Team:Valencia Biocampus/Project
From 2013.igem.org
| (38 intermediate revisions not shown) | |||
| Line 28: | Line 28: | ||
preloader:true | preloader:true | ||
}); | }); | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
}); | }); | ||
</script> | </script> | ||
| Line 44: | Line 39: | ||
<li><a href="#Results">Results</a></li> | <li><a href="#Results">Results</a></li> | ||
<li><a href="#Parts">Biobricks</a></li> | <li><a href="#Parts">Biobricks</a></li> | ||
| - | + | <li><a href="#Notebook">Notebook</a></li> | |
</ul> | </ul> | ||
| Line 55: | Line 50: | ||
<html> | <html> | ||
<div style="width:540px;margin-right:15px;float:left;"> | <div style="width:540px;margin-right:15px;float:left;"> | ||
| - | Bacteria are essential in biotechnology, but they can hardly move. Nematodes, such as <i>Caenorhabditis elegans</i>, are fast crawling organisms, but they have limited biotechnological applications. By combining the best from both organisms, we present the first artificial synthetic symbiosis with bacteria engineered to ride on worms, which concentrate in hotspots where bacteria perform a desired biotechnological process, such as bioplastic (PHA) production. We have engineered <i>Pseudomonas putida</i> with a whole operon that allows the formation of a biofilm on the worm. Biofilm formation is switched on and off depending on the media, and thus bacteria get on and off the worm like travellers on a bus. We have also engineered a third partner, <i>Escherichia coli</i>, to express an interference RNA that promotes clumping. Taken together, our artificial symbiosis allows biotechnologically interesting bacteria to travel on nematodes, reach nutrient-rich biomass spots and maximize the efficiency of biotechnological fermentations in | + | Bacteria are essential in biotechnology, but they can hardly move. Nematodes, such as <i>Caenorhabditis elegans</i>, are fast crawling organisms, but they have limited biotechnological applications. By combining the best from both organisms, we present the first artificial synthetic symbiosis with bacteria engineered to ride on worms, which concentrate in hotspots where bacteria perform a desired biotechnological process, such as bioplastic (PHA) production. We have engineered <i>Pseudomonas putida</i> with a whole operon that allows the formation of a biofilm on the worm. Biofilm formation is switched on and off depending on the media, and thus bacteria get on and off the worm like travellers on a bus. We have also engineered a third partner, <i>Escherichia coli</i>, to express an interference RNA that promotes clumping. Taken together, our artificial symbiosis allows biotechnologically interesting bacteria to travel on nematodes, reach nutrient-rich biomass spots and maximize the efficiency of biotechnological fermentations in heterogeneous substrates. |
</div> | </div> | ||
| Line 70: | Line 65: | ||
<p> | <p> | ||
<b><i>Escherichia coli</i></b> | <b><i>Escherichia coli</i></b> | ||
| - | is a model organism widely | + | is a model organism widely used in fields such as microbiology, molecular biology and genetics. Because of this, there is a great range of genetic manipulation |
| - | techniques | + | techniques related with this Gram negative bacterium. There is also a lot of information about the biochemistry and genetics of this |
| - | microorganism. All this allows to work easily with it in the lab and | + | microorganism. All this allows us to work easily with it in the lab and perform assays testing different conditions that will be tried out in other |
| - | organisms unstudied in such depth. Moreover, <i>E.coli</i> is the main food of C.elegans, organism that is able to incorporate nucleic acids from the | + | organisms unstudied in such depth. Moreover, <i>E.coli</i> is the main food of <i>C.elegans</i>, organism that is able to incorporate nucleic acids from the |
| - | bacteria, RNA, to its cells (Mello et al., 2004). This is the usual mechanism | + | bacteria, RNA, to its cells (Mello et al., 2004). This is the usual mechanism to transform C.elegans, introducing exogenous genetic material from |
| - | the bacteria altering the expression of the worm’s genome in order to make modifications of interest. | + | the bacteria and altering the expression of the worm’s genome in order to make modifications of interest. |
</p> | </p> | ||
<h2>Controlling the mechanism</h2> | <h2>Controlling the mechanism</h2> | ||
<p> | <p> | ||
| - | The role of <i>E.coli</i> in our project is to carry out the synthesis of an iRNA | + | The role of <i>E.coli</i> in our project is to carry out the synthesis of an iRNA to induce the social feeding behaviour of <i>Caenorhabditis elegans</i> |
