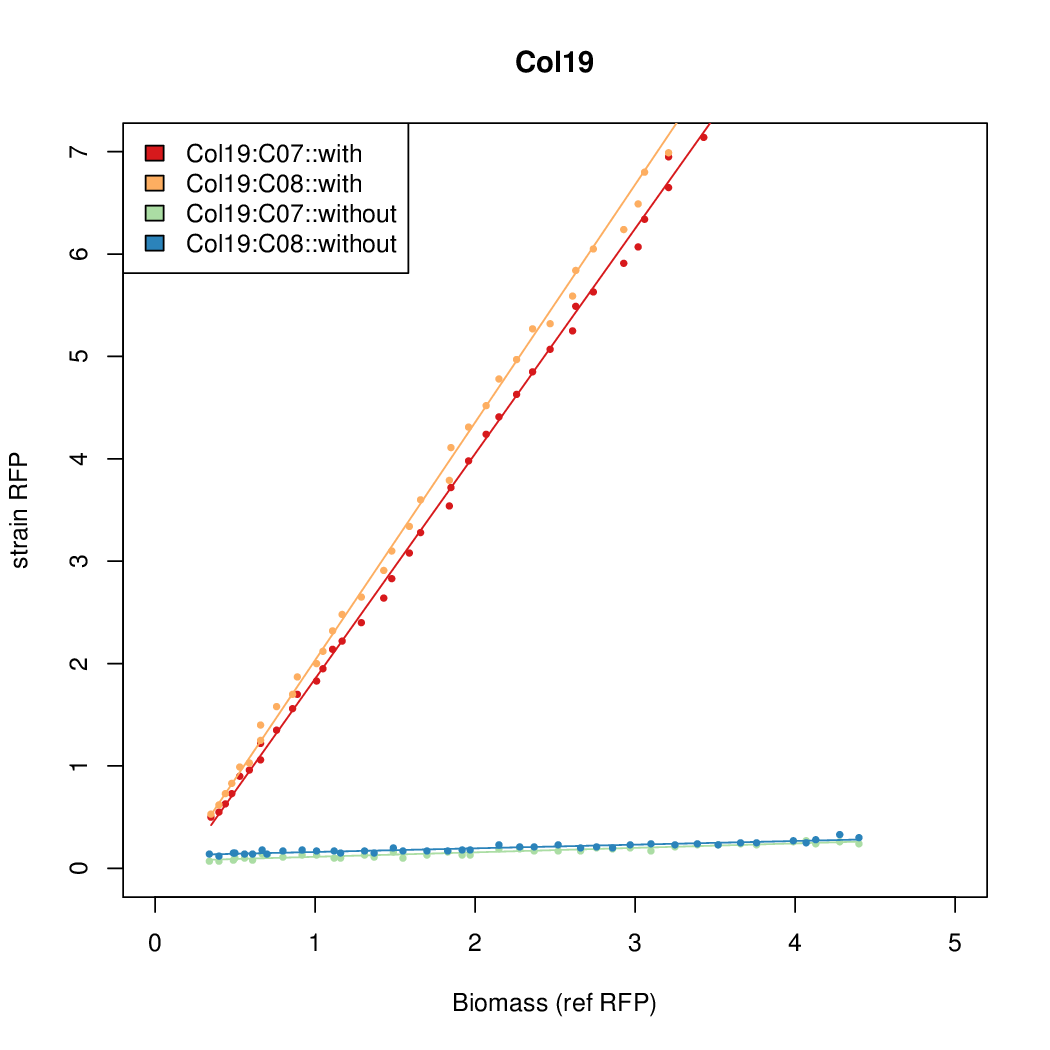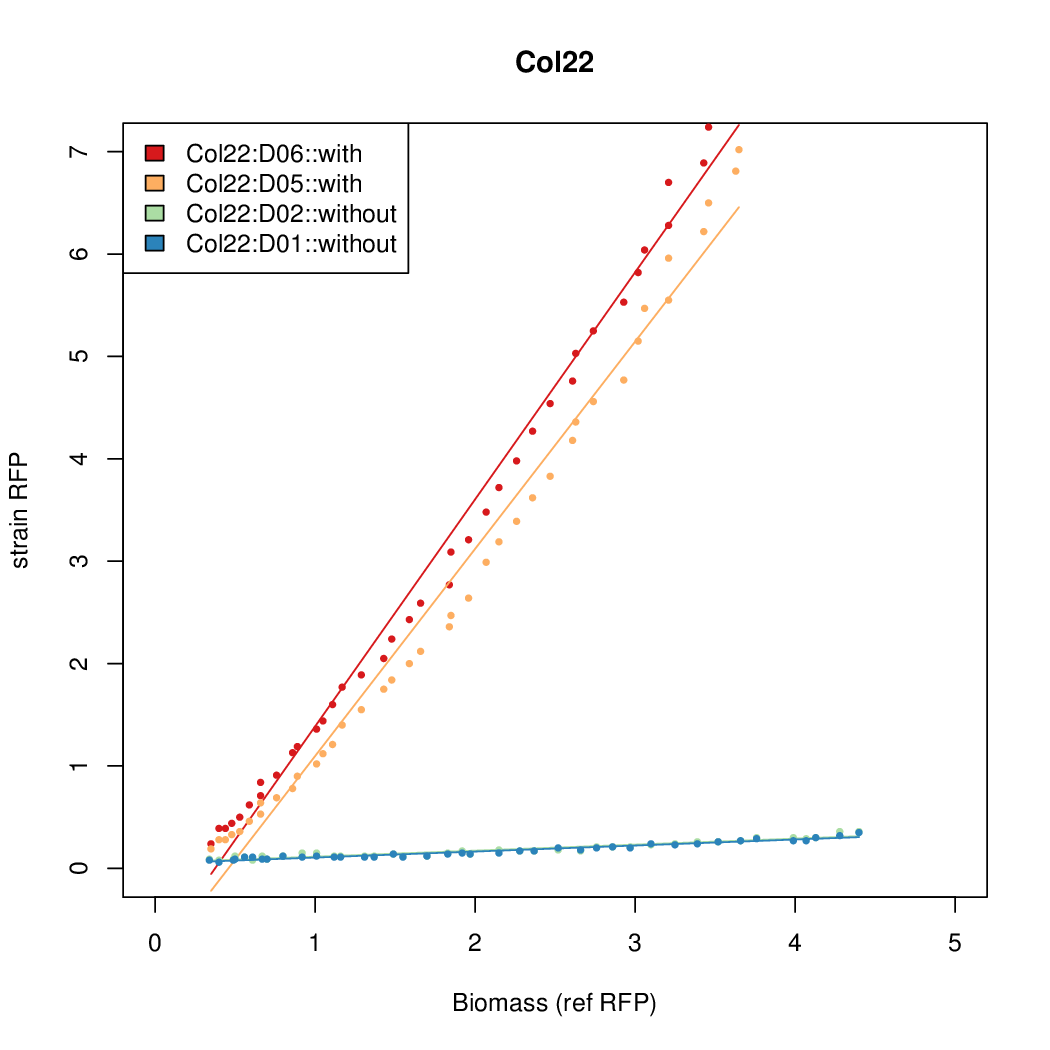Team:DTU-Denmark/pBAD SPL
From 2013.igem.org
(→Details) |
(→Summary) |
||
| Line 29: | Line 29: | ||
=== Summary === | === Summary === | ||
| + | Promoter activity when induced (with arabinose added) plotted vs basal activity (without arabinose; ie leakiness of the promoter). The colonies that we selected all show less activity than the the constitutive promoter, and when induced, show higher activity than the constitutive promoter. | ||
[[File:Induced_vs_basal.png]] | [[File:Induced_vs_basal.png]] | ||
Revision as of 08:40, 1 October 2013
pBAD SPL
Contents |
pBAD synthetic promoter library
As a tool for expressing lethal proteins in E. coli we made a synthetic promoter library (SPL, [http://dspace.mit.edu/handle/1721.1/60080 RFC 63]) with the pBAD arabinose inducible promoter. The concept was taken from the DTU iGEM team from 2010.
Methods
Experimental procedure
- Random promoter sequences were ordered matching the sequence CTGACGNNNNNNNNNNNNNNNNNNTAWWATNNNNA.
- USER cloning to add RFP downstream of promoter.
- Colonies were plated.
- Colonies that were not red prior to induction with arabinose but that did turn red after induction with arabinose were selected.
- Wells were inoculated from overnight cultures of each of the selected colonies
- One plate was run with arabinose, and another plate was run without arabinose.
