Team:Bielefeld-Germany/Biosafety/Biosafety Strain
From 2013.igem.org
| (70 intermediate revisions not shown) | |||
| Line 9: | Line 9: | ||
<html> | <html> | ||
<style> | <style> | ||
| - | h1{ | + | h1{margin-top:70px; } |
| - | + | ||
#globalwrapper ul {padding-left:40px; padding-right:40px;} | #globalwrapper ul {padding-left:40px; padding-right:40px;} | ||
| Line 68: | Line 67: | ||
<p align="justify"> | <p align="justify"> | ||
| - | The | + | The basis for our Safety-System is an auxotrophic safety strain combined with a RNase from ''Bacillus amyloliquefaciens'' as a degradation part. In this K12 derived Safety-Strain the constitutive alanine racemase (''alr'') and the catabolic alanine racemase (''dadX'') are deleted. As the alanine racemase catalyzes the reversible isomerization from L-alanine into the enantiomer D-alanine our safety strain is no longer able to synthesis the essential amino acid D-alanine. D-alanine is an essential molecule for the Gram-negative bacterium ''E. coli'', because it is responsible for the cross-linkage of the peptidoglycan layer. Therefore a lack of D-alanine inhibits cell division and leads to cell lysis in growing bacteria. This effect is comparable to bacteriolytic effect known from beta-lactam-antbiotics.<br> |
| - | An important parameter, when expressing toxic gene products, like | + | An important parameter, when expressing toxic gene products, like Rnase Ba (Barnase), is the maintenance of plasmid stability. The strict dependence on D-alanine ensures plasmid stability, when a plasmid containing the coding sequence for the alanine racemase (''alr'') is brought into the Safety-Strain and no D-alanine is supplemented. Therefore only plasmid carrying bacteria are able to grow. This constitutes a double kill-switch system, because not only the Barnase would ensure cell death, but also loss of the alanine racemase (''alr'') encoded on the plasmid.<br> |
| - | + | ||
</p> | </p> | ||
| - | + | <br> | |
==Theory== | ==Theory== | ||
<p align="justify"> | <p align="justify"> | ||
| - | The cell wall is essential for every living bacteria | + | The cell wall is essential for every living bacteria as it confers stability and structure, protects against osmotic pressure and regulates the transport of molecules.<br> |
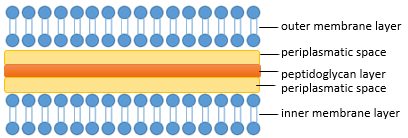
| - | The composition of the cell wall differs between | + | The composition of the cell wall differs between bacteria, a feature commonly used in taxonomy. The most common division is based on the Gram-staining into Gram-negative, for example ''Escherichia coli'' and Gram-positive Bacteria for example ''Bacillus subtillis''. Gram-negative bacteria are characterized by an inner plasma membrane, a thin peptidoglycan layer, periplasmatic spaces and the outer membrane. In contrast Gram-positive bacteria generally lack the outer membrane but have a thicker peptidoglycan layer (see Figure 2 and 3).<br> |
</p> | </p> | ||
<br> | <br> | ||
| - | [[File:IGEM Bielefeld 2013 Biosafety gram- ohnehintergrund 2.png|400px|thumb|center|'''Figure 2:''' | + | [[File:IGEM Bielefeld 2013 Biosafety gram- ohnehintergrund 2.png|400px|thumb|center|'''Figure 2:''' The cell wall of a Gram-negative bacteria is characterized by a thin peptidoglycan-layer and a second membran. ]] |
| - | [[File:IGEM Bielefeld 2013 Biosafety gram+ ohnehintergrund 2.png|400px|thumb|center|'''Figure 3:'''' | + | [[File:IGEM Bielefeld 2013 Biosafety gram+ ohnehintergrund 2 verbessert.png|400px|thumb|center|'''Figure 3:'''' The cell wall of a Gram-positive bacteria instead is characterized by a thick peptidoglycan layer and only one membrane]] |
<br> | <br> | ||
| - | + | <p align="justify"> | |
| - | + | Hence, the peptidoglycan layer is an interesting approach to control bacterial growth. Peptidoglycan itself is a polymer consisting of a linear chain of polysaccharides and short peptides. The polysaccharides component are of alternating residues of beta-(1,4) linked ''N''-acetylglucosamine and ''N''-acetylmuramic acid and they are cross-linked in ''E. coli'' by a tetra-peptide of L-alanine, D-glutamic acid, ''meso''-diaminopimelic acid and finally D-alanine. The cross-linkage is thereby realized by a transpeptide-linkage of ''meso''-diaminopimelic acid and D-alanine.([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#References Cava ''et al.'', 2011])<br> | |