| - | + | , the <b>clumping. </b>To achieve this, we cloned the biobrick Bba_K1112000 in <i>E.coli</i>, </b></i><b>XL1-Blue strain (fig. 1). </b> | |
</p> | </p> | ||
<div style="clear:both;width:100%;height:auto;overflow:auto;"> | <div style="clear:both;width:100%;height:auto;overflow:auto;"> | ||
| Line 107: | Line 102: | ||
<p> | <p> | ||
This<b> construction</b> consists of the <i>E.coli </i>fadA promoter, which is <b>activated in the presence of fatty acids</b> (Clark, 1981), and the | This<b> construction</b> consists of the <i>E.coli </i>fadA promoter, which is <b>activated in the presence of fatty acids</b> (Clark, 1981), and the | ||
| - | antisense sequence of the mRNA <i>FLP-21 </i>from C.elegans which encodes a protein involved in the solitary feeding behaviour of the worm, the formation | + | antisense sequence of the mRNA <i>FLP-21 </i>from <i>C.elegans</i> which encodes a protein involved in the solitary feeding behaviour of the worm, the formation |
| - | of the dsRNA complex inhibits the expression of this protein, inducing the social feeding | + | of the dsRNA complex inhibits the expression of this protein, inducing the social feeding behaviour (<b>fig. 2)</b>. |
</p> | </p> | ||
<p style="text-align:center"> | <p style="text-align:center"> | ||
| Line 123: | Line 118: | ||
<p> | <p> | ||
<b><i>Pseudomonas putida</i> </b> is a gram-negative bacterium that is found in most soil and water habitats where there is oxygen. Its diverse metabolism and its capacity to break down | <b><i>Pseudomonas putida</i> </b> is a gram-negative bacterium that is found in most soil and water habitats where there is oxygen. Its diverse metabolism and its capacity to break down | ||
| - | organic harmful solvents ( | + | organic harmful solvents (most of its genes are involved in degrading aromatic or aliphatic hydrocarbons) in contaminated soils make this microorganism |
| - | irreplaceable for research studies in the field of bioremediation but also for biosynthesis of value-added products. In addition, <i>Pseudomonas putida</i> has several strains including KT2440, the one we have worked with. This strain can colonize plant roots, from which they take nutrients, while at the same time it offers protection | + | irreplaceable for research studies in the field of bioremediation but also for biosynthesis of value-added products. In addition, <i>Pseudomonas putida</i> has several strains including KT2440, the one we have worked with. This strain can colonize plant roots, from which they take nutrients, while at the same time it offers protection for the plant against pathogens. |
</p> | </p> | ||
<html> | <html> | ||
| Line 130: | Line 125: | ||
<div style="width:550px;float:left;"> | <div style="width:550px;float:left;"> | ||
<p> | <p> | ||
| - | For example, it is capable of converting <a href="http://en.wikipedia.org/wiki/Styrene" target="_blank">styrene</a> oil into the <b><a href="http://en.wikipedia.org/wiki/Biodegradable_plastic" target="_blank">biodegradable | + | For example, it is capable of converting <a href="http://en.wikipedia.org/wiki/Styrene" target="_blank">styrene</a> oil into the <b><a href="http://en.wikipedia.org/wiki/Biodegradable_plastic" target="_blank">biodegradable <a href="http://en.wikipedia.org/wiki/Polyhydroxyalkanoates" target="_blank">PHA</a> plastic</a> </b>. This helps the degradation of the polystyrene foam which |
was thought to be non-biodegradable. Styrene is a major environment toxic pollutant released from industrial sites. The conversion to PHA allows the cure | was thought to be non-biodegradable. Styrene is a major environment toxic pollutant released from industrial sites. The conversion to PHA allows the cure | ||
| - | of styrene pollution but it is also beneficial | + | of styrene pollution but it is also beneficial for society because of its applications in tissue engineering. |
<br/> | <br/> | ||
| - | PHA is also | + | PHA is also environmentally friendly and has a long self-life therefore it is also used in everyday items. Unlike styrene, PHA can break down in soil or water. |
</p> | </p> | ||
<p> | <p> | ||
Within <i>Pseudomonas putida</i>, PHA accumulates under unbalanced growth conditions as a means of intracellular storage, storing excess carbon and energy. | Within <i>Pseudomonas putida</i>, PHA accumulates under unbalanced growth conditions as a means of intracellular storage, storing excess carbon and energy. | ||
| - | These PHA polymers are synthesized by enzyme PHA synthase which is bound to the surface of the PHA granules and uses coenzyme A thioesters of | + | These PHA polymers are synthesized by the enzyme PHA synthase which is bound to the surface of the PHA granules and uses coenzyme A thioesters of |