Data analysis
- Data was collected from the Biolector, and analyzed using a series of R scripts written by Chris Workman (unpublished).
- The maturation and degradation times for mCherry were both assumed to be 40 min. TODOref
- The growth rate, mu, was estimated to be 1.28 (from an average of all wells on all plates) since we expect each strain to grow at the same rate.
- A time window representing exponential growth was selected (between 1 and 4.5 hours).
- The RFP measurement for a constitutively expressed strain was used as a standard measure of growth. This is plotted on the x-axis in the detailed plots per colony below.
- Figures were plotted using R.
Results
Summary
Promoter activity when induced (with arabinose added) plotted vs basal activity (without arabinose; ie leakiness of the promoter). The colonies that we selected all show less activity than the the constitutive promoter, and when induced, show higher activity than the constitutive promoter.

Details
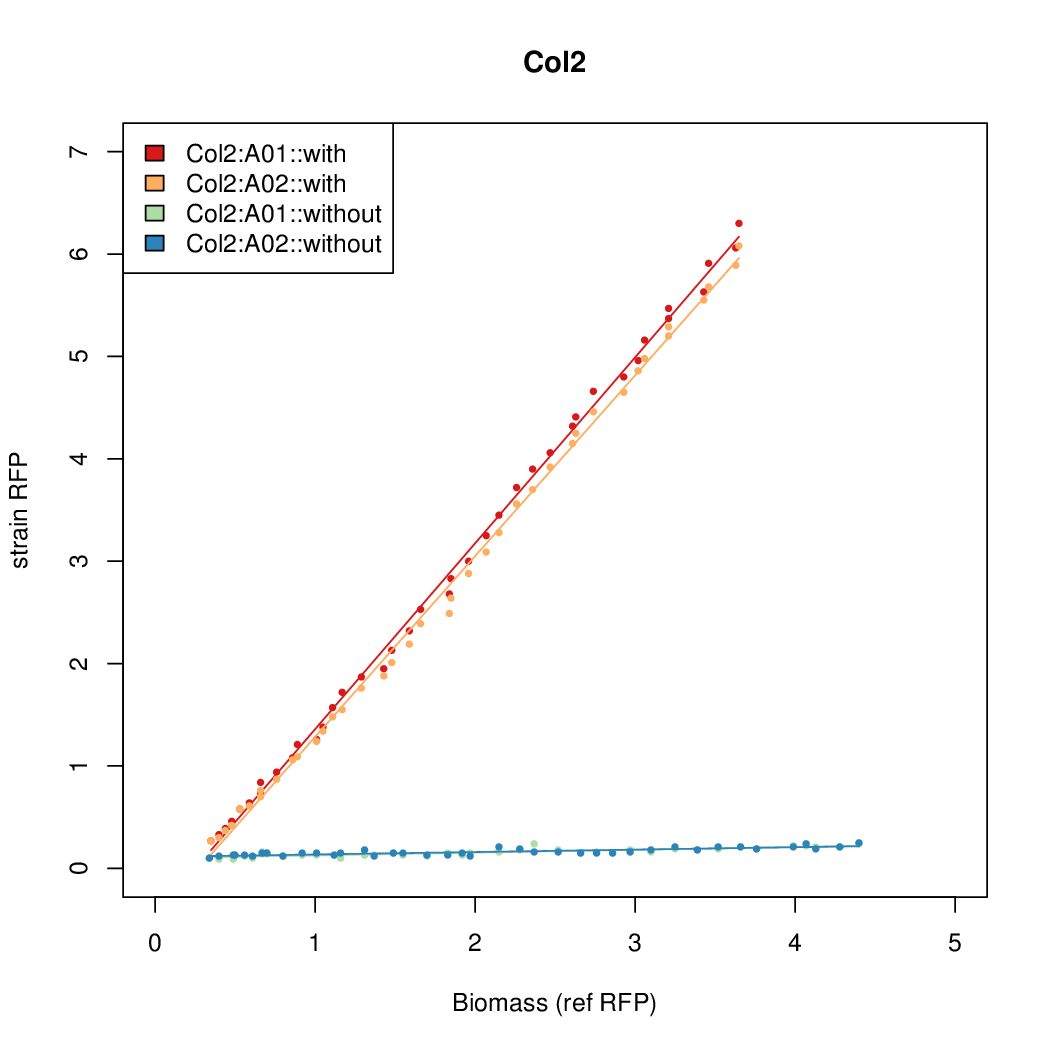
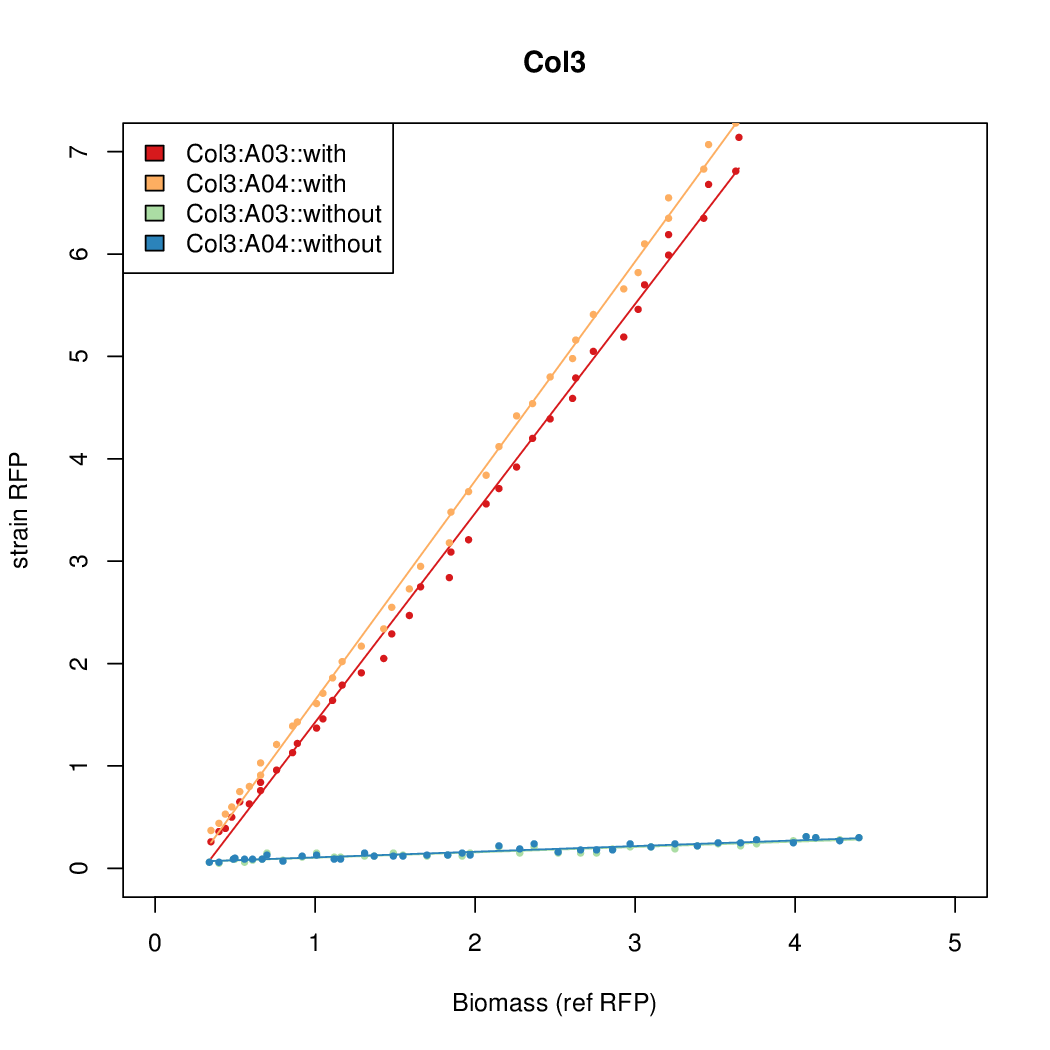
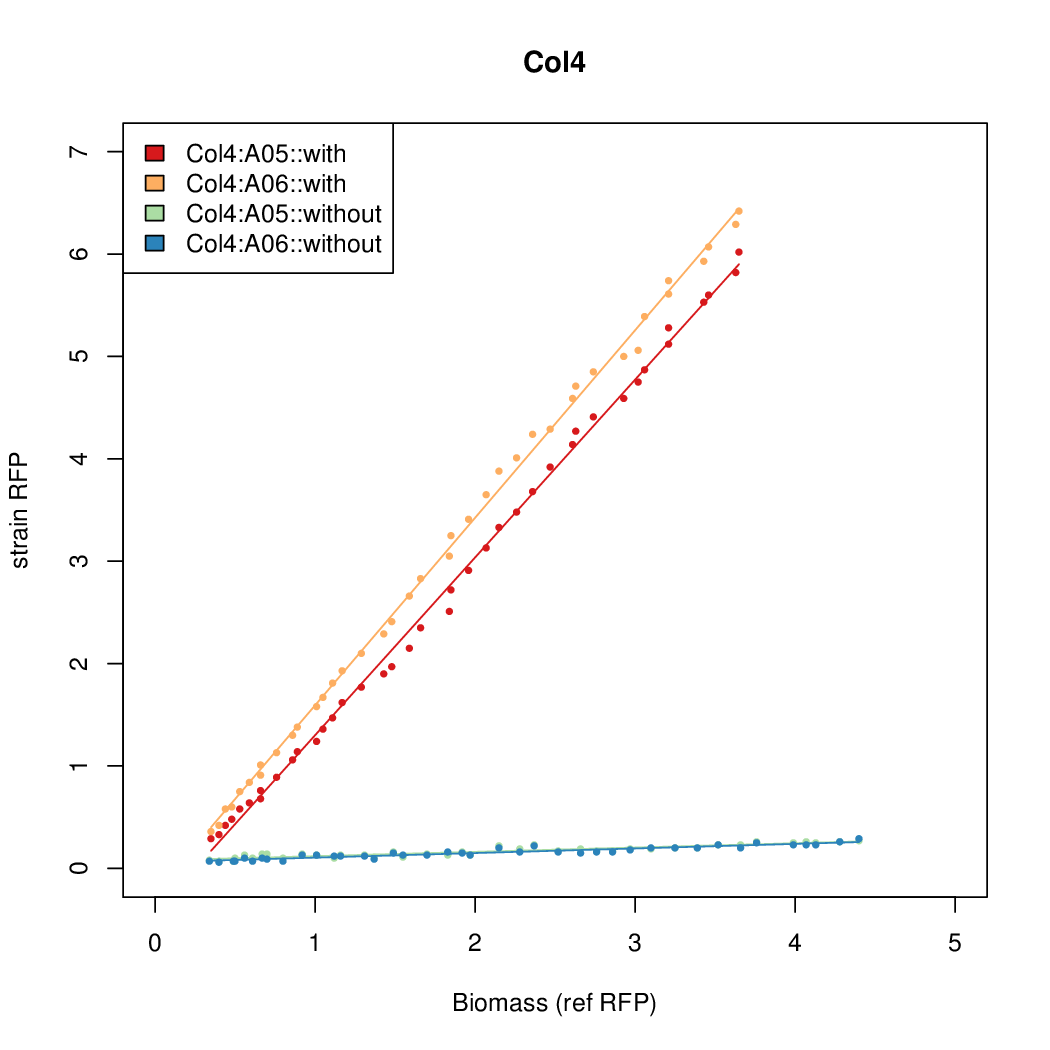
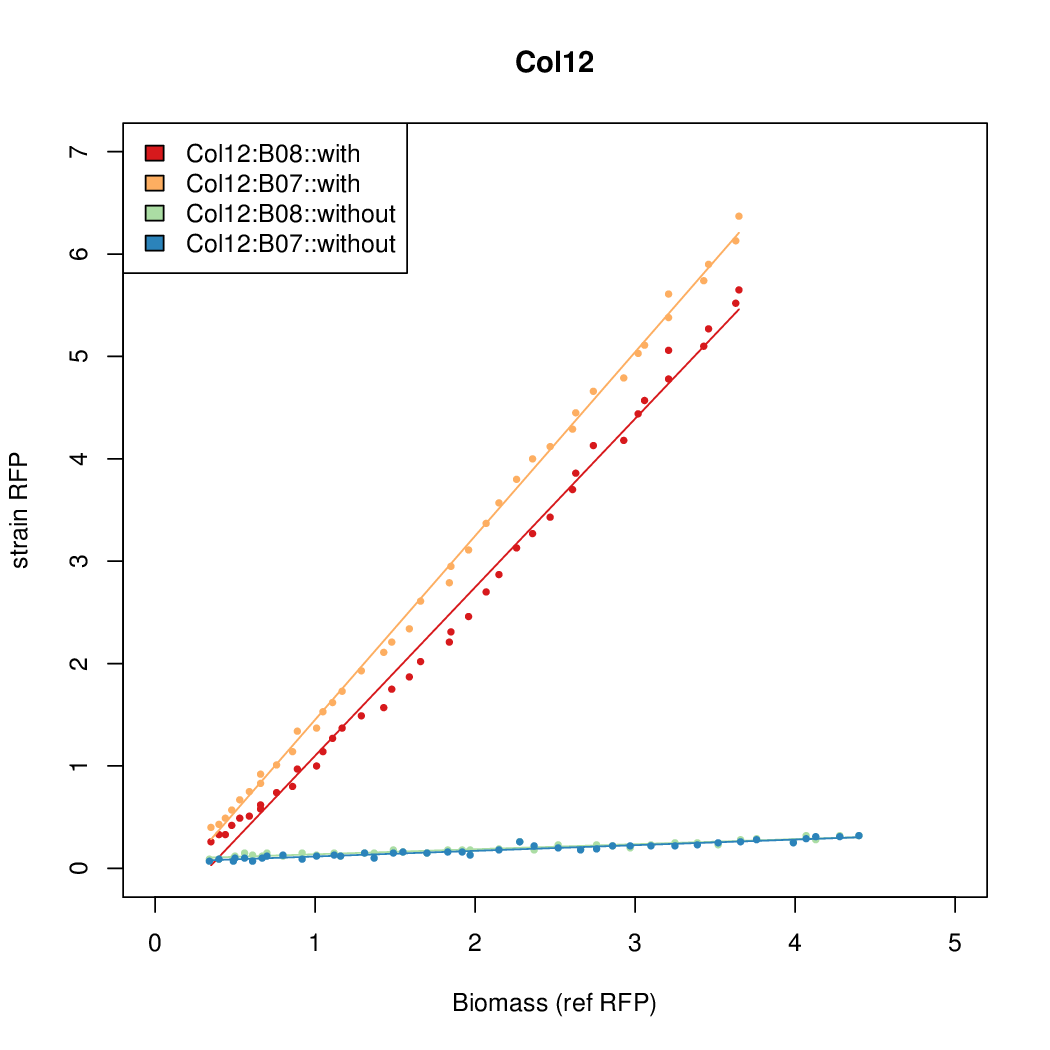
