||16.5||25 | ||
|} | |} | ||
| + | {| class="wikitable" | ||
| + | !genome DNA||KOD plus||10x buffer||dNTP||MgSO4||Prefix-PkaiBC_fwd primer ver.3||PkaiBC-suffix_rev primer ver.3||MilliQ||total | ||
| + | |- | ||
| + | |RpaB||0.5||2.5||2.5||1.5||0.75||0.75||16.5||25 | ||
| + | |} | ||
| + | |||
| + | |||
{| class="wikitable" | {| class="wikitable" | ||
!PreDenature||Denature||Annealing||Extension||cycle | !PreDenature||Denature||Annealing||Extension||cycle | ||
Latest revision as of 20:43, 27 September 2013
Contents |
Sep 13
Miniprep
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 9/12 Fusion6 antisense | 245.9 | 1.88 | 1.41 |
| 9/12 Plux-3 | 115.5 | 1.58 | 1.32 |
| 9/12 Plux-4 | 125.3 | 1.93 | 1.44 |
| 9/12 Fusion3n2 attenuator-1 | 231.9 | 1.81 | 1.31 |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
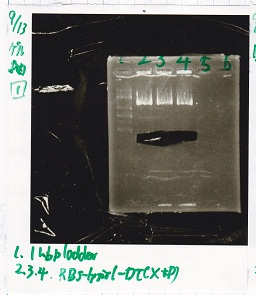
| 1 | 1kb ladder | -- |
| 2 | RBS-lysis1-DT | XbaI&PstI |
| 3 | RBS-lysis1-DT | XbaI&PstI |
| 4 | RBS-lysis1-DT | XbaI&PstI |
| Lane | DNA | Enzyme |
|---|---|---|
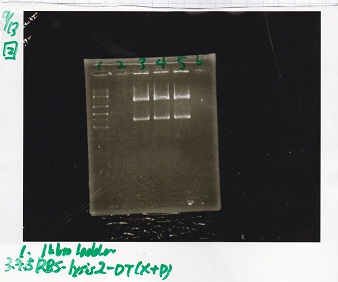
| 1 | 1kb ladder | -- |
| 3 | RBS-lysis2-DT | XbaI&PstI |
| 4 | RBS-lysis2-DT | XbaI&PstI |
| 5 | RBS-lysis2-DT | XbaI&PstI |
| Lane | DNA | Enzyme |
|---|---|---|
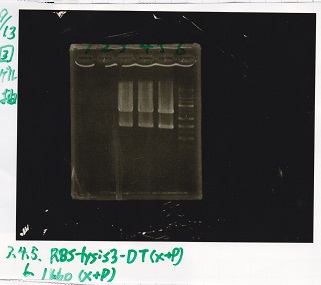
| 3 | RBS-lysis3-DT | XbaI&PstI |
| 4 | RBS-lysis3-DT | XbaI&PstI |
| 5 | RBS-lysis3-DT | XbaI&PstI |
| 6 | 1kb ladder | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 2 | 1kb ladder | -- |
| 3 | pSB1C3 | EcoRI&SpeI |
| 4 | pSB1C3 | EcoRI&SpeI |
| 5 | pSB1C3 | EcoRI&SpeI |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kb ladder | -- |
| 2 | pSB1C3 | XbaI&PstI |
| 3 | pSB1C3 | XbaI&PstI |
| 4 | pSB1C3 | XbaI&PstI |
Restriction Enzyme Digestion
| 9/13 Plac | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cuts(PstI) | 16 | 0µL | 0µL | 0µL | 1µL | 3µL | 3µL | 7µL | 30µL |
| NC | 0.8 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.2µL | 10µL |
| 9/10 Pcon attenuator | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cut(PstI) | 10 | 0µL | 0µL | 0µL | 1µL | 3µL | 3µL | 13µL | 30µL |
| NC | 0.5 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.5µL | 10µL |
| 9/12 Plux | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cuts(PstI) | 12 | 0µL | 0µL | 0µL | 1µL | 3µL | 3µL | 11µL | 30µL |
| NC | 0.6 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.4µL | 10µL |
| 9/12 DT-E | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cut(XbaI) | 23.7 | 0µL | 0µL | 1µL | 0µL | 3µL | 3µL | 0µL | 30.7µL |
| NC | 1.2 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 6.8µL | 10µL |
| 9/12 Pcon-RBS-luxR-bTE | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cuts(XbaI) | 22.4 | 0µL | 0µL | 1µL | 0µL | 3µL | 3µL | 0.6µL | 30µL |
| NC | 2.2 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 5.8µL | 10µL |
| 9/12 Pcon-P | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cut(SpeI) | 21 | 0µL | 1µL | 0µL | 0µL | 3µL | 3µL | 12.0µL | 30µL |