hydroxyalkanoic acids as substrates. | hydroxyalkanoic acids as substrates. | ||
</p> | </p> | ||
| Line 148: | Line 143: | ||
<p> | <p> | ||
| - | The role of <i>P. putida </i>in the Synthetic symbiosis that we have designed is to be carried by <i>C. elegans </i>to hotspots of interest where would | + | The role of <i>P. putida </i>in the Synthetic symbiosis that we have designed is to be carried by <i>C. elegans </i>to hotspots of interest where it would |
| - | produce bioplastic PHA. | + | produce bioplastic, PHA. |
</p> | </p> | ||
| Line 160: | Line 155: | ||
<p> | <p> | ||
One of the main characters of our work is a nematode known as <b><i>Caenorhabditis elegans, </i></b>from the Rhabditidae family. It was first used as an | One of the main characters of our work is a nematode known as <b><i>Caenorhabditis elegans, </i></b>from the Rhabditidae family. It was first used as an | ||
| - | experimental model in Developmental Genetics studies and nowadays is also used in other fields such as Clinical Biology, Neurobiology and Cell Biology, | + | experimental model in Developmental Genetics studies and nowadays it is also used in other fields such as Clinical Biology, Neurobiology and Cell Biology, |
being a good model to study Alzheimer disease, obesity, diabetes and aging, among others. | being a good model to study Alzheimer disease, obesity, diabetes and aging, among others. | ||
</p> | </p> | ||
| Line 189: | Line 184: | ||
<p> | <p> | ||
When we were considering on creating a new system for the transport of bacteria, we found different key advantages that made the nematode the best option. | When we were considering on creating a new system for the transport of bacteria, we found different key advantages that made the nematode the best option. | ||
| - | For example, <i>C. elegans</i> is able to move very fast around solid substrates (soil | + | For example, <i>C. elegans</i> is able to move very fast around solid substrates (soil actually is its natural habitat) and also in agar, so it’s very |
useful for both lab experiments and real-environment tests. | useful for both lab experiments and real-environment tests. | ||
</p> | </p> | ||
| Line 199: | Line 194: | ||
</p> | </p> | ||
<p> | <p> | ||
| - | The situation changes when there is an attractant in the medium. Here, our 'transport' begins to direct | + | The situation changes when there is an attractant in the medium. Here, our 'transport system' begins to direct its movements to the focus of the substance |
| - | (volatile or soluble) which acts as an attractant in a process known as <b>'chemotaxis'</b> thanks to the amazing smell of our nematode. This allows us to | + | (volatile or soluble) which acts as an attractant in a process known as <b>'chemotaxis'</b> thanks to the amazing sense of smell of our nematode. This allows us to |
‘guide’ the nematodes <b>towards defined spots in irregular substrates.</b> | ‘guide’ the nematodes <b>towards defined spots in irregular substrates.</b> | ||
</p> | </p> | ||
| Line 218: | Line 213: | ||
</p> | </p> | ||
<p> | <p> | ||
| - | There are several strains of our nematode. The one that drew our attention was called 'N2', which had a fairly interesting | + | There are several strains of our nematode. The one that drew our attention was called 'N2', which had a fairly interesting behaviour: under normal |
| - | conditions, it eats individually, whereas under certain conditions (such as starvation), a social feeding | + | conditions, it eats individually, whereas under certain conditions (such as starvation), a social feeding behaviour known as <b>“clumping”</b> is induced. |
But this phenomenon can also be induced if the expression of some particular genes is interfered. This fact gave us the opportunity to develop the <b>first artificial symbiosis between worms and bacteria</b>, based on the manipulation of the behaviour of <i>C. elegans</i> by simply nourishing it with | But this phenomenon can also be induced if the expression of some particular genes is interfered. This fact gave us the opportunity to develop the <b>first artificial symbiosis between worms and bacteria</b>, based on the manipulation of the behaviour of <i>C. elegans</i> by simply nourishing it with | ||
transformed <i>E.coli </i>able to synthesize the iRNA. | transformed <i>E.coli </i>able to synthesize the iRNA. | ||
| Line 240: | Line 235: | ||
<div class="tab-pane fade" id="Results"> | <div class="tab-pane fade" id="Results"> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
<p> | <p> | ||