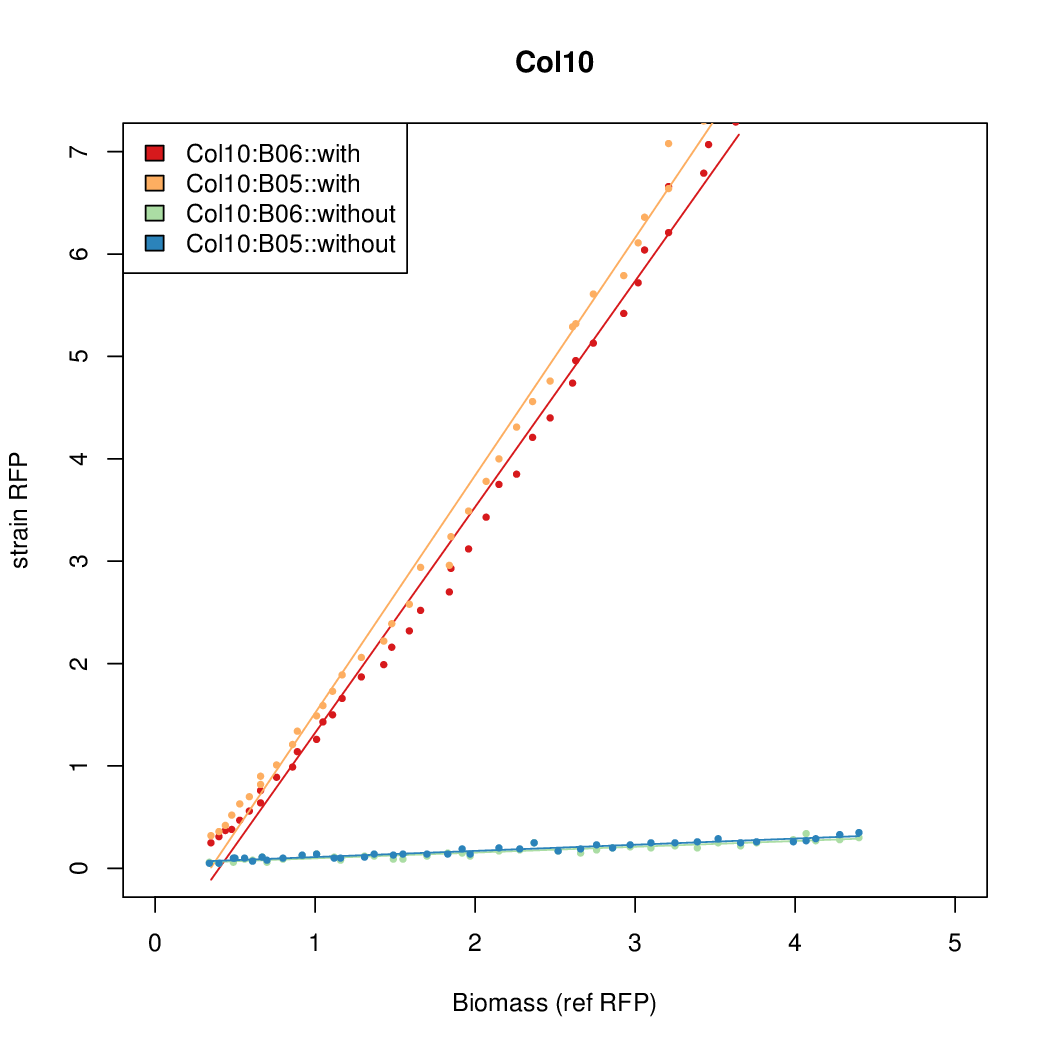
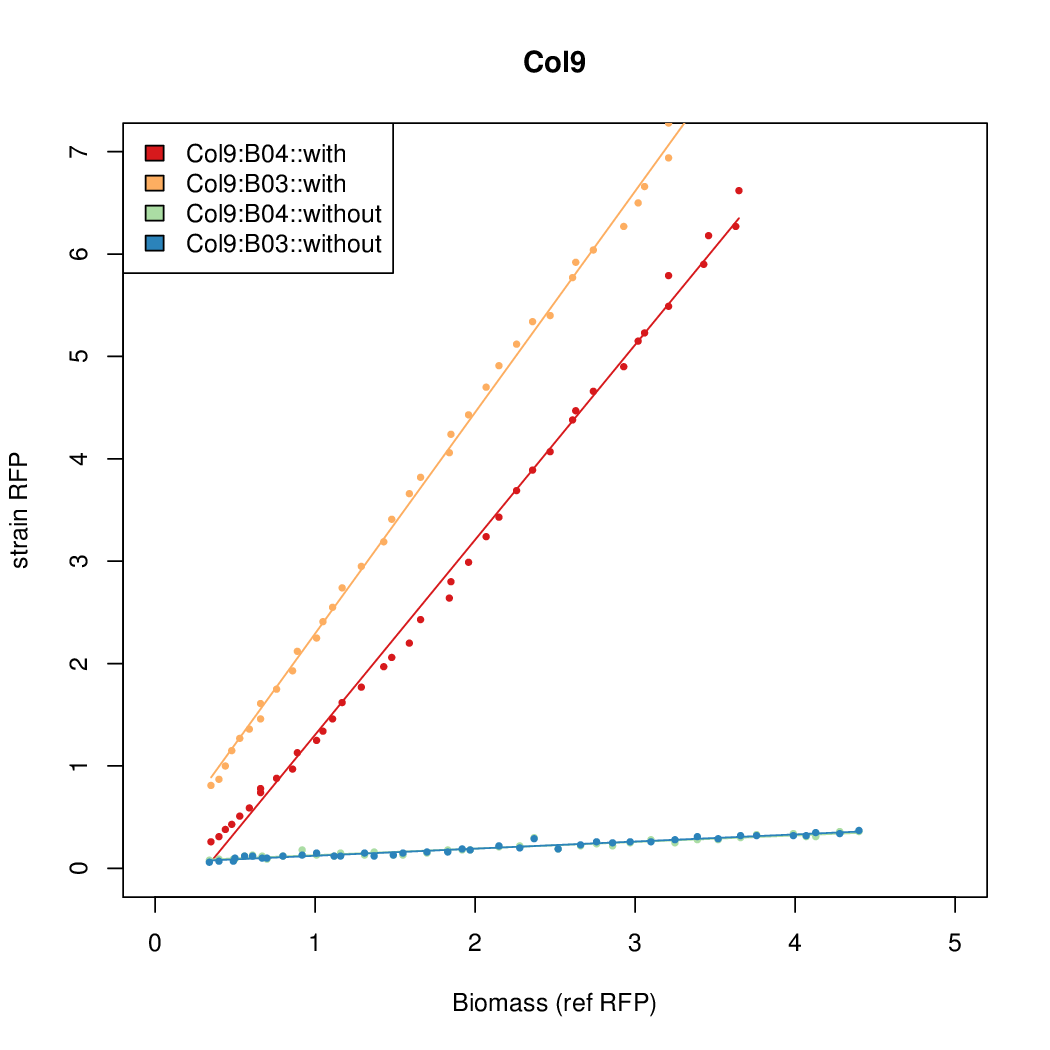
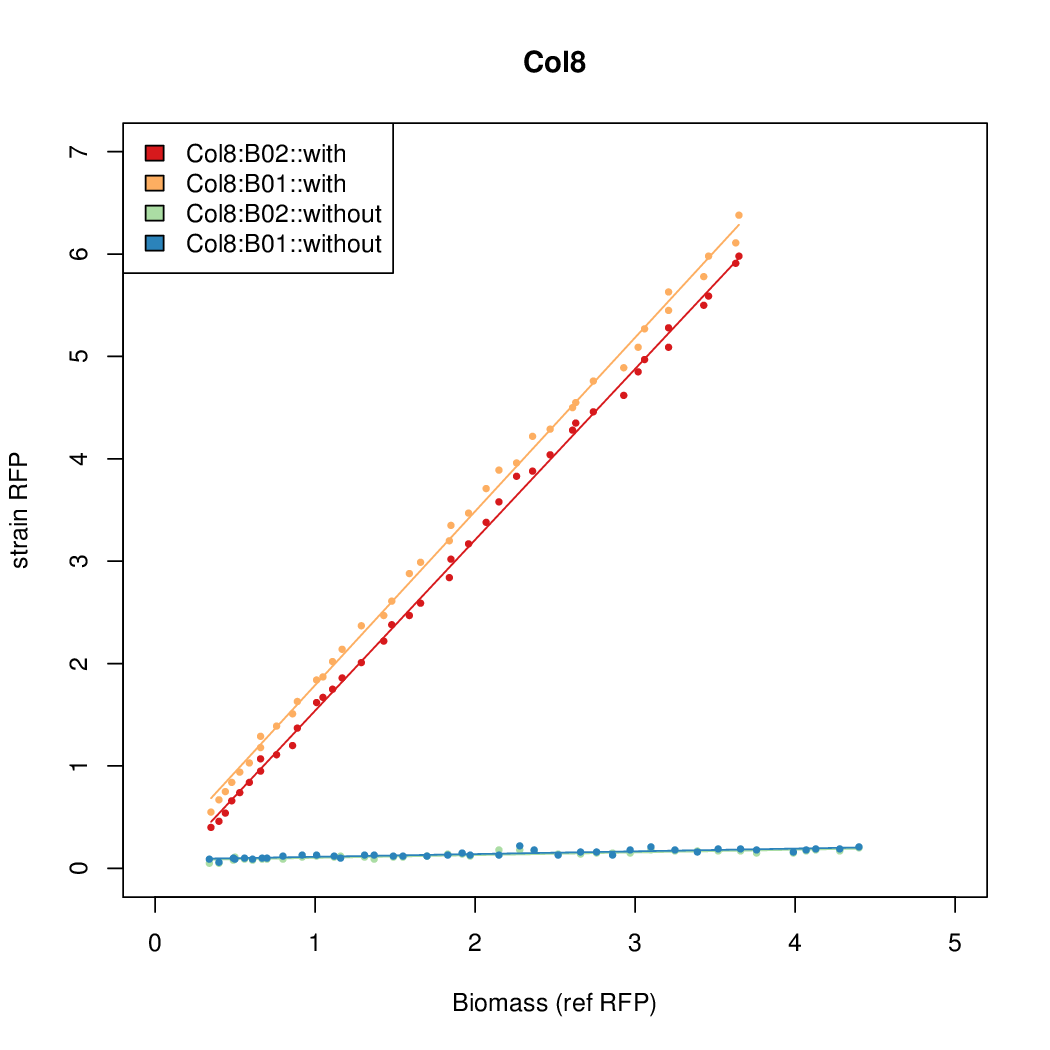
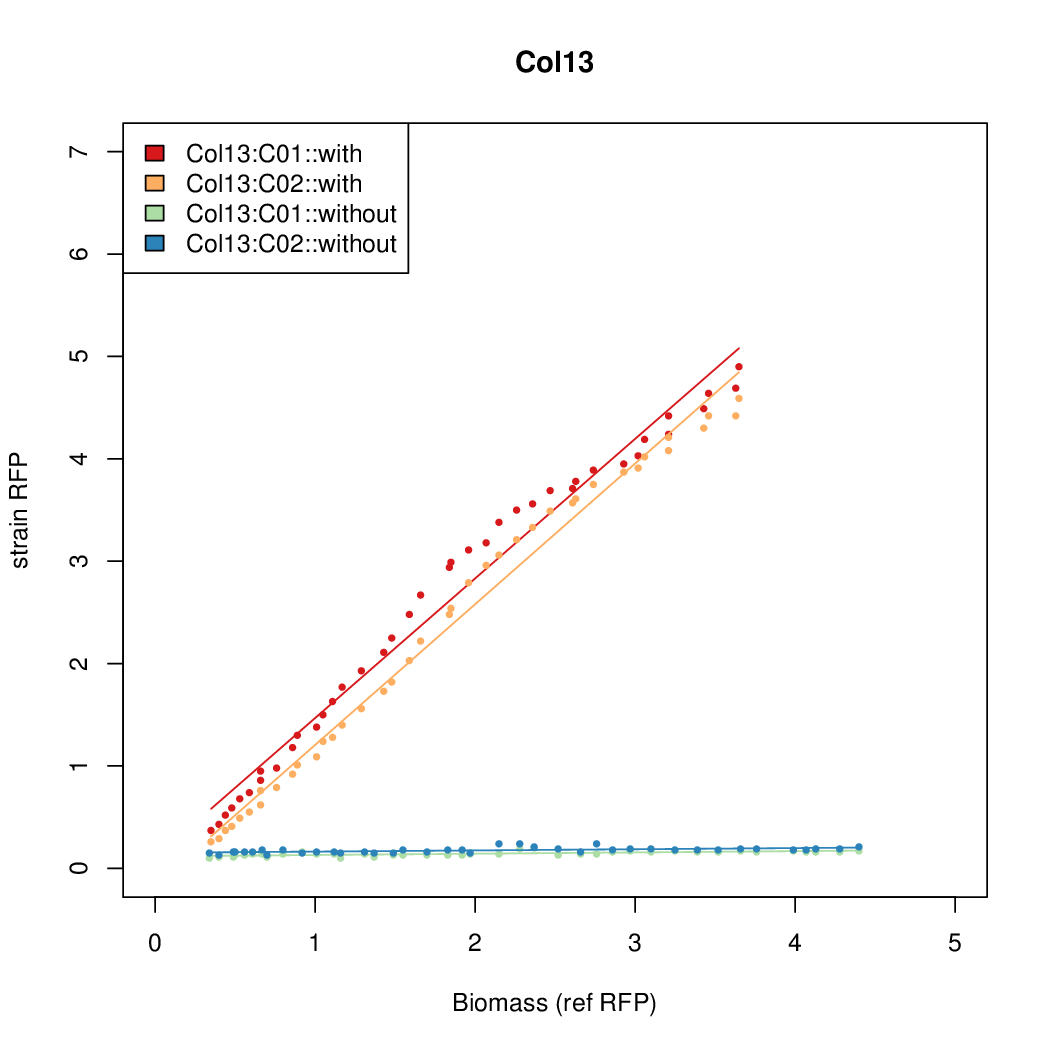
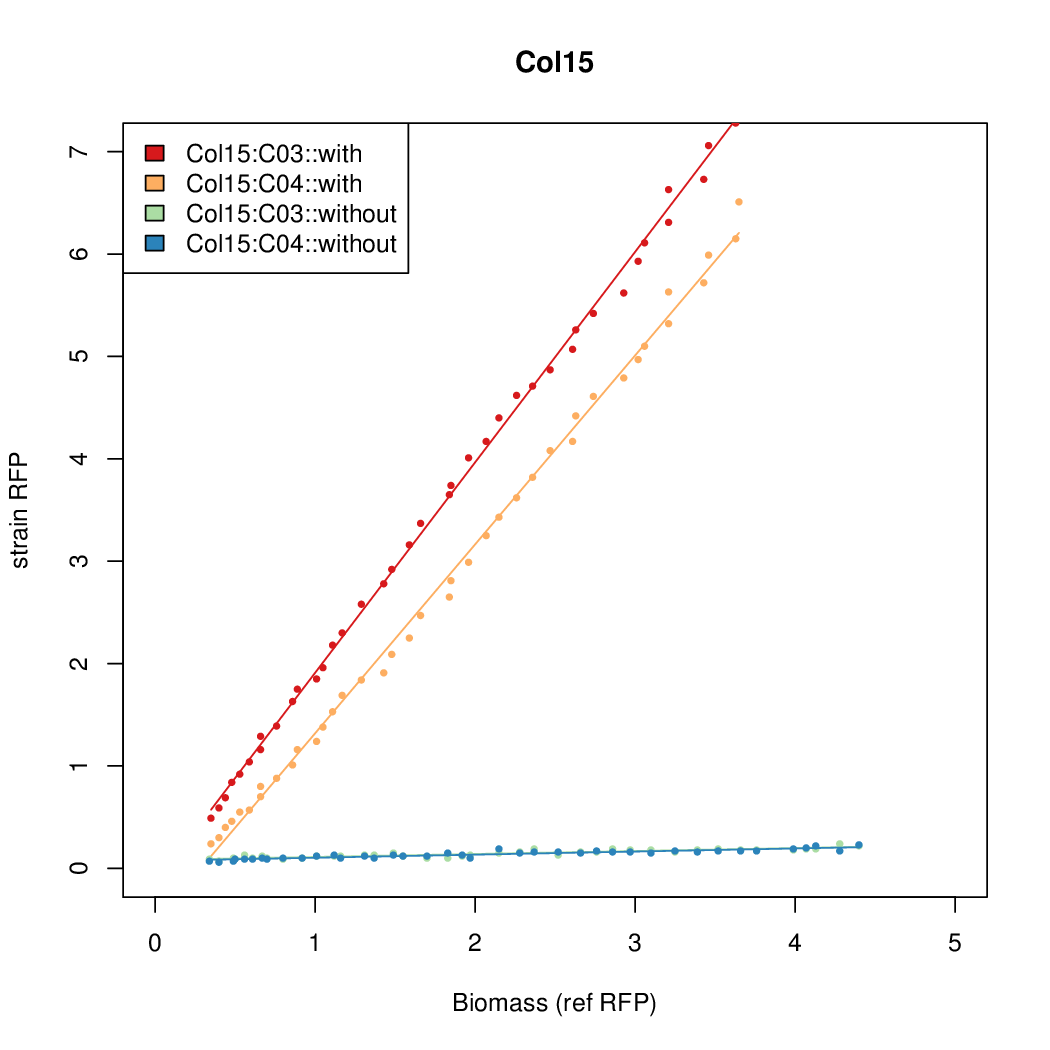
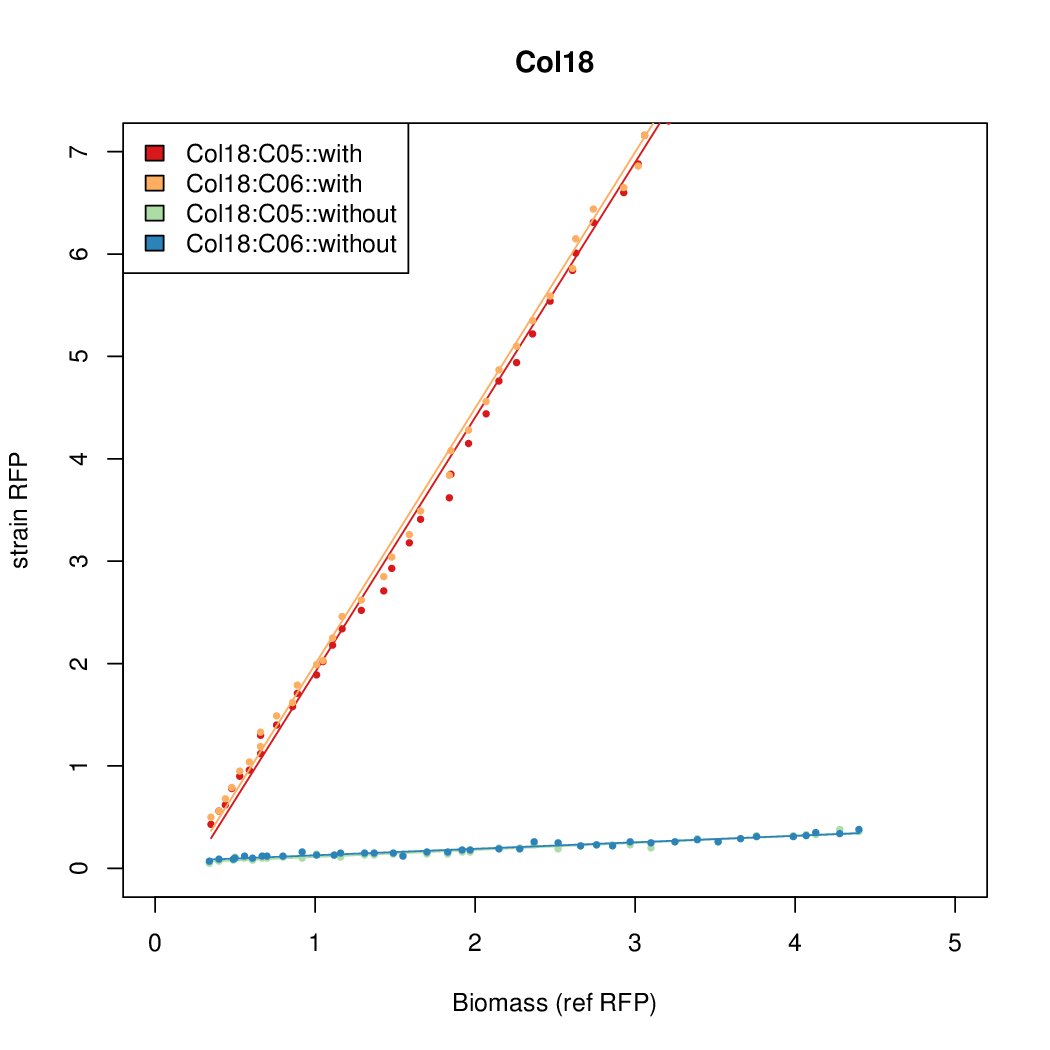
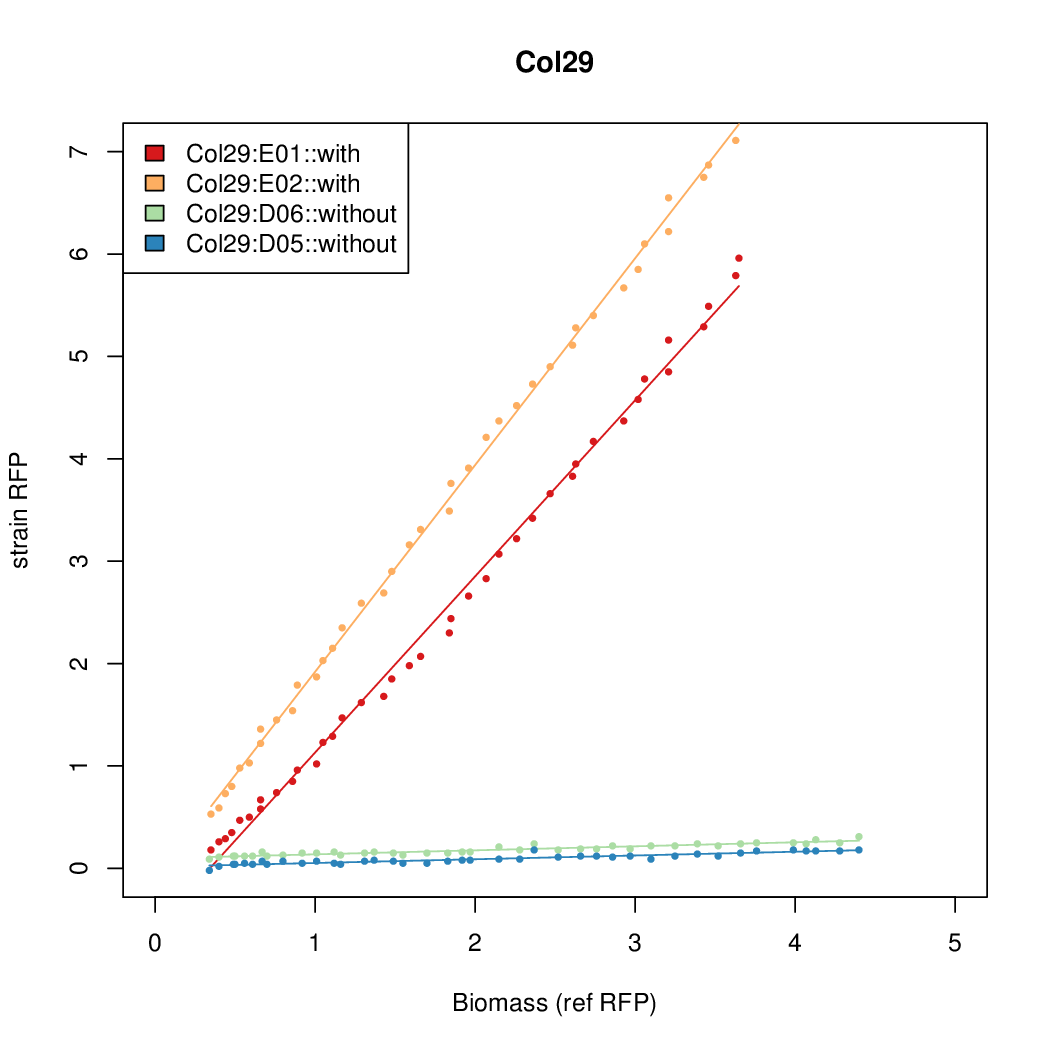
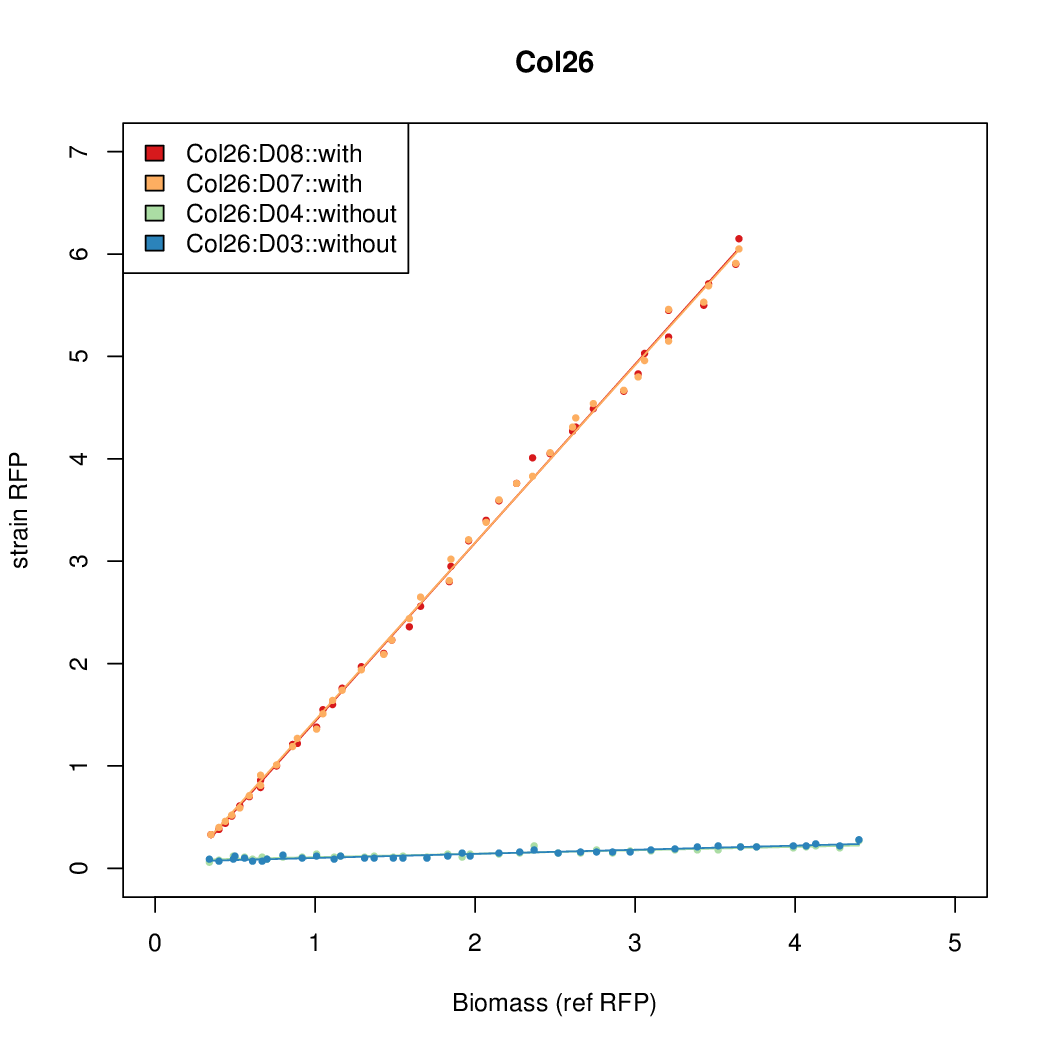
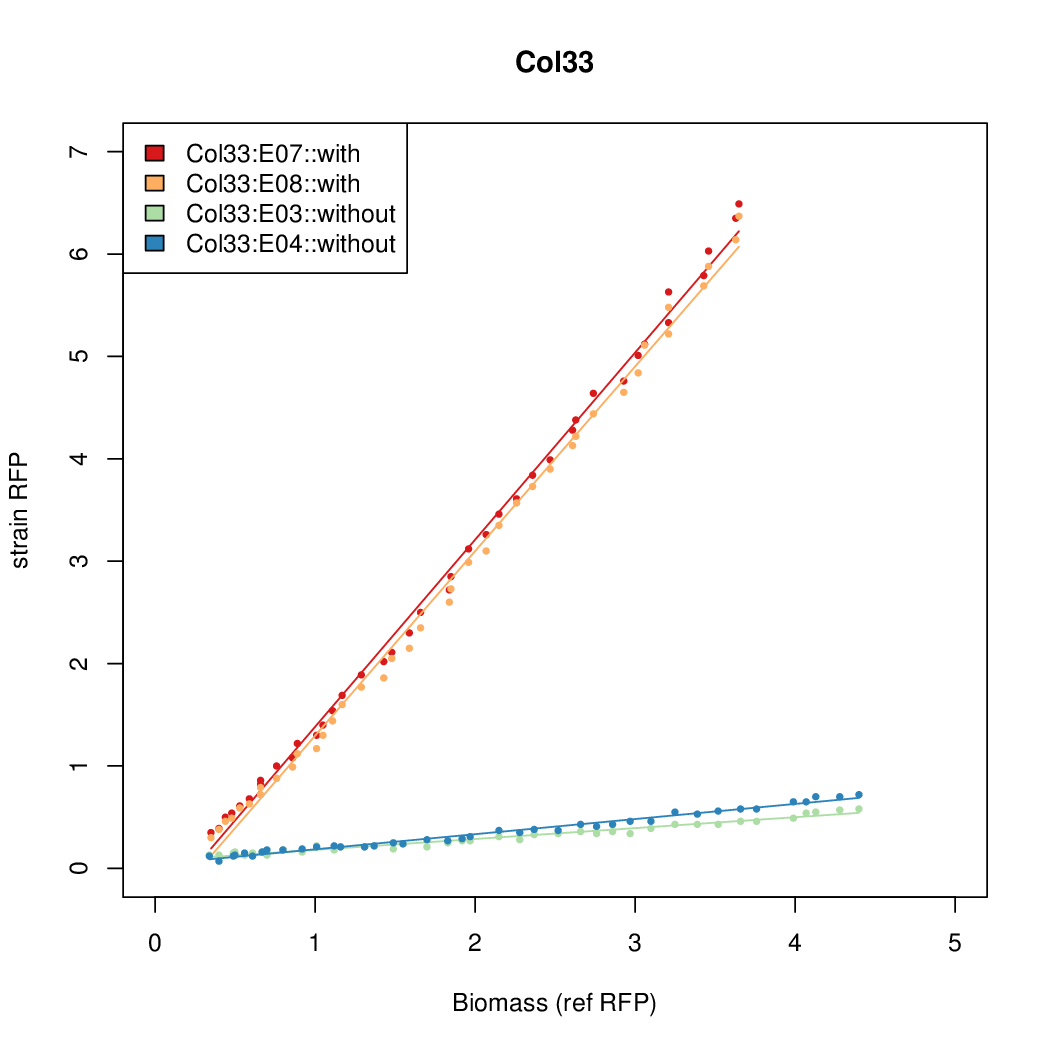
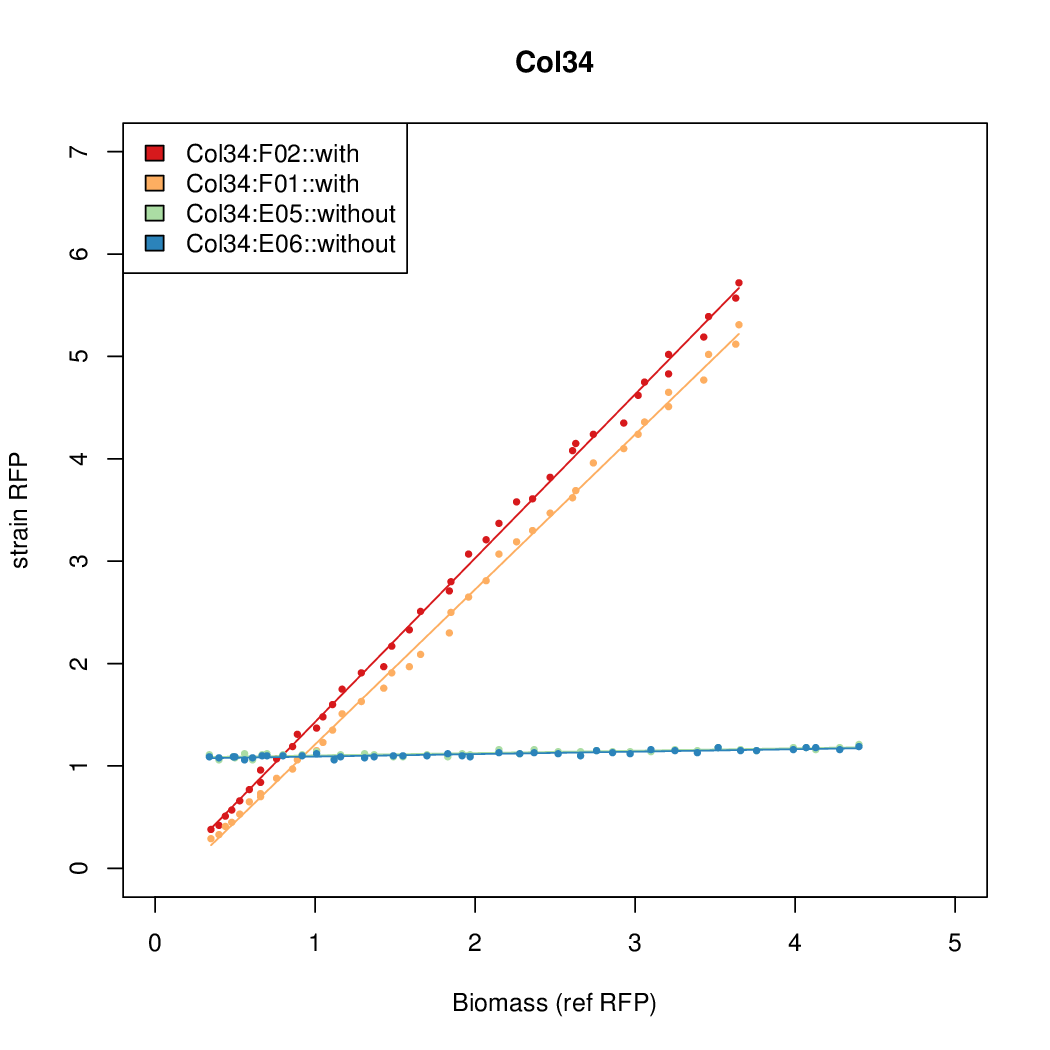
