Team:UChicago/Plan
From 2013.igem.org
(→Test keratinase expression and activity in E. coli BL21-De3 kerA transformants) |
(→Assembling plasmid for expression in B. subtilis) |
||
| (87 intermediate revisions not shown) | |||
| Line 26: | Line 26: | ||
<!-- insert your text between these two 'html' tags --> | <!-- insert your text between these two 'html' tags --> | ||
| - | + | = '''Summer Plan''' = | |
| + | |||
| + | |||
== Constructing the kerA BioBricks == | == Constructing the kerA BioBricks == | ||
| - | + | <p>As we approached synthesizing our construct, we realized that it artificial synthesis the kerA gene was cheaper than obtaining the <i>B. subtilis</i> PWD-1 strain and isolating the gene from there. Furthermore, the natural kerA sequence contained cut sites that were incompatible with 3A assembly (EcoRI and a PstI). Therefore, we opted use the Gibson assembly method with geneblocks ordered from IDT to synthesize the kerA gene to our liking. Once synthesized, we amplified with PCR and checked the DNA fragment's length using gel electrophoresis to confirm we had the right DNA.</p> | |
| - | Furthermore, the kerA sequence | + | |
| - | + | ||
| - | + | <p>Once we had sufficient kerA DNA to work with, we decided it would be best to insert the BioBrick into an Amp resistant backbone. To do so, we digested the PCR product and the backbone with EcoRI and PstI. We set up a ligation and transformed the plasmid into competent <i>E. coli</i> DH5-alpha. Unfortunately, we were unable to get any transformants. Prior to the transformation with the plasmid containing kerA, we tested the transformation efficiency of our competent cells so we discarded the possibility that we were performing the transformations incorrectly. Therefore, we thought that maybe the problem was with the assembly, PCR, digestion, or ligation of kerA. We tried using different conditions (temperatures, incubation time, and different enzymes) to eliminate our sources of error, but we were unable to get any transformants.</p> | |
| - | |||
| - | + | <p>We were contemplating making several variants of our construct depending on how our needs changed as the project progressed. For example, the inclusion of the secretion signal peptide depended on which compartment of the cell was best suited to the proper folding of keratinase. Another example would be the inclusion of a His-tag for detection & purification purposes—which could later be removed in the final product. Some of the kerA variants we had planned for are pictured below. The DNA sequences for the 4 variants can also be found <i>[[Team:UChicago/Parts|here]]</i>.</p> | |
| - | + | ||
| - | [[File: | + | [[File:Keratinase gene diagrams.jpg|center|700px]] |
| - | + | ||
| - | [ | + | |
| + | <html><div class="dividerNotebook"><p><a href="#top">[top]</a></p></div></html> | ||
== Constructing the pUB110 backbone == | == Constructing the pUB110 backbone == | ||
| - | In order to express kerA in B. subtilis in high amounts, we need to use a high copy number plasmid. Since high copy number plasmids for B. subtilis are not available | + | <p>In order to express kerA in <i>B. subtilis</i> in high amounts, we need to use a high copy number plasmid. Since high copy number plasmids for <i>B. subtilis</i> are not available from the iGEM HQ, we decided to design our own. We derived our design from pUB110 because it has been widely used in <i>B. subtilis</i></p> |
| - | [[File:pub110 | + | [[File:pub110 original-1.jpg]] |
| + | [[File:pub110 modified-1.jpg]] | ||
| + | Adapted from: | ||
| + | Boe, L. A. R. S., Gros, M. F., te Riele, H. E. I. N., Ehrlich, S. D., & Gruss, A. (1989). [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC210059/ Replication origins of single-stranded-DNA plasmid pUB110.] Journal of bacteriology, 171(6), 3366-3372. | ||
| - | |||