| + | Therefore D-alanine is an essential component of the bacterial peptidoglycan layer. In conclusion a lack of D-alanine would have a bacteriolytic effect, comparable to beta-lactam-antibiotics, because it inhibits the building of the peptidoglycan-linkage. Hence, the bacteria without the ability to obtain D-alanine will lyse during cell division.</p><br> | ||
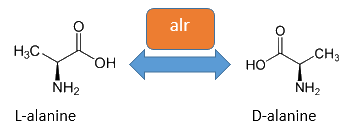
| - | [[Image:IGEM Bielefeld 2013 alr isomerase bearbeitet.png|600px|thumb|center|'''Figure | + | [[Image:IGEM Bielefeld 2013 alr isomerase bearbeitet.png|600px|thumb|center|'''Figure 4:''' The alanine racemase from ''E. coli'' catalyses the reversible reaction from L-alanine to D-alanine. Pyridoxal-5'-phosphate is an essential cofactor for this reaction. ]] |
| - | + | ||
| - | + | ||
| - | + | ||
| + | <p align="justify"> | ||
| + | The accumulation of D-alanine in ''E. coli'' can be catalyzed by an alanine racemase (EC 5.1.1.1). This enzyme enables the reversible reaction from L-alanine into the enantiomer D-alanine. For this reaction the cofactor pyridoxal-5'-phosphate (PLP) is also needed. ''E. coli'' posses two alanine racemases. One, encoded by ''alr'' is constitutively expressed and therefore normally responsible for the accumulation of D-alanine, while the other one encoded by ''dadX'' is under control of the ''dad''-operon and usually used in catabolism ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#References Walsh ''et al.'', 1989]).<br> | ||
| + | The deletion of the constitutive alanine racemase (''alr'') and the catabolic alanine racemase (''dadX'') will lead to a strict dependence of D-alanine, so that the bacteria with these mutations are only able to grow on media with D-alanine supplementation or by complementation of the alanine racemase on a separate plasmid. As shown in [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#Results Figure 5] below the D-alanine-auxotrophic mutant shows a strict dependence on D-alanine.</p><br> | ||
==Genetic Approach== | ==Genetic Approach== | ||
| + | <br> | ||
| + | ==='''Deletion of the alanine racemases'''=== | ||
<p align="justify"> | <p align="justify"> | ||
| - | For the construction of our Biosafety- | + | For the construction of our Biosafety-Chassis we deleted both alanine racemases via homologous recombination with the plasmid pKO4 (a derivative of pKO3 containing the coding sequence of the ''lacZ-alpha'' fragment and a kanamycin resistance marker) to establish a D-alanine-auxotrophic mutant. For the deletion of the alanine racemase we used the following primers.<br> |
| - | By following the protocol for the double-cross over using the heat-sensitive RepA and the sacB | + | The primers alr_Ec_d1 and alr_Ec_d2 will amplify an about 600 bp long homologous fragment upstream of the coding sequence of the alanine racemase, while the primers alr_Ec_d3 and alr_Ec_d4 will yield an about 600 bp long homologous fragment downstream of the corresponding alanine racemase. This two fragments can be ligated by Gibson-Assembly or alternative classically by gene SOEing (Gene splicing by overlap extension, [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#References Horton ''et al.'', 1990]). Afterwards the fuesd fragments flanking the alanine racemase were cloned into the vector pKO4 using again primer-extension (see box below).<br> |
| + | By following the protocol for the double-cross over using the heat-sensitive RepA protein and the ''sacB'' gene from the vector [http://arep.med.harvard.edu/labgc/pko3.html pKO3], the two alanine racemases could be successfully deleted ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#References Link ''et al.'', 1990]).</p> | ||