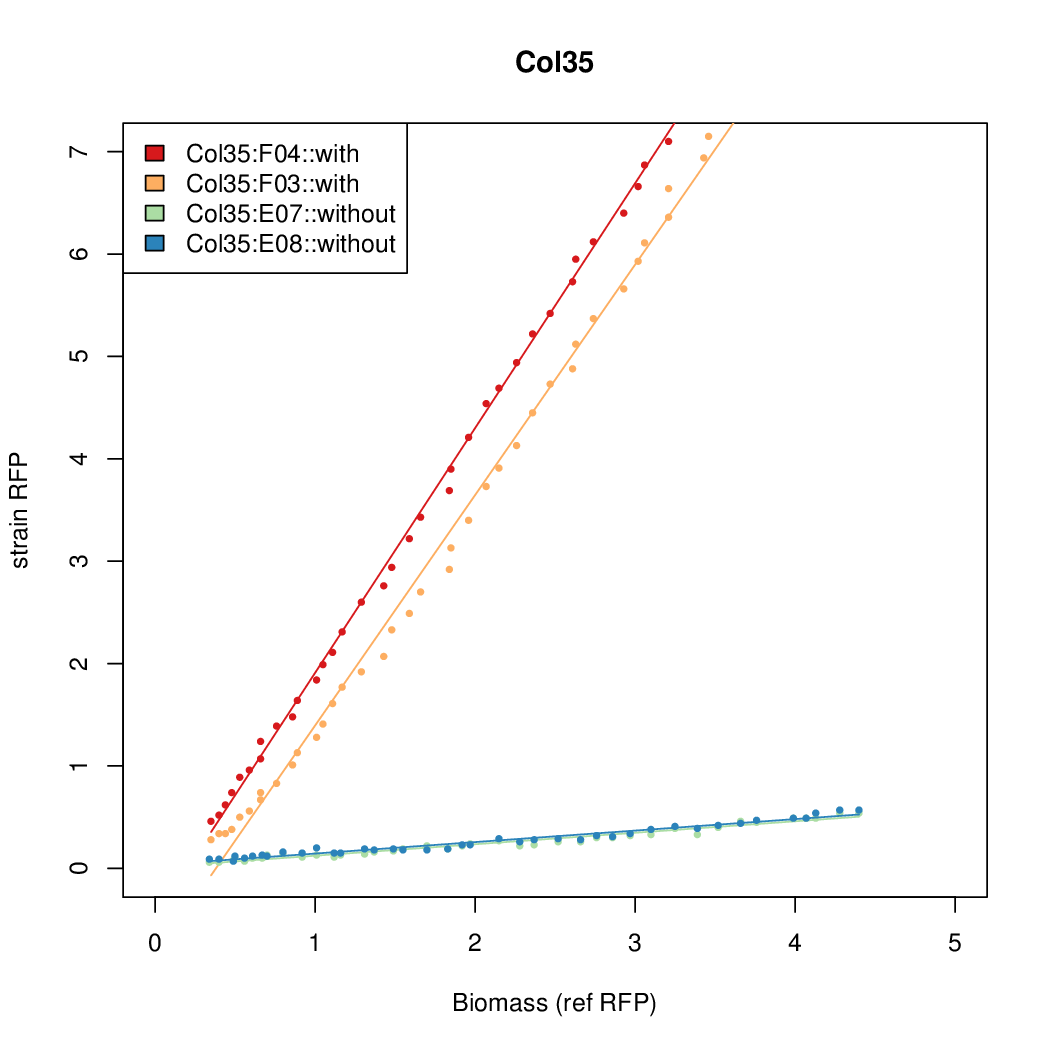
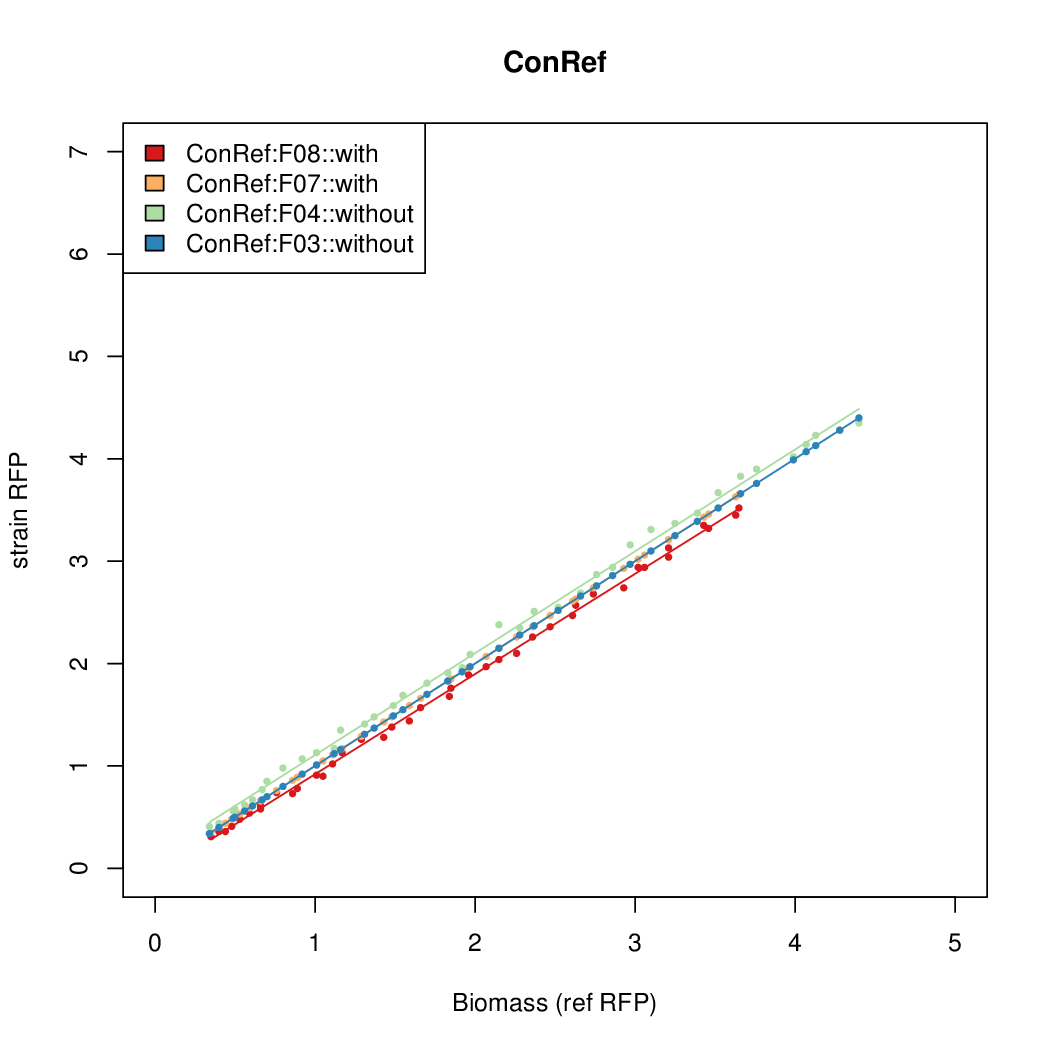
Promoter strengths for two trials of each colony with and without arabinose.
| Colony Number | With arabinose 1 | With arabinose 2 | Without arabinose 1 | Without arabinose 2 |
|---|---|---|---|---|
| Col2 | 8.9025 | 7.8699 | 0.727 | 0.9552 |
| Col3 | 9.2724 | 12.1142 | 0.5248 | 0.6982 |