| - | |||
| + | The original pUB110 plasmid is pictured above on the left, while our proposed BioBrick version is modeled on the right. Because the pUB110 vector includes EcoRI and XbaI sites, we had to find a way to remove them before we turned the vector into a BioBrick backbone. We chose to digest pUB110 with NdeI and AflII, which would remove the Pre gene along with the EcoRI and XbaI sites from the vector. Luckily, this gene is not essential for replication in <i>B. subtilis</i>. We then ran the digest in a gel and did a gel extraction to isolate the part of the vector that we needed. | ||
| - | |||
In order to add in the BioBrick prefix and suffix to the vector, we designed a linker that was synthesized as two separate oligos. The DNA sequence of the two oligos are as follows: | In order to add in the BioBrick prefix and suffix to the vector, we designed a linker that was synthesized as two separate oligos. The DNA sequence of the two oligos are as follows: | ||
| - | 5' attcgtcttaaggaattcgcggccgcttctagagtactagtagcggccgctgcagcatatgtcatac 3' | + | <pre>5' attcgtcttaaggaattcgcggccgcttctagagtactagtagcggccgctgcagcatatgtcatac 3' |
| - | 5' gtatgacatatgctgcagcggccgctactagtactctagaagcggccgcgaattccttaagacgaat 3' | + | 5' gtatgacatatgctgcagcggccgctactagtactctagaagcggccgcgaattccttaagacgaat 3'</pre> |
| - | |||
| - | |||
| - | + | We annealed the two oligos in duplex buffer (recipe from IDT) to use the duplex as a linker. Once the oligos were annealed, the linker was digested with NdeI and AflII. We then set up an overnight ligation of the digested, gel purified pUB110 and digested pUB110 linker, which would ideally lead to the formation of a backbone compatible with BioBricks. Unfortunately, we ran out of time and we were unable to get any transformants with our modified vector. To make sure that we weren't doing anything wrong with the transformation procedure of <i>B. subtilis</i>, we tried to transform the original pUB110. The transformation was successful, which led us to think that the problem may be with our modified vector. The failure of transformations could have been attributed to incomplete digestions, too little DNA retrieved from the gel extraction, or incomplete ligation. We attempted to address these potential problems by using new AflII and NdeI restriction enzymes with varying incubation times, different kits for gel extractions, and overnight ligations. Despite all our attempts, we were unable to transform <i>B. subtilis</i> with our modified backbone. | |
| - | |||
| - | + | If our transformation of the modified pUB110 vector worked, we would have gone on to pick transformants, grow them overnight to amplify the quantity of backbone DNA, and then do a miniprep to isolate the modified pUB110 backbone. We would then send the vector for sequencing. Once we verified that the backbone sequence were correct, we would have tested expression of the backbone. Using the NEB assembly kit, we would have digested an upstream part or promoter, a RFP BioBrick, and the modified pUB110 backbone. Once the construct were assembled, we would have transformed it into <i>B. subtilis</i> and checked that it worked by using it to express RFP. | |
| - | |||
| - | + | <html><div class="dividerNotebook"><p><a href="#top">[top]</a></p></div></html> | |
| - | + | == Assembling plasmid for expression in <i>B. subtilis</i> == | |
| - | + | In order to express kerA in <i>B. subtilis</i>, we would have used the NEB assembly kit for the following parts: | |
| - | |||
| - | + | :*Upstream part: constitutive promoter Pveg +RBS spoVG + SacB ([http://parts.igem.org/Part:BBa_K541501 BBa_K541501]) found in a Cam resistant vector. | |
| + | :*Downstream part: kerA variants without the signal peptide in the Amp resistant vector. | ||
| + | :*Destination plasmid: the modified pUB110 with Kan resistance. | ||