| - | '''Primer for the complete | + | '''Primer for the complete deletion of the constitutive alanine racemase (''alr'') and the catabolic alanine racemase (''dadX'') in ''E. coli''<br> |
'''Primer: alr_Ec_d1 (39 bp)'''<br> | '''Primer: alr_Ec_d1 (39 bp)'''<br> | ||
5'-'''TAGCTCACTCATTAGGCACCC'''AGCTCGATGACGAAGACT-3'<br> | 5'-'''TAGCTCACTCATTAGGCACCC'''AGCTCGATGACGAAGACT-3'<br> | ||
| Line 121: | Line 123: | ||
<p align="justify"> | <p align="justify"> | ||
| - | To verify the | + | To verify the deletion of the alanine racemases in ''E. coli'', we designed some additional primers, which anneal outside the deletion region so that a successful deletion can be seen by a 1370 bp (''alr'') or a 1312 bp (''dadX'') amplified DNA fragment, while the wild type shows typically a DNA fragment with a length of 2439 bp (''alr'') or 2375 bp (''dadX''). The primers for the control are listed in the box below.</p> |
<br> | <br> | ||
| - | '''Primer for the control of the successful | + | '''Primer for the control of the successful deletion of the constitutive alanine racemase (''alr'') and the catabolic alanine racemase (''dadX'') in ''E. coli''<br> |
'''Primer: alr_Ec_control1 (20 bp)'''<br> | '''Primer: alr_Ec_control1 (20 bp)'''<br> | ||
5'-GCTGGAGATGCCATCAGAAC-3'<br> | 5'-GCTGGAGATGCCATCAGAAC-3'<br> | ||
| Line 134: | Line 136: | ||
5'-CTGGATCAACGCTTCTTTGG-3'<br> | 5'-CTGGATCAACGCTTCTTTGG-3'<br> | ||
| + | <br> | ||
| + | |||
| + | ==='''Deletion of araC'''=== | ||
<p align="justify"> | <p align="justify"> | ||
| - | + | In addition to the deletion of the alanine racemases we deleted the coding sequence for ''araC'' in the Safety-Strain. As the AraC protein is the repressor of the arabinose promoter P<sub>BAD</sub>, this mutation effects only the Biosafety-System araCactive, but is very useful because of two reasons: On the one hand, the repressor AraC works also as an activator, so that there is no basal transcription behind the P<sub>BAD</sub> promoter in the strain ''E. coli'' K-12 Δ''alr'' Δ''dadX'' Δ''araC'' as [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#Results verified]. This allows to transform the Biosafety-System araCative with the barnase as the toxic gene product in the bacteria without triggering cell death. On the other hand the P<sub>BAD</sub> promoter is possible too tightly regulated by the repressor AraC, so that a genomic deletion is necessary for the expression of the toxic gene barnase, when there is still a basal transcription of the ''araC'' gene from the plasmid.</p> | |
<br> | <br> | ||
<p align="justify"> | <p align="justify"> | ||
| - | For the deletion of the repressor | + | For the deletion of the repressor AraC we therefore used the same methods as for the deletion of the alanine racemases in ''E. coli''. The primers araC_d1 and araC_d2 will amplify a 600 bp long homologous fragment upstream of the coding sequence for ''araC'', while the primers araC_d3 and araC_d4 will amplify an about 600 bp long fragment downstream of ''araC''. This two fragments can be ligated by Gibson-Assembly or gene SOEing (Gene splicing overlap Extension, [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#References Horton ''et al.'', 1990]) and then cloned into to gene-targeting vector [http://arep.med.harvard.edu/labgc/pko3.html pKO3]. |
| - | By following the protocol for the double-cross over using the heat-sensitive | + | By following the protocol for the double-cross over by using the heat-sensitive RepA protein and the ''sacB'' from the vector pKO3, the repressor AraC could be successfully deleted ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#References Link ''et al.'', 1990]).</p> |
| - | '''Primer for the complete | + | '''Primer for the complete deletion of the repressor coding gene ''araC'' in ''E. coli''<br> |
'''Primer: araC_d1 (40 bp)'''<br> | '''Primer: araC_d1 (40 bp)'''<br> | ||
5'-'''TAGCTCACTCATTAGGCACCC'''CGGCAGGAATATCGATCGC-3'<br> | 5'-'''TAGCTCACTCATTAGGCACCC'''CGGCAGGAATATCGATCGC-3'<br> | ||
| Line 153: | Line 158: | ||
<p align="justify"> | <p align="justify"> | ||
