Team:Heidelberg/Tyrocidine week16 ms
From 2013.igem.org
(Created page with " == Validation of constructs by sequencing == * '''The sequences of for our NRPSs arrived: The alignments for the [[Media:Di TriI TriII TetII.zip|Di-, TriI-, TriII- and TetII -...") |
|||
| Line 1: | Line 1: | ||
| - | |||
== Validation of constructs by sequencing == | == Validation of constructs by sequencing == | ||
| Line 18: | Line 17: | ||
=== Protocol === | === Protocol === | ||
| - | [[File: | + | [[File:Heidelberg_Dipeptide 1&3B restr EcoRI PstI.jpg|right|100px|thumb|restriction with EcoRI and PstI]] |
{| class="wikitable" | {| class="wikitable" | ||
|- | |- | ||
| Line 152: | Line 151: | ||
<gallery> | <gallery> | ||
| - | File: | + | File:Heidelberg_Restriction digest NotI Di TriII TetI.png|restriction digest with NotI. Lanes 1 to 8: Dipeptide-NRPS, lanes 9 to 15: Tripeptide II-NRPS, lanes 16 to 23: Tetrapeptide-I-NRPS. |
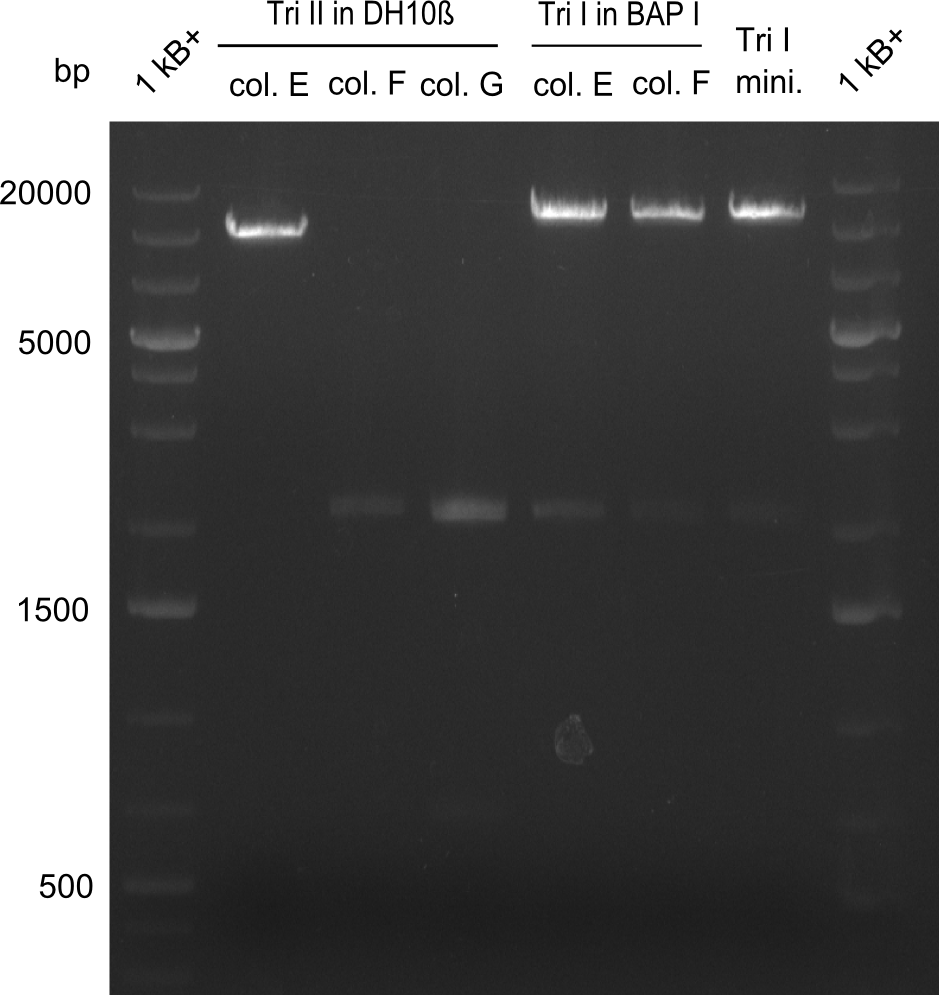
| - | File: | + | File:Heidelberg_Tri II in DH10 and Tri I in BAPI restriction.png|Restriction digest with NotI. Lanes 1 to 3: Tripeptide-II-NRPS |