| + | [[File:NEB.jpg|center|frame|Schematic of the BioBrick® assembly method. Source: New England Bio Labs]] | ||
| - | == Testing KerA expression and activity in B. subtilis == | + | |
| + | [[File:assembly of our parts 1.jpg|center|frame|Assembly of our parts]] | ||
| + | |||
| + | Note: In order to test expression of kerA with its native signal peptide, we would have instead used the constitutive promoter Pveg + RBS spoVG ([http://parts.igem.org/Part:BBa_K143053 BBa_K143053]) as the upstream part. | ||
| + | |||
| + | |||
| + | <html><div class="dividerNotebook"><p><a href="#top">[top]</a></p></div></html> | ||
| + | |||
| + | == Assembling plasmid for expression in <i>E. coli</i> BL21-De3 == | ||
| + | |||
| + | To test expression in <i>E. coli</i>, we would have used the constitutive expression cassette ([http://parts.igem.org/Part:BBa_K314100 BBa_K314100]) as the upstream part, the kerA variants without the signal peptide, and a high copy number vector with Kan resistance as a destination plasmid. If kerA expression in <i>E. coli</i> were toxic, we would have instead used the lac induced expression cassette ([http://partsregistry.org/Part:BBa_K314103 BBa_K314103]) as the upstream part. | ||
| + | |||
| + | |||
| + | <html><div class="dividerNotebook"><p><a href="#top">[top]</a></p></div></html> | ||
| + | |||
| + | == Testing KerA expression and activity in <i>B. subtilis</i> == | ||
| - | + | Ideally, we would have gone on to grow our <i>B. subtilis</i> kerA transformants on keratin plates (made with LB agar + keratin derived from feathers) to determine secretion and activity of KerA protein. As a negative control for non-specific protease activity we would have instead used non-transformed <i>B. subtilis</i>. As a positive control, we would have also gotten B. licheniformis PWD-1 and grown it on the keratin plates. We would have used the clearing of keratin from the plate as a sign of kerA activity and secretion. | |
| - | [[File:keratin palte.jpg]] | + | [[File:keratin palte 1.jpg]] |
Source: | Source: | ||
| - | Wawrzkiewicz, K., Wolski, T., & Łobarzewski, J. (1991). Screening the keratinolytic activity of dermatophytes in vitro. Mycopathologia, 114(1), 1-8. | + | Wawrzkiewicz, K., Wolski, T., & Łobarzewski, J. (1991). [http://link.springer.com/article/10.1007%2FBF00436684 Screening the keratinolytic activity of dermatophytes in vitro.] Mycopathologia, 114(1), 1-8. |
| - | To be completely certain that kerA (his tagged) is being expressed in B. subtilis, we would have done a western blot on the cell lysis as well as on the liquid medium in which the transformants were grown. The western blot of the medium would indicate whether there was | + | To be completely certain that kerA (his tagged) is being expressed in <i>B. subtilis</i>, we would have also done a western blot on the cell lysis as well as on the liquid medium in which the transformants were grown. The western blot of the medium would indicate whether there was KerA secretion. |
| - | |||
| - | + | In order to have a more quantitative test of KerA activity, we would have used a keratin azure assay, which demonstrates keratinase activity based on a colorimetric reading. In this assay, keratin azure is used, which is a keratin protein that is conjugated to a specific molecule such that when it is digested by a proteinase, it breaks down into a colored reaction product. The concentration of this product, and thus the activity of the proteinase, can then be deduced by measuring the change in color using a spectrophotometer. | |
| - | |||
| - | == | + | <html><div class="dividerNotebook"><p><a href="#top">[top]</a></p></div></html> |
| + | == Testing KerA expression and activity in <i>E. coli</i> BL21-De3 == | ||