| - | To verify the deletion of the repressor | + | To verify the deletion of the repressor AraC in ''E. coli'', we designed primers, which anneal outside the deletion region so that a successful deletion can be seen by a 1404 bp amplified DNA fragment, while the wild type shows typically a DNA fragment with a length of 2268 bp. The primers for the control are listed in the box below.</p> |
<br> | <br> | ||
| Line 163: | Line 168: | ||
==Results== | ==Results== | ||
| + | <br> | ||
| + | ==='''Characterization of the D-alanine auxotrophy'''=== | ||
| + | <p align="justify"> | ||
| + | The deletion of the alanine racemases and ''araC'' in ''E. coli'' was not possible in the commonly used strains like JM109, Top10 or KRX, but in the wild type strain K-12. This might be due to the ''recA''1 mutations in these strains, which guarantees a better plasmid maintenance because of a defect in the RecA recombinase. <br> | ||
| + | To avoid a second recombination of the alanine racemase (''alr'') from the plasmid with the genome, the whole coding sequence was deleted in the genome and the characterization of the alanine racemase was performed with the antibiotic chloramphenicol. For the complementation the alanine racemase (''alr'') was brought under the control of the P<sub>''tac''</sub> promoter. The P<sub>''tac''</sub> promoter is a fusion promoter of the -35 region of the ''trp'' promoter and the -10 region the ''lac'' promoter, so that there only slight repression and the expression of the alanine racemase is highly activated ([https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_Strain#References De Boer ''et al.'', 1983]). Therefore an induction with IPTG was not necessary on M9 minimal medium, but surprisingly it was essential on LB-agar.<br> | ||
| + | The deletion of the constitutive alanine racemase (''alr'') and the catabolic alanine racemase (''dadX'') in ''E. coli'' leads to a strict dependence on the amino acid D-alanine, as expected. As shown in Figure 5 below the bacteria with this deletions are not any more able to grow on normal M9 medium without D-alanine supplementation (purple curve), whereas the wild type does (red curve). The auxotrophic Safety-Strain grows only on media with D-alanine (5 mM) supplemented (blue curve) or by a complementation of the alanine racemase via plasmid. Furthermore, it could be demonstrated, that the auxotrophic mutant K-12 ∆''alr'' ∆''dadX'' grows slightly slower, than the wild type K-12. In contrast the bacteria containing the alanine racemase (''alr'') on the plasmid <bbpart>BBa_K1172902</bbpart> does hardly show a disadvantage in the cell division compared to the wild type.</p> | ||
| + | <br> | ||
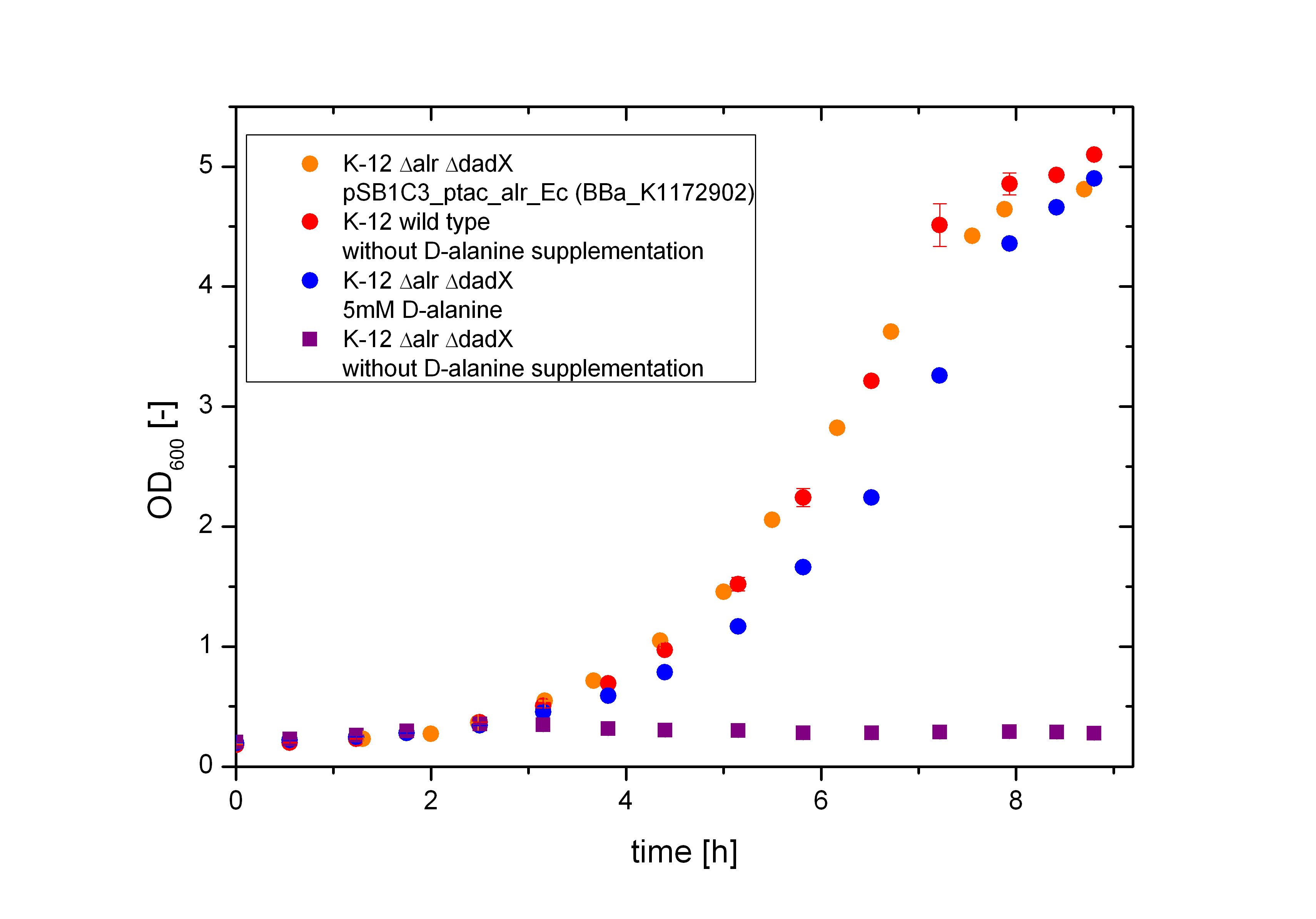
| + | [[File:Team-Bielefeld-Biosafety-Strain-D-Alanine-Complementation.jpg|600px|thumb|center|'''Figure 5:''' Characterization of the D-alanine auxotrophic Biosafety-Strain. The Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' depends strict on the presence of D-alanine in the media or a complementation via plasmid containing an intact alanine racemase.]] | ||
| + | <br> | ||
| - | == | + | ==='''Characterization of the araC-deletion'''=== |
| + | <p align="justify"> | ||
| + | The deletion of the ''araC'' gene was added to the D-alanine auxotrophic Biosafety-Strain resulting in the ''E. coli'' strain K-12 ∆''alr'' ∆''dadX'' ∆''araC''. This strain is, in addition to the D-alanine auxotrophy, not able any more to use the pentose L-arabinose as carbon source. This opens the possibility to maintain a plasmid with a toxic gene behind the P<sub>BAD</sub> promoter without causing cell death. This huge advantage results from the AraC protein functioning as repressor and activator of the arabinose operon ''araBAD'' simultaneously. Lacking this activator, the transcription of the genes behind this promoter is shutdown completely.<br> | ||