| NC | 2.1 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 5.7µL | 10µL |
M9 (EDTA)
1mM EDTA
| volume | 10ml |
|---|---|
| Na2HPO4 | 300mg |
| KH2PO4 | 150mg |
| NaCl | 25mg |
| NH4Cl | 50mg |
| Fe(3)-EDTA | 21mg |
| volume | 10ml |
|---|---|
| Na2HPO4 | 300mg |
| KH2PO4 | 150mg |
| NaCl | 25mg |
| NH4Cl | 50mg |
| Fe(3)-EDTA | 105mg |
</div>
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kb ladder | -- |
| 3 | Pcon-P | SpeI |
| 4 | Pcon-P | SpeI |
| 5 | Pcon-P | SpeI |
Liquid Culture
| Sample | medium |
|---|---|
| 9/11 aptamer12_1M(pSB1C3)-1 | -- |
| 9/11 aptamer 12_P1 | -- |
| 9/11 Fusaion1 antisense-4 | -- |
| 9/12 pSB4K5-1 | -- |
| 9/12 pSB4K5-2 | -- |
Restriction Enzyme Digestion
| 9/13 Plac-P | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cuts(SpeI) | 23.5 | 0µL | 0.5µL | 0µL | 0µL | 3µL | 3µL | 0µL | 30µL |
| NC | 8 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 0µL | 10µL |
| 9/13 Pcon-pT181 attenuator-P | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cut(SpeI) | 23.5 | 0µL | 0.5µL | 0µL | 0µL | 3µL | 3µL | 0µL | 30µL |
| NC | 02.3 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 5.7µL | 10µL |
| 9/13 Plux-P | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 1cuts(SpeI) | 23.8 | 0µL | 0.2µL | 0µL | 1µL | 3µL | 3µL | 0µL | 30µL |
| NC | 8 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 0µL | 10µL |
| 9/5 RBS-lacZα-DT | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cut(XbaI&PstI) | 10.8 | 0µL | 0µL | 1µL | 1µL | 3µL | 3µL | 11.2µL | 30µL |
| NC | 0.5 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.5µL | 10µL |
Electrophoresis
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/13 Plac (SpeI&PstI) | 8.5µL | 9/13 pT181 antisense (XbaI&PstI) | 2.1 µL | 5.3 µL |
| experiment | 9/13 Plac (SpeI&PstI) | 8.5 µL | 9/10 aptamer 12_1R-DT (XbaI&PstI) | 1.8 µL | 5.15µL |
| experiment | 9/13 Plac (SpeI&PstI) | 8.5 µL | 9/2 pT181 attenuator(XbaI&PstI) | 2.3 µL | 10.4 µL |
| experiment | 9/13 Plux (SpeI&PstI) | 11.4 µL | 9/13 RBS-lysis1-DT (XbaI&PstI) | 5.0 µL | 8.2 µL |
| experiment | 9/13 Plux (SpeI&PstI) | 11.4 µL | 9/13 RBS-lysis2-DT (XbaI&PstI) | 7.0 µL | 9.2 µL |
| experiment | 9/13 Plux (SpeI&PstI) | 7.2 µL | 9/13 RBS-lysis3-DT (XbaI&PstI) | 6.0 µL | 6.6 µL |
| experiment | 9/13 Pcon-RBS-luxR-DT (EcoRI&XbaI) | 2.0 µL | 9/4 Plux-RBS-GFP-DT (EcoRI&SpeI) | 5.2 µL | 3.6 µL |
| experiment | 9/13 Pcon-RBS-luxR-DT (EcoRI&XbaI) | 2.0 µL | 9/10 aptamer 12_1R-DT (XbaI&PstI) | 1.6 µL | 1.8 µL |
| experiment | 9/12 Ptet-pT181 antisense (--) | 1.0 µL | 9/10 spinach-DT (--) | 2.4 µL | 1.7 µL |
PCR