| - | + | In <i>E. coli</i> transformants, we would have detected keratinase (his tagged) expression in the cell lysate by western blot. If expression of his tagged KerA is detected, we would have proceeded to do a keratin azure assay on the cell lysate. | |
| - | <html> | + | <html><div class="dividerNotebook"><p><a href="#top">[top]</a></p></div></html> |
| + | == Improve KerA activity by mutagenesis == | ||
| + | |||
| + | Our ideal enzymatic degradation method was to grow <i>B. subtilis</i> directly from the feathers, so that feather waste could be continually added. To do this, KerA's activity and its secretion should occur at a similar temperature. However, optimal KerA secretion occurs between 25 to 37°C, while its highest activity is observed at 50°C. In the future, we would like to use error prone PCR to introduce mutations in kerA as a way to isolate kerA variants with increased activity at lower temperatures. We would then transform the mutated kerA gene into <i>B. subtilis</i>. The KerA mutants would then be tested using a keratin azure assay at temperatures ranging from 25 to 37°C to quantitatively determine keratinase activity. | ||
| + | |||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | <div id="The_Advisors" class="divider"><p><a href="#top">[top]</a></p></div> | ||
| + | <div class="button wrapperContent"> | ||
| + | <h3><a href="/Team:UChicago/Sponsor_Us">Sponsor Us for Next Year's Competition!</a></h3> | ||
</div> | </div> | ||
| - | + | <div id="" class="subNav"> | |
| + | <ul> | ||
| + | <li class=""> | ||
| + | <a href="/Team:UChicago/Project">Return to Project Overview</a> | ||
| + | </li> | ||
| + | </ul> | ||
| + | </div> | ||
| - | |||
| + | </html> | ||
{{:Team:UChicago/Templates/Standard_footer}} | {{:Team:UChicago/Templates/Standard_footer}} | ||
Latest revision as of 03:12, 26 October 2013
Summer Plan
Constructing the kerA BioBricks
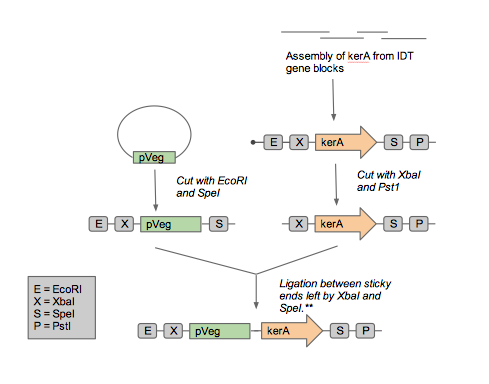
As we approached synthesizing our construct, we realized that it artificial synthesis the kerA gene was cheaper than obtaining the B. subtilis PWD-1 strain and isolating the gene from there. Furthermore, the natural kerA sequence contained cut sites that were incompatible with 3A assembly (EcoRI and a PstI). Therefore, we opted use the Gibson assembly method with geneblocks ordered from IDT to synthesize the kerA gene to our liking. Once synthesized, we amplified with PCR and checked the DNA fragment's length using gel electrophoresis to confirm we had the right DNA.
Once we had sufficient kerA DNA to work with, we decided it would be best to insert the BioBrick into an Amp resistant backbone. To do so, we digested the PCR product and the backbone with EcoRI and PstI. We set up a ligation and transformed the plasmid into competent E. coli DH5-alpha. Unfortunately, we were unable to get any transformants. Prior to the transformation with the plasmid containing kerA, we tested the transformation efficiency of our competent cells so we discarded the possibility that we were performing the transformations incorrectly. Therefore, we thought that maybe the problem was with the assembly, PCR, digestion, or ligation of kerA. We tried using different conditions (temperatures, incubation time, and different enzymes) to eliminate our sources of error, but we were unable to get any transformants.
We were contemplating making several variants of our construct depending on how our needs changed as the project progressed. For example, the inclusion of the secretion signal peptide depended on which compartment of the cell was best suited to the proper folding of keratinase. Another example would be the inclusion of a His-tag for detection & purification purposes—which could later be removed in the final product. Some of the kerA variants we had planned for are pictured below. The DNA sequences for the 4 variants can also be found here.