| + | To verify if the P<sub>BAD</sub> promoter is really shutdown, the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' ∆''araC'' was cultivated on M9-minimal medium containing only L-arabinose as carbon source. As shown in Figure 6 below without the ''araC'' gene there is not even a slight basal transcription (blue curve). The Biosafety-Strain can not grow on L-arabinose, where as the positive control (red curve) does. In addition, the growth of the wild-type on L-arabinose was compared to D-glucose (black curve). The faster growth of ''E. coli'' on D-glucose in contrast to L-arabinose explains why the ''araBAD'' operon is naturally shut down. Because D-glucose is more efficient to metabolize, a tight regulation of the arabinose catabolism is needed to save the cell resources while the better substrate is present. But in synthetic biology this regulon can be used for different approaches. The Biosafety-System [https://2013.igem.org/Team:Bielefeld-Germany/Biosafety/Biosafety_System_S araCtive] is only one interesting application.</p><br> | ||
| - | + | <br> | |
| - | + | ||
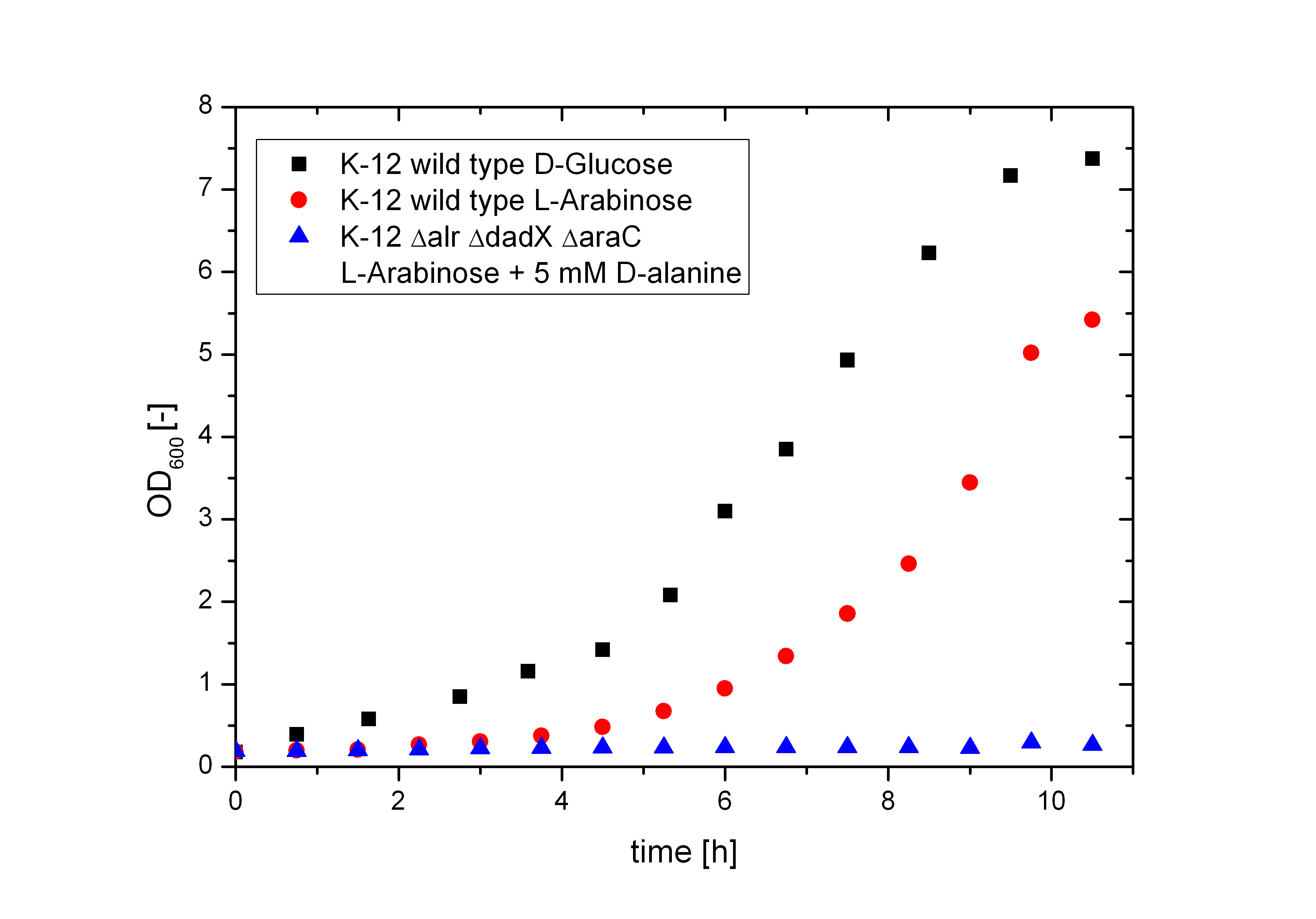
| + | [[File:Team-Bielefeld-Biosafety-Strain-araCharacterization.jpg|600px|thumb|center|'''Figure 6:''' Characterization of the ''araC'' deletion. The AraC protein is essential for the transcription of the P<sub>BAD</sub> promoter, so that the Biosafety-Strain K-12 ∆''alr'' ∆''dadX'' ∆''araC'' can not use L-arabinose as carbon source, while the wildtype K-12 does. To compare the efficiency of L-arabinose metabolism with D-glucose this cultivation is also shown. ]] | ||
| - | < | + | |
| + | ==References== | ||
| + | <p align="justify"> | ||
| + | *Cava F, Lam H, de Pedro MA, Waldor MK (2011) Emerging knowledge of regulatory roles of d-amino acids in bacteria [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3037491/pdf/18_2010_Article_571.pdf|''Cell and Molecular Life Sciences 68: 817 - 831.''] | ||
| + | *De Boer Hermann, Comstock J. Lisa and Vasser Mark (1983) The tac promoter: a functional hybrid derived from the trp and lac promoters. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC393301/pdf/pnas00627-0036.pdf|''Proceedings of the National Academy of Science of the United States of America 80: 21 - 25.''] | ||
| + | *Horton R., Cai Z., Ho S. and Pease L. (1990) Gene splicing by overlap extension: tailor-made genes using the polymerase chain reaction [http://www.ncbi.nlm.nih.gov/pubmed/2357375|''BioTechniques 8: 528 - 535.''] | ||
| + | *Link, A.J., Phillips, D. and Church, G.M. (1997) Methods for generating precise deletions and insertions in the genome of wild-type Escherichia coli: Application to open reading frame characterization. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC179534/pdf/1796228.pdf|''Journal of Bacteriology 179: 6228-6237.''] | ||
| + | *Walsh, Christopher (1989) Enzymes in the D-alanine branch of bacterial cell wall peptidoglycan assembly. [http://www.jbc.org/content/264/5/2393.long|''Journal of biological chemistry 264: 2393 - 2396.''] | ||
| + | |||
| + | </p> | ||
| + | <br><br><br> | ||
</div> | </div> | ||
| Line 180: | Line 207: | ||
| - | <div id=" | + | <div id="asdf"> |
| + | <html> | ||
| + | <div id="nav2" style="width:210px; padding-bottom:5px; padding-left:15px;"> | ||
| + | |||
| + | <div class="navbutton" id="home" style="float:left; padding-left:55px; margin-left:10px; padding-top:0px;"> | ||
| + | <a href="https://2013.igem.org/Team:Bielefeld-Germany" title="Jump to Frontpage"> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/f/f6/Bielefeld-Germany2013-ButtonHome.png" height="15px"> | ||
| + | </a> | ||
| + | </div> | ||
| + | <div class="navbutton" id="top" style="float:left; padding-left:10px; margin-left:10px; padding-top:0px;"> | ||
| + | <a href="#" title="Jump to top"> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/a/ab/Bielefeld-Germany2013-Up_orange_new.png" height="15px"> | ||
| + | </a> | ||
| + | </div> | ||
| + | </div> | ||
| + | </html> | ||
| + | |||
| + | |||
| + | <div id="rightcol" style="width:210px; height:100%; overflow-y:auto; box-shadow:0px 0px 2px 0px grey;" padding:0px 20px;> | ||
__TOC__ | __TOC__ | ||
| - | + | </div> | |
| + | |||
| + | |||