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/7 tetR aptamer 12_1R(pSB1C3) | 0.5 | 6.05 | 1.75 | 1.7 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/6 pT181 antisense-1(pSB1C3) | 0.5 | 6.05 | 1.75 | 1.7 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/6 pT181 attenuator (pSB1C3) | 0.5 | 3.85 | 1.75 | 3.9 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 8/27 Pλ-RBS-luxI-DT-1 | 0.5 | 2.55 | 1.75 | 5.2 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 8/31 Pbad/araC-RBS-RFP-2 | 0.5 | 3.75 | 1.75 | 4 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/13 Plac(pSB1A2) | 0.5 | 4.2 | 1.75 | 3.5 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 8/20 J23100-RBS-lacZα-DT | 0.5 | 6.35 | 1.75 | 1.4 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/7 Ptet-RBS-lacZα-DT | 0.5 | 6.05 | 1.75 | 1.7 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/8 J23100-spinach-DT | 0.5 | 5.75 | 1.75 | 2 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/11 RBS-lysis1-DT | 0.5 | 6.05 | 1.75 | 1.7 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/12 RBS-lysis2-DT | 0.5 | 6.75 | 1.75 | 1.0 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/12 RBS-lysis3-DT | 0.5 | 6.75 | 1.75 | 1.0 | 0.5 | 0 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/8 J23100-spinach-DT | 0.5 | 6.35 | 1.75 | 1.4 | 0 | 0.5 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 8/27 Pλ-RBS-luxI-DT-1 | 0.5 | 2.55 | 1.75 | 5.2 | 0 | 0.5 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/12 RBS-lysis2-DT | 0.5 | 6.75 | 1.75 | 1.0 | 0 | 0.5 | 10.5 |
| genome DNA | Big Dye | MilliQ | 5x buffer | temp(400mg) | SasA_fwd primer | SasA_rev primer | total |
|---|---|---|---|---|---|---|---|
| 9/11 RBS-lysis1-DT | 0.5 | 6.05 | 1.75 | 1.7 | 0 | 0.5 | 10.5 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 96°C | 96°C | 50°C | 68°C | -- |
| 2m | 10s | 5s | 4m | 30 |
PCR
| genome DNA | KOD plus | 10x buffer | dNTP | MgSO4 | Prefix-RBS-RpaB_fwd primer | RpaB-GG4-suffix_rev primer | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| RpaB | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 16.5 | 25 |
| genome DNA | KOD plus | 10x buffer | dNTP | MgSO4 | Prefix-PkaiBC_fwd primer ver.3 | PkaiBC-suffix_rev primer ver.3 | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| RpaB | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 16.5 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 98°C | 58°C | 68°C | -- |
| 2m | 10s | 30s | 30s | 30 |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
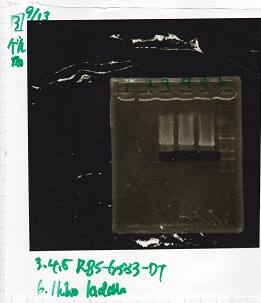
| 1 | 100bp ladder | -- |
| 3 | RBS-lacZα-DT | XbaI&PstI |
| 4 | RBS-lacZα-DT | XbaI&PstI |
| 5 | RBS-lacZα-DT | XbaI&PstI |
| 6 | RBS-lacZα-DT | XbaI&PstI |
Transformation
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| 9/13 Plac+pT181 antisense | 2µL | 20µL | 22µL | Amp |
| 9/13 Plac+apt12_1R | 2µL | 20µL | 22µL | Amp |
| 9/13 Plac+pT181 attenuator | 2µL | 20µL | 22µL | Amp |
| 9/13 Plux+pT181 antisense | 2µL | 20µL | 22µL | CP |
| 9/13 Plux+RBS-lysis2-DT | 2µL | 20µL | 22µL | CP |
| 9/13 Plux+RBS-lysis2-DT | 2µL | 20µL | 22µL | CP |
| 9/13 Pcon-RBS-lacR-DT+Plux-RBS-GFP-DT | 2µL | 20µL | 22µL | Amp |
| 9/13 Pcon-pT181 antisense+apt12_1R-DT | 2µL | 20µL | 22µL | Amp |
| 9/13 Ptet-pT181antisense+spinach-DT | 2µL | 20µL | 22µL | CP |
| Pλ-luxI-1 | 2µL | 20µL | 22µL | Amp |
 "
"