| Error creating thumbnail: /websites/2013.igem.org/wiki/bin/ulimit4.sh: line 4: convert: command not found |
Constructing the pUB110 backbone
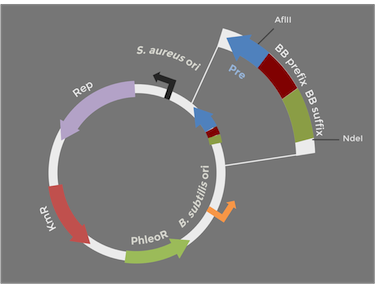
In order to express kerA in B. subtilis in high amounts, we need to use a high copy number plasmid. Since high copy number plasmids for B. subtilis are not available from the iGEM HQ, we decided to design our own. We derived our design from pUB110 because it has been widely used in B. subtilis
Adapted from:
Boe, L. A. R. S., Gros, M. F., te Riele, H. E. I. N., Ehrlich, S. D., & Gruss, A. (1989). [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC210059/ Replication origins of single-stranded-DNA plasmid pUB110.] Journal of bacteriology, 171(6), 3366-3372.
The original pUB110 plasmid is pictured above on the left, while our proposed BioBrick version is modeled on the right. Because the pUB110 vector includes EcoRI and XbaI sites, we had to find a way to remove them before we turned the vector into a BioBrick backbone. We chose to digest pUB110 with NdeI and AflII, which would remove the Pre gene along with the EcoRI and XbaI sites from the vector. Luckily, this gene is not essential for replication in B. subtilis. We then ran the digest in a gel and did a gel extraction to isolate the part of the vector that we needed.
In order to add in the BioBrick prefix and suffix to the vector, we designed a linker that was synthesized as two separate oligos. The DNA sequence of the two oligos are as follows:
5' attcgtcttaaggaattcgcggccgcttctagagtactagtagcggccgctgcagcatatgtcatac 3' 5' gtatgacatatgctgcagcggccgctactagtactctagaagcggccgcgaattccttaagacgaat 3'
We annealed the two oligos in duplex buffer (recipe from IDT) to use the duplex as a linker. Once the oligos were annealed, the linker was digested with NdeI and AflII. We then set up an overnight ligation of the digested, gel purified pUB110 and digested pUB110 linker, which would ideally lead to the formation of a backbone compatible with BioBricks. Unfortunately, we ran out of time and we were unable to get any transformants with our modified vector. To make sure that we weren't doing anything wrong with the transformation procedure of B. subtilis, we tried to transform the original pUB110. The transformation was successful, which led us to think that the problem may be with our modified vector. The failure of transformations could have been attributed to incomplete digestions, too little DNA retrieved from the gel extraction, or incomplete ligation. We attempted to address these potential problems by using new AflII and NdeI restriction enzymes with varying incubation times, different kits for gel extractions, and overnight ligations. Despite all our attempts, we were unable to transform B. subtilis with our modified backbone.
If our transformation of the modified pUB110 vector worked, we would have gone on to pick transformants, grow them overnight to amplify the quantity of backbone DNA, and then do a miniprep to isolate the modified pUB110 backbone. We would then send the vector for sequencing. Once we verified that the backbone sequence were correct, we would have tested expression of the backbone. Using the NEB assembly kit, we would have digested an upstream part or promoter, a RFP BioBrick, and the modified pUB110 backbone. Once the construct were assembled, we would have transformed it into B. subtilis and checked that it worked by using it to express RFP.
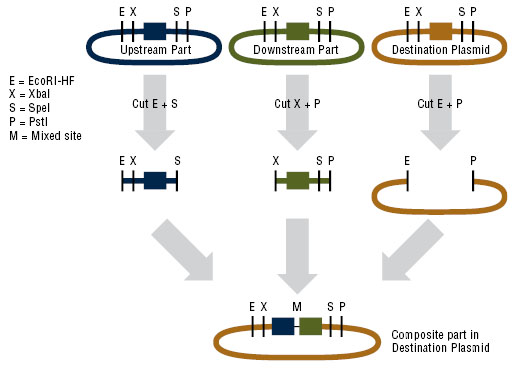
Assembling plasmid for expression in B. subtilis
In order to express kerA in B. subtilis, we would have used the NEB assembly kit for the following parts:
- Upstream part: constitutive promoter Pveg +RBS spoVG + SacB ([http://parts.igem.org/Part:BBa_K541501 BBa_K541501]) found in a Cam resistant vector.