</div> | </div> | ||
</div> | </div> | ||
Latest revision as of 09:44, 26 October 2013
Safety Strain
Overview
The basis for our Safety-System is an auxotrophic safety strain combined with a RNase from Bacillus amyloliquefaciens as a degradation part. In this K12 derived Safety-Strain the constitutive alanine racemase (alr) and the catabolic alanine racemase (dadX) are deleted. As the alanine racemase catalyzes the reversible isomerization from L-alanine into the enantiomer D-alanine our safety strain is no longer able to synthesis the essential amino acid D-alanine. D-alanine is an essential molecule for the Gram-negative bacterium E. coli, because it is responsible for the cross-linkage of the peptidoglycan layer. Therefore a lack of D-alanine inhibits cell division and leads to cell lysis in growing bacteria. This effect is comparable to bacteriolytic effect known from beta-lactam-antbiotics.
An important parameter, when expressing toxic gene products, like Rnase Ba (Barnase), is the maintenance of plasmid stability. The strict dependence on D-alanine ensures plasmid stability, when a plasmid containing the coding sequence for the alanine racemase (alr) is brought into the Safety-Strain and no D-alanine is supplemented. Therefore only plasmid carrying bacteria are able to grow. This constitutes a double kill-switch system, because not only the Barnase would ensure cell death, but also loss of the alanine racemase (alr) encoded on the plasmid.
Theory
The cell wall is essential for every living bacteria as it confers stability and structure, protects against osmotic pressure and regulates the transport of molecules.
The composition of the cell wall differs between bacteria, a feature commonly used in taxonomy. The most common division is based on the Gram-staining into Gram-negative, for example Escherichia coli and Gram-positive Bacteria for example Bacillus subtillis. Gram-negative bacteria are characterized by an inner plasma membrane, a thin peptidoglycan layer, periplasmatic spaces and the outer membrane. In contrast Gram-positive bacteria generally lack the outer membrane but have a thicker peptidoglycan layer (see Figure 2 and 3).
Hence, the peptidoglycan layer is an interesting approach to control bacterial growth. Peptidoglycan itself is a polymer consisting of a linear chain of polysaccharides and short peptides. The polysaccharides component are of alternating residues of beta-(1,4) linked N-acetylglucosamine and N-acetylmuramic acid and they are cross-linked in E. coli by a tetra-peptide of L-alanine, D-glutamic acid, meso-diaminopimelic acid and finally D-alanine. The cross-linkage is thereby realized by a transpeptide-linkage of meso-diaminopimelic acid and D-alanine.(Cava et al., 2011)
Therefore D-alanine is an essential component of the bacterial peptidoglycan layer. In conclusion a lack of D-alanine would have a bacteriolytic effect, comparable to beta-lactam-antibiotics, because it inhibits the building of the peptidoglycan-linkage. Hence, the bacteria without the ability to obtain D-alanine will lyse during cell division.
The accumulation of D-alanine in E. coli can be catalyzed by an alanine racemase (EC 5.1.1.1). This enzyme enables the reversible reaction from L-alanine into the enantiomer D-alanine. For this reaction the cofactor pyridoxal-5'-phosphate (PLP) is also needed. E. coli posses two alanine racemases. One, encoded by alr is constitutively expressed and therefore normally responsible for the accumulation of D-alanine, while the other one encoded by dadX is under control of the dad-operon and usually used in catabolism (Walsh et al., 1989).
The deletion of the constitutive alanine racemase (alr) and the catabolic alanine racemase (dadX) will lead to a strict dependence of D-alanine, so that the bacteria with these mutations are only able to grow on media with D-alanine supplementation or by complementation of the alanine racemase on a separate plasmid. As shown in Figure 5 below the D-alanine-auxotrophic mutant shows a strict dependence on D-alanine.