| Col4 | 9.4571 | 11.4522 | 0.5231 | 0.2508 |
| Col12 | 6.3641 | 10.5389 | 0.5897 | 0.6869 |
| Col10 | 7.9697 | 9.7949 | 0.3392 | 0.733 |
| Col9 | 7.8563 | 20.1094 | 0.6995 | 0.7432 |
| Col8 | 12.2318 | 15.4548 | 0.4203 | 0.4538 |
| Col13 | 11.0377 | 7.3343 | 0.482 | 0.4641 |
| Col15 | 15.6817 | 8.2707 | 0.8169 | 0.1343 |
| Col18 | 14.7916 | 15.5736 | 0.6674 | 0.6745 |
| Col19 | 14.2126 | 16.4898 | 0.4545 | 0.3566 |
| Col29 | 7.1853 | 16.3467 | 0.5445 | 0.5013 |
| Col26 | 9.7724 | 9.6269 | 0.7118 | 0.7865 |
| Col22 | 8.4168 | 5.5958 | 0.6049 | 0.5645 |
| Col33 | 9.1982 | 8.9987 | 0.6508 | 1.374 |
| Col34 | 10.6987 | 7.883 | 0.5067 | 0.5031 |
| Col35 | 13.8427 | 7.5469 | 0.4363 | 0.6281 |
| ConRef | 6.506 | 7.9323 | 8.7811 | 7.9323 |
Example of use
See also "Hello World project".
 "
"