- Downstream part: kerA variants without the signal peptide in the Amp resistant vector.
- Destination plasmid: the modified pUB110 with Kan resistance.
Note: In order to test expression of kerA with its native signal peptide, we would have instead used the constitutive promoter Pveg + RBS spoVG ([http://parts.igem.org/Part:BBa_K143053 BBa_K143053]) as the upstream part.
Assembling plasmid for expression in E. coli BL21-De3
To test expression in E. coli, we would have used the constitutive expression cassette ([http://parts.igem.org/Part:BBa_K314100 BBa_K314100]) as the upstream part, the kerA variants without the signal peptide, and a high copy number vector with Kan resistance as a destination plasmid. If kerA expression in E. coli were toxic, we would have instead used the lac induced expression cassette ([http://partsregistry.org/Part:BBa_K314103 BBa_K314103]) as the upstream part.
Testing KerA expression and activity in B. subtilis
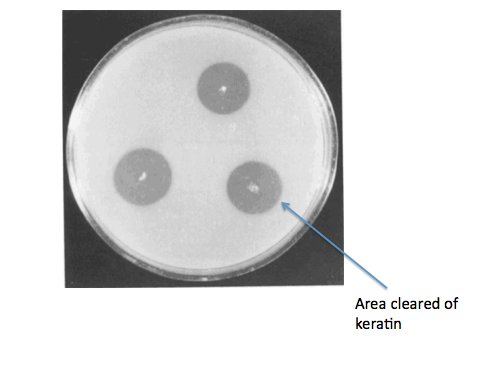
Ideally, we would have gone on to grow our B. subtilis kerA transformants on keratin plates (made with LB agar + keratin derived from feathers) to determine secretion and activity of KerA protein. As a negative control for non-specific protease activity we would have instead used non-transformed B. subtilis. As a positive control, we would have also gotten B. licheniformis PWD-1 and grown it on the keratin plates. We would have used the clearing of keratin from the plate as a sign of kerA activity and secretion.
Source:
Wawrzkiewicz, K., Wolski, T., & Łobarzewski, J. (1991). [http://link.springer.com/article/10.1007%2FBF00436684 Screening the keratinolytic activity of dermatophytes in vitro.] Mycopathologia, 114(1), 1-8.
To be completely certain that kerA (his tagged) is being expressed in B. subtilis, we would have also done a western blot on the cell lysis as well as on the liquid medium in which the transformants were grown. The western blot of the medium would indicate whether there was KerA secretion.
In order to have a more quantitative test of KerA activity, we would have used a keratin azure assay, which demonstrates keratinase activity based on a colorimetric reading. In this assay, keratin azure is used, which is a keratin protein that is conjugated to a specific molecule such that when it is digested by a proteinase, it breaks down into a colored reaction product. The concentration of this product, and thus the activity of the proteinase, can then be deduced by measuring the change in color using a spectrophotometer.
Testing KerA expression and activity in E. coli BL21-De3
In E. coli transformants, we would have detected keratinase (his tagged) expression in the cell lysate by western blot. If expression of his tagged KerA is detected, we would have proceeded to do a keratin azure assay on the cell lysate.
Improve KerA activity by mutagenesis
Our ideal enzymatic degradation method was to grow B. subtilis directly from the feathers, so that feather waste could be continually added. To do this, KerA's activity and its secretion should occur at a similar temperature. However, optimal KerA secretion occurs between 25 to 37°C, while its highest activity is observed at 50°C. In the future, we would like to use error prone PCR to introduce mutations in kerA as a way to isolate kerA variants with increased activity at lower temperatures. We would then transform the mutated kerA gene into B. subtilis. The KerA mutants would then be tested using a keratin azure assay at temperatures ranging from 25 to 37°C to quantitatively determine keratinase activity.
 "
"