Genetic Approach
Deletion of the alanine racemases
For the construction of our Biosafety-Chassis we deleted both alanine racemases via homologous recombination with the plasmid pKO4 (a derivative of pKO3 containing the coding sequence of the lacZ-alpha fragment and a kanamycin resistance marker) to establish a D-alanine-auxotrophic mutant. For the deletion of the alanine racemase we used the following primers.
The primers alr_Ec_d1 and alr_Ec_d2 will amplify an about 600 bp long homologous fragment upstream of the coding sequence of the alanine racemase, while the primers alr_Ec_d3 and alr_Ec_d4 will yield an about 600 bp long homologous fragment downstream of the corresponding alanine racemase. This two fragments can be ligated by Gibson-Assembly or alternative classically by gene SOEing (Gene splicing by overlap extension, Horton et al., 1990). Afterwards the fuesd fragments flanking the alanine racemase were cloned into the vector pKO4 using again primer-extension (see box below).
By following the protocol for the double-cross over using the heat-sensitive RepA protein and the sacB gene from the vector [http://arep.med.harvard.edu/labgc/pko3.html pKO3], the two alanine racemases could be successfully deleted (Link et al., 1990).
Primer for the complete deletion of the constitutive alanine racemase (alr) and the catabolic alanine racemase (dadX) in E. coli
Primer: alr_Ec_d1 (39 bp)
5'-TAGCTCACTCATTAGGCACCCAGCTCGATGACGAAGACT-3'
Primer: alr_Ec_d2 (20 bp)
5'-GCCGCTTGCATTTGTGTTCC-3'
Primer: alr_Ec_d3 (40 bp)
5'-GGAACACAAATGCAAGCGGCTTGATTGTCTGTGCCGGATG-3'
Primer: alr_Ec_d4 (40 bp)
5'-GCTTTCTACGTGTTCCGCTTCCGGGAAGTAGCGTTTCAGG-3'
Primer: dadX_Ec_d1 (39 bp)
5'-TAGCTCACTCATTAGGCACCTGAAGTGTGGCGATGAAGT-3'
Primer:dadX_Ec_d2 (20 bp)
5'-GGGTCATCTCGTTTCCTTAG-3'
Primer: dadX_Ec_d3 (40 bp)
5'-CTAAGGAAACGAGATGACCCACTTGTTGTAAGCCGGATCG-3'
Primer: dadX_Ec_d4 (40 bp)
5'-GCTTTCTACGTGTTCCGCTTCGAAGCCAGCGCCAAATATG-3'
To verify the deletion of the alanine racemases in E. coli, we designed some additional primers, which anneal outside the deletion region so that a successful deletion can be seen by a 1370 bp (alr) or a 1312 bp (dadX) amplified DNA fragment, while the wild type shows typically a DNA fragment with a length of 2439 bp (alr) or 2375 bp (dadX). The primers for the control are listed in the box below.
Primer for the control of the successful deletion of the constitutive alanine racemase (alr) and the catabolic alanine racemase (dadX) in E. coli
Primer: alr_Ec_control1 (20 bp)
5'-GCTGGAGATGCCATCAGAAC-3'
Primer: alr_Ec_control2 (20 bp)
5'-CCGGCGAATATTGCTACGTG-3'
Primer: dadX_Ec_control1 (20 bp)
5'-GCTTTAATACGCCCGTTGAC-3'
Primer: dadX_Ec_control2 (20 bp)
5'-CTGGATCAACGCTTCTTTGG-3'
Deletion of araC
In addition to the deletion of the alanine racemases we deleted the coding sequence for araC in the Safety-Strain. As the AraC protein is the repressor of the arabinose promoter PBAD, this mutation effects only the Biosafety-System araCactive, but is very useful because of two reasons: On the one hand, the repressor AraC works also as an activator, so that there is no basal transcription behind the PBAD promoter in the strain E. coli K-12 Δalr ΔdadX ΔaraC as verified. This allows to transform the Biosafety-System araCative with the barnase as the toxic gene product in the bacteria without triggering cell death. On the other hand the PBAD promoter is possible too tightly regulated by the repressor AraC, so that a genomic deletion is necessary for the expression of the toxic gene barnase, when there is still a basal transcription of the araC gene from the plasmid.
For the deletion of the repressor AraC we therefore used the same methods as for the deletion of the alanine racemases in E. coli. The primers araC_d1 and araC_d2 will amplify a 600 bp long homologous fragment upstream of the coding sequence for araC, while the primers araC_d3 and araC_d4 will amplify an about 600 bp long fragment downstream of araC. This two fragments can be ligated by Gibson-Assembly or gene SOEing (Gene splicing overlap Extension, Horton et al., 1990) and then cloned into to gene-targeting vector [http://arep.med.harvard.edu/labgc/pko3.html pKO3]. By following the protocol for the double-cross over by using the heat-sensitive RepA protein and the sacB from the vector pKO3, the repressor AraC could be successfully deleted (Link et al., 1990).