| - | + | We have created a <b>Results Section</b> to organise our results depending on the activity they belong to (<b>The Riding, The Calling, The Clumping and The Building</b>) in order to make it more intuitive. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</p> | </p> | ||
<p style="text-align:center"> | <p style="text-align:center"> | ||
| - | < | + | <a href="https://2013.igem.org/Team:Valencia_Biocampus/Results" class="btn btn-primary">Go to Results section</a> |
</p> | </p> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| Line 697: | Line 257: | ||
Remember that the goal of proper part documentation is to describe and define a part such that it can be used without a need to refer to the primary literature. The next iGEM team should be able to read your documentation and be able to use the part successfully. Also, you should provide proper references to acknowledge previous authors and to provide for users who wish to know more. | Remember that the goal of proper part documentation is to describe and define a part such that it can be used without a need to refer to the primary literature. The next iGEM team should be able to read your documentation and be able to use the part successfully. Also, you should provide proper references to acknowledge previous authors and to provide for users who wish to know more. | ||
| + | <groupparts>iGEM013 Valencia_Biocampus</groupparts> | ||
| + | |||
--> | --> | ||
</html> | </html> | ||
| - | |||
<html> | <html> | ||
| Line 712: | Line 273: | ||
<th> Designer </th> | <th> Designer </th> | ||
<th> Length </th> | <th> Length </th> | ||
| - | <th> | + | <th> Fav </th> |
</tr> | </tr> | ||
</thead> | </thead> | ||
| Line 721: | Line 282: | ||
<td> fadB promoter + FLP-21 iRNA </td> | <td> fadB promoter + FLP-21 iRNA </td> | ||
<td> Pedro Luis Dorado Morales </td> | <td> Pedro Luis Dorado Morales </td> | ||
| - | <td> </td> | + | <td> 344 </td> |
| - | <td> </td> | + | <td> <img src="https://2013.igem.org/common/tablesorter/themes/groupparts/heart13.gif" alt="" /></td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
<td><img class="table-cross" src="https://static.igem.org/mediawiki/2013/0/01/CheckIcon.gif" alt="Yes" height="24" width="24"/></td> | <td><img class="table-cross" src="https://static.igem.org/mediawiki/2013/0/01/CheckIcon.gif" alt="Yes" height="24" width="24"/></td> | ||
<td> <a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K1112001">BBa_K1112001</a> </td> | <td> <a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K1112001">BBa_K1112001</a> </td> | ||
| - | <td> | + | <td> Coding </td> |
<td> pGlnA + hmsHFRS operon </td> | <td> pGlnA + hmsHFRS operon </td> | ||
<td> Alba Iglesias Vilches </td> | <td> Alba Iglesias Vilches </td> | ||
| - | <td> </td> | + | <td> 6342</td> |
<td> </td> | <td> </td> | ||
</tr> | </tr> | ||
| Line 736: | Line 297: | ||
<td><img class="table-check" src="https://static.igem.org/mediawiki/2013/0/01/CheckIcon.gif" alt="Yes" height="24" width="24"/></td> | <td><img class="table-check" src="https://static.igem.org/mediawiki/2013/0/01/CheckIcon.gif" alt="Yes" height="24" width="24"/></td> | ||
<td> <a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K1112002">BBa_K1112002</a> </td> | <td> <a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K1112002">BBa_K1112002</a> </td> | ||
| - | <td> | + | <td> Coding </td> |
<td> Cluster PHA </td> | <td> Cluster PHA </td> | ||
<td> Alba Iglesias Vilches </td> | <td> Alba Iglesias Vilches </td> | ||
| - | <td> </td> | + | <td> 4978</td> |
<td> </td> | <td> </td> | ||
</tr> | </tr> | ||
| Line 754: | Line 315: | ||
<p> | <p> | ||
<a href="http://parts.igem.org/Part:BBa_K1112001"><h3>BBa_K1112001: pGlnA + hmsHFRS operon</h3></a> | <a href="http://parts.igem.org/Part:BBa_K1112001"><h3>BBa_K1112001: pGlnA + hmsHFRS operon</h3></a> | ||
| - | We reused our nitrogen sensitive promoter from one of our last year constructs and we | + | We reused our nitrogen sensitive promoter from one of our last year constructs and we used it to control the expression of the operon that triggers the formation of a biofilm over <i>C.elegans</i>. |
</p> | </p> | ||
| Line 761: | Line 322: | ||
This BioBrick contains the complete natural sequence that induces the production of PHA in <i>Pseudomonas putida</i> (KT2440). | This BioBrick contains the complete natural sequence that induces the production of PHA in <i>Pseudomonas putida</i> (KT2440). | ||
</p> | </p> | ||
| + | |||
| + | </div> | ||
| + | |||
| + | |||
| + | <div class="tab-pane fade" id="Notebook"> | ||
| + | </html> | ||
| + | |||
| + | ==Notebook== | ||
| + | <html> | ||
| + | <p> | ||
| + | During the whole project, we have been working on a notebook in order to show the world what we have been working on and to collect all our progress. | ||
| + | </p> | ||
| + | <p>You'll be able to see our results, the picture of the day, the song that was being played in the lab while we were working and our iGEM News.</p> | ||
| + | <p style="text-align:center"> | ||
| + | <a href="https://2013.igem.org/Team:Valencia_Biocampus/Notebook" class="btn btn-primary">Go to Notebook</a> | ||
| + | </p> | ||
| + | |||
</div> | </div> | ||
Latest revision as of 19:18, 24 October 2013
Project Overview
Why E.coli
Escherichia coli is a model organism widely used in fields such as microbiology, molecular biology and genetics. Because of this, there is a great range of genetic manipulation techniques related with this Gram negative bacterium. There is also a lot of information about the biochemistry and genetics of this microorganism. All this allows us to work easily with it in the lab and perform assays testing different conditions that will be tried out in other organisms unstudied in such depth. Moreover, E.coli is the main food of C.elegans, organism that is able to incorporate nucleic acids from the bacteria, RNA, to its cells (Mello et al., 2004). This is the usual mechanism to transform C.elegans, introducing exogenous genetic material from the bacteria and altering the expression of the worm’s genome in order to make modifications of interest.
Controlling the mechanism
The role of E.coli in our project is to carry out the synthesis of an iRNA to induce the social feeding behaviour of Caenorhabditis elegans , the clumping. To achieve this, we cloned the biobrick Bba_K1112000 in E.coli, XL1-Blue strain (fig. 1).

Fig. 1.
- pSB1C3 is a high copy number plasmid (RFC [10]) carrying chloramphenicol resistance.
- The replication origin is a pUC19-derived pMB1 (copy number of 100-300 per cell).
- pSB1C3 has terminators bracketing its MCS which are designed to prevent transcription from *inside* the MCS from reading out into the vector. The efficiency of these terminators is known to be < 100%. Ideally we would construct a future set of terminators for bracketing a MCS that were 100% efficient in terminating both into and out of the MCS region.
This construction consists of the E.coli fadA promoter, which is activated in the presence of fatty acids (Clark, 1981), and the antisense sequence of the mRNA FLP-21 from C.elegans which encodes a protein involved in the solitary feeding behaviour of the worm, the formation of the dsRNA complex inhibits the expression of this protein, inducing the social feeding behaviour (fig. 2).

Pseudomonas putida and PHA production
Pseudomonas putida is a gram-negative bacterium that is found in most soil and water habitats where there is oxygen. Its diverse metabolism and its capacity to break down organic harmful solvents (most of its genes are involved in degrading aromatic or aliphatic hydrocarbons) in contaminated soils make this microorganism irreplaceable for research studies in the field of bioremediation but also for biosynthesis of value-added products. In addition, Pseudomonas putida has several strains including KT2440, the one we have worked with. This strain can colonize plant roots, from which they take nutrients, while at the same time it offers protection for the plant against pathogens.
For example, it is capable of converting styrene oil into the biodegradable PHA plastic . This helps the degradation of the polystyrene foam which
was thought to be non-biodegradable. Styrene is a major environment toxic pollutant released from industrial sites. The conversion to PHA allows the cure
of styrene pollution but it is also beneficial for society because of its applications in tissue engineering.
PHA is also environmentally friendly and has a long self-life therefore it is also used in everyday items. Unlike styrene, PHA can break down in soil or water.
Within Pseudomonas putida, PHA accumulates under unbalanced growth conditions as a means of intracellular storage, storing excess carbon and energy. These PHA polymers are synthesized by the enzyme PHA synthase which is bound to the surface of the PHA granules and uses coenzyme A thioesters of hydroxyalkanoic acids as substrates.

The role of P. putida in the Synthetic symbiosis that we have designed is to be carried by C. elegans to hotspots of interest where it would produce bioplastic, PHA.
An overview about our nematode
One of the main characters of our work is a nematode known as Caenorhabditis elegans, from the Rhabditidae family. It was first used as an experimental model in Developmental Genetics studies and nowadays it is also used in other fields such as Clinical Biology, Neurobiology and Cell Biology, being a good model to study Alzheimer disease, obesity, diabetes and aging, among others.