Primer for the complete deletion of the repressor coding gene araC in E. coli
Primer: araC_d1 (40 bp)
5'-TAGCTCACTCATTAGGCACCCCGGCAGGAATATCGATCGC-3'
Primer: araC_d2 (40 bp)
5'-CTTCTCTGAATGGTGGGAGTGTCATAATTGGTAACGAATC-3'
Primer: araC_d3 (40 bp)
5'-GATTCGTTACCAATTATGACACTCCCACCATTCAGAGAAG-3'
Primer: araC_d4 (40 bp)
5'-GCTTTCTACGTGTTCCGCTTAACGCCAATCCCGACCACAG-3'
To verify the deletion of the repressor AraC in E. coli, we designed primers, which anneal outside the deletion region so that a successful deletion can be seen by a 1404 bp amplified DNA fragment, while the wild type shows typically a DNA fragment with a length of 2268 bp. The primers for the control are listed in the box below.
Primer for the control of the successful Deletion of repressor araC in E. coli
Primer: araC_control1 (20 bp)
5'-CTTTACCGCTGCGCCATAAC-3'
Primer: araC_control2 (20 bp)
5'-AACCGCAGTGTGGTCTTTCC-3'
Results
Characterization of the D-alanine auxotrophy
The deletion of the alanine racemases and araC in E. coli was not possible in the commonly used strains like JM109, Top10 or KRX, but in the wild type strain K-12. This might be due to the recA1 mutations in these strains, which guarantees a better plasmid maintenance because of a defect in the RecA recombinase.
To avoid a second recombination of the alanine racemase (alr) from the plasmid with the genome, the whole coding sequence was deleted in the genome and the characterization of the alanine racemase was performed with the antibiotic chloramphenicol. For the complementation the alanine racemase (alr) was brought under the control of the Ptac promoter. The Ptac promoter is a fusion promoter of the -35 region of the trp promoter and the -10 region the lac promoter, so that there only slight repression and the expression of the alanine racemase is highly activated (De Boer et al., 1983). Therefore an induction with IPTG was not necessary on M9 minimal medium, but surprisingly it was essential on LB-agar.
The deletion of the constitutive alanine racemase (alr) and the catabolic alanine racemase (dadX) in E. coli leads to a strict dependence on the amino acid D-alanine, as expected. As shown in Figure 5 below the bacteria with this deletions are not any more able to grow on normal M9 medium without D-alanine supplementation (purple curve), whereas the wild type does (red curve). The auxotrophic Safety-Strain grows only on media with D-alanine (5 mM) supplemented (blue curve) or by a complementation of the alanine racemase via plasmid. Furthermore, it could be demonstrated, that the auxotrophic mutant K-12 ∆alr ∆dadX grows slightly slower, than the wild type K-12. In contrast the bacteria containing the alanine racemase (alr) on the plasmid <bbpart>BBa_K1172902</bbpart> does hardly show a disadvantage in the cell division compared to the wild type.
Characterization of the araC-deletion
The deletion of the araC gene was added to the D-alanine auxotrophic Biosafety-Strain resulting in the E. coli strain K-12 ∆alr ∆dadX ∆araC. This strain is, in addition to the D-alanine auxotrophy, not able any more to use the pentose L-arabinose as carbon source. This opens the possibility to maintain a plasmid with a toxic gene behind the PBAD promoter without causing cell death. This huge advantage results from the AraC protein functioning as repressor and activator of the arabinose operon araBAD simultaneously. Lacking this activator, the transcription of the genes behind this promoter is shutdown completely.
To verify if the PBAD promoter is really shutdown, the Biosafety-Strain K-12 ∆alr ∆dadX ∆araC was cultivated on M9-minimal medium containing only L-arabinose as carbon source. As shown in Figure 6 below without the araC gene there is not even a slight basal transcription (blue curve). The Biosafety-Strain can not grow on L-arabinose, where as the positive control (red curve) does. In addition, the growth of the wild-type on L-arabinose was compared to D-glucose (black curve). The faster growth of E. coli on D-glucose in contrast to L-arabinose explains why the araBAD operon is naturally shut down. Because D-glucose is more efficient to metabolize, a tight regulation of the arabinose catabolism is needed to save the cell resources while the better substrate is present. But in synthetic biology this regulon can be used for different approaches. The Biosafety-System araCtive is only one interesting application.
References
- Cava F, Lam H, de Pedro MA, Waldor MK (2011) Emerging knowledge of regulatory roles of d-amino acids in bacteria [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3037491/pdf/18_2010_Article_571.pdf|Cell and Molecular Life Sciences 68: 817 - 831.]
- De Boer Hermann, Comstock J. Lisa and Vasser Mark (1983) The tac promoter: a functional hybrid derived from the trp and lac promoters. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC393301/pdf/pnas00627-0036.pdf|Proceedings of the National Academy of Science of the United States of America 80: 21 - 25.]
- Horton R., Cai Z., Ho S. and Pease L. (1990) Gene splicing by overlap extension: tailor-made genes using the polymerase chain reaction [http://www.ncbi.nlm.nih.gov/pubmed/2357375|BioTechniques 8: 528 - 535.]
- Link, A.J., Phillips, D. and Church, G.M. (1997) Methods for generating precise deletions and insertions in the genome of wild-type Escherichia coli: Application to open reading frame characterization. [http://www.ncbi.nlm.nih.gov/pmc/articles/PMC179534/pdf/1796228.pdf|Journal of Bacteriology 179: 6228-6237.]
- Walsh, Christopher (1989) Enzymes in the D-alanine branch of bacterial cell wall peptidoglycan assembly. [http://www.jbc.org/content/264/5/2393.long|Journal of biological chemistry 264: 2393 - 2396.]
 "
"