Another interesting thing is that it feeds on Escherichia coli. Its “favorite” strain is OP50, although we checked that it’s also able to feed on XL1-Blue strain, the one that we used in all our molecular biology experiments.
Some advantages of C. elegans when compared with other model organisms are:
- Lifespan ranges between 2 and 3 weeks, so experimentation times are reduced.
- Its maintenance and study is cheap and simple (it is transparent, which facilitates microscopic observation).
- It is very small (1 mm), so it is possible to carry out experiments with a huge number of worms in a small Petri dish having a great statistical support.
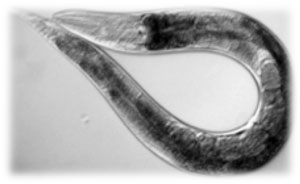
Why use C. elegans as a transport?
When we were considering on creating a new system for the transport of bacteria, we found different key advantages that made the nematode the best option. For example, C. elegans is able to move very fast around solid substrates (soil actually is its natural habitat) and also in agar, so it’s very useful for both lab experiments and real-environment tests.
Its movement, in addition to being very fast, has two modes: random and directed.
When there is no attractant in the medium, C. elegans moves doing uncoordinated movements in several directions in what is known as 'random walk'.
The situation changes when there is an attractant in the medium. Here, our 'transport system' begins to direct its movements to the focus of the substance (volatile or soluble) which acts as an attractant in a process known as 'chemotaxis' thanks to the amazing sense of smell of our nematode. This allows us to ‘guide’ the nematodes towards defined spots in irregular substrates.
Chemotaxis is the foundation to guide the transport of bacteria and is therefore the focus of experimentation with C. elegans, with the aim of finding the best attractant. This ability makes C. elegans a perfect 'bus' for bacteria. (simuelegans online here)
Moreover, we found two pathogens (Yersinia pestis and Xhenorhabus nematophila) with the ability to form biofilms on C. elegans thanks to the proteins of the operon hmsHFRS. When genetically-engineered strains of commonly used bacteria such as Escherichia coli or Pseudomonas putida express this hmsHFRS operon, they have the 'ticket' to travel: they are able to adhere to the worm’s surface by means of forming a synthetic biofilm.
Expanding our knowledge about C. elegans...
Once we knew that C. elegans was the best option, we began to discover interesting things for further study.
There are several strains of our nematode. The one that drew our attention was called 'N2', which had a fairly interesting behaviour: under normal conditions, it eats individually, whereas under certain conditions (such as starvation), a social feeding behaviour known as “clumping” is induced. But this phenomenon can also be induced if the expression of some particular genes is interfered. This fact gave us the opportunity to develop the first artificial symbiosis between worms and bacteria, based on the manipulation of the behaviour of C. elegans by simply nourishing it with transformed E.coli able to synthesize the iRNA.
With it, while C. elegans acts as transport, bacteria return the favour giving it the ability to eat in company.

We have created a Results Section to organise our results depending on the activity they belong to (The Riding, The Calling, The Clumping and The Building) in order to make it more intuitive.
Parts
These are the BioBricks we have designed, constructed, and characterized. We have submitted them to the Registry of Standard Biological Parts
| Works? | Name | Type | Description | Designer | Length | Fav |
|---|---|---|---|---|---|---|
| BBa_K1112000 | Regulatory | fadB promoter + FLP-21 iRNA | Pedro Luis Dorado Morales | 344 |  |
|
| BBa_K1112001 | Coding | pGlnA + hmsHFRS operon | Alba Iglesias Vilches | 6342 | ||
| BBa_K1112002 | Coding | Cluster PHA | Alba Iglesias Vilches | 4978 |
BBa_K1112000: fadB promoter + FLP-21 iRNA
This construction is made up by a fatty acid-sensitive promoter that directs the transcription of an iRNA responsible for the social or solitary behavior of C.elegans.
BBa_K1112001: pGlnA + hmsHFRS operon
We reused our nitrogen sensitive promoter from one of our last year constructs and we used it to control the expression of the operon that triggers the formation of a biofilm over C.elegans.
BBa_K1112002: cluster PHA
This BioBrick contains the complete natural sequence that induces the production of PHA in Pseudomonas putida (KT2440).
Notebook
During the whole project, we have been working on a notebook in order to show the world what we have been working on and to collect all our progress.
You'll be able to see our results, the picture of the day, the song that was being played in the lab while we were working and our iGEM News.
 "
"