| - | File: | + | File:Heidelberg_pPW02 DH10β ACD.png|Restriction digest with PstI & EcoRI. All lanes: Tetrapeptide-II-NRPS |
| - | File: | + | File:Heidelberg_pPW02 DH10β B.png|Restriction digest with PstI & EcoRI. All lanes: Tetrapeptide-II-NRPS |
</gallery> | </gallery> | ||
| Line 181: | Line 180: | ||
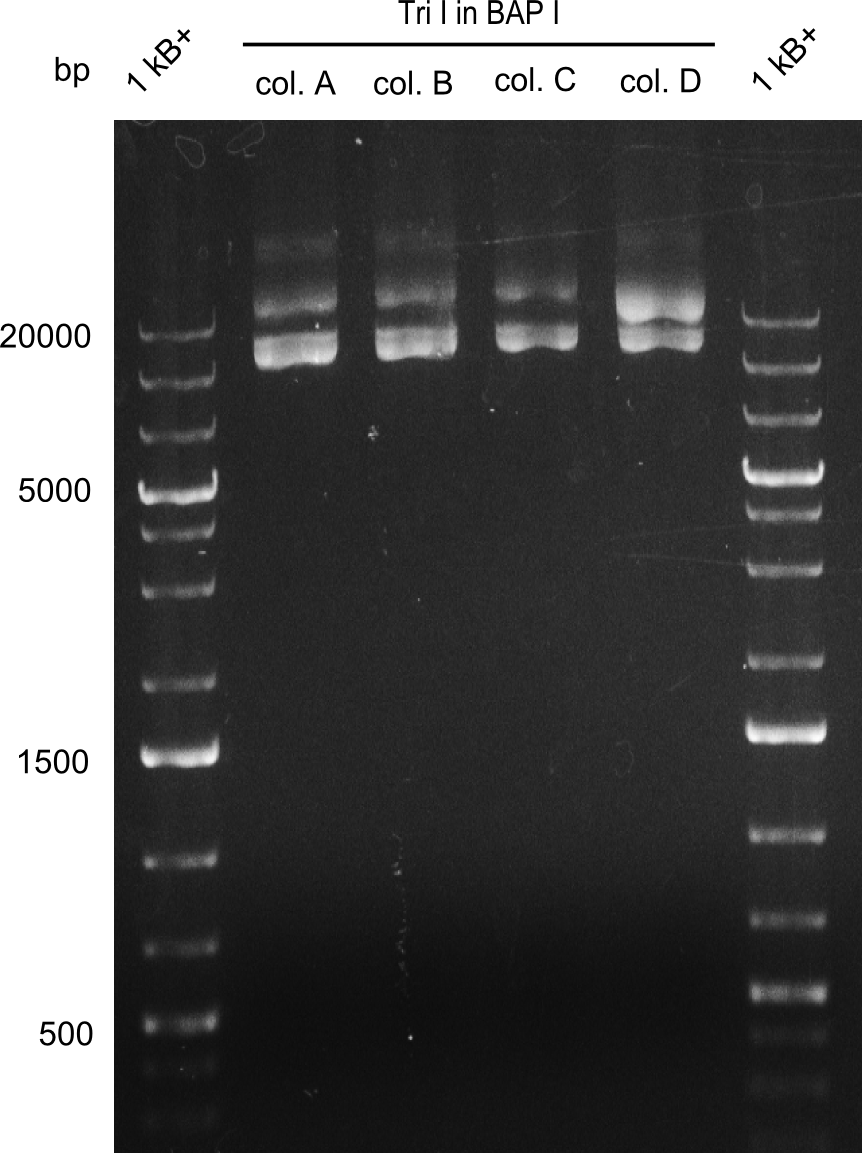
| - | [[File: | + | [[File:Heidelberg_Restriction NotI TriI in BAPI.png|300px]] |
| - | [[File: | + | [[File:Heidelberg_Tri II in DH10 and Tri I in BAPI restriction.png|300px]] |
==== Results ==== | ==== Results ==== | ||
| Line 191: | Line 190: | ||
===Dipeptide === | ===Dipeptide === | ||
| - | [[File: | + | [[File:Heidelberg_DiinBAP_neg_TetIinBAP_20130818.png|300px]] |
| - | [[File: | + | [[File:Heidelberg_DiinBAP_TetIinBAP_pos_20130819.png|300px]] |
==== Results ==== | ==== Results ==== | ||
| Line 200: | Line 199: | ||
===Tetrapeptide I === | ===Tetrapeptide I === | ||
| - | [[File: | + | [[File:Heidelberg_DiinBAP_neg_TetIinBAP_20130818.png|300px]] |
| - | [[File: | + | [[File:Heidelberg_DiinBAP_TetIinBAP_pos_20130819.png|300px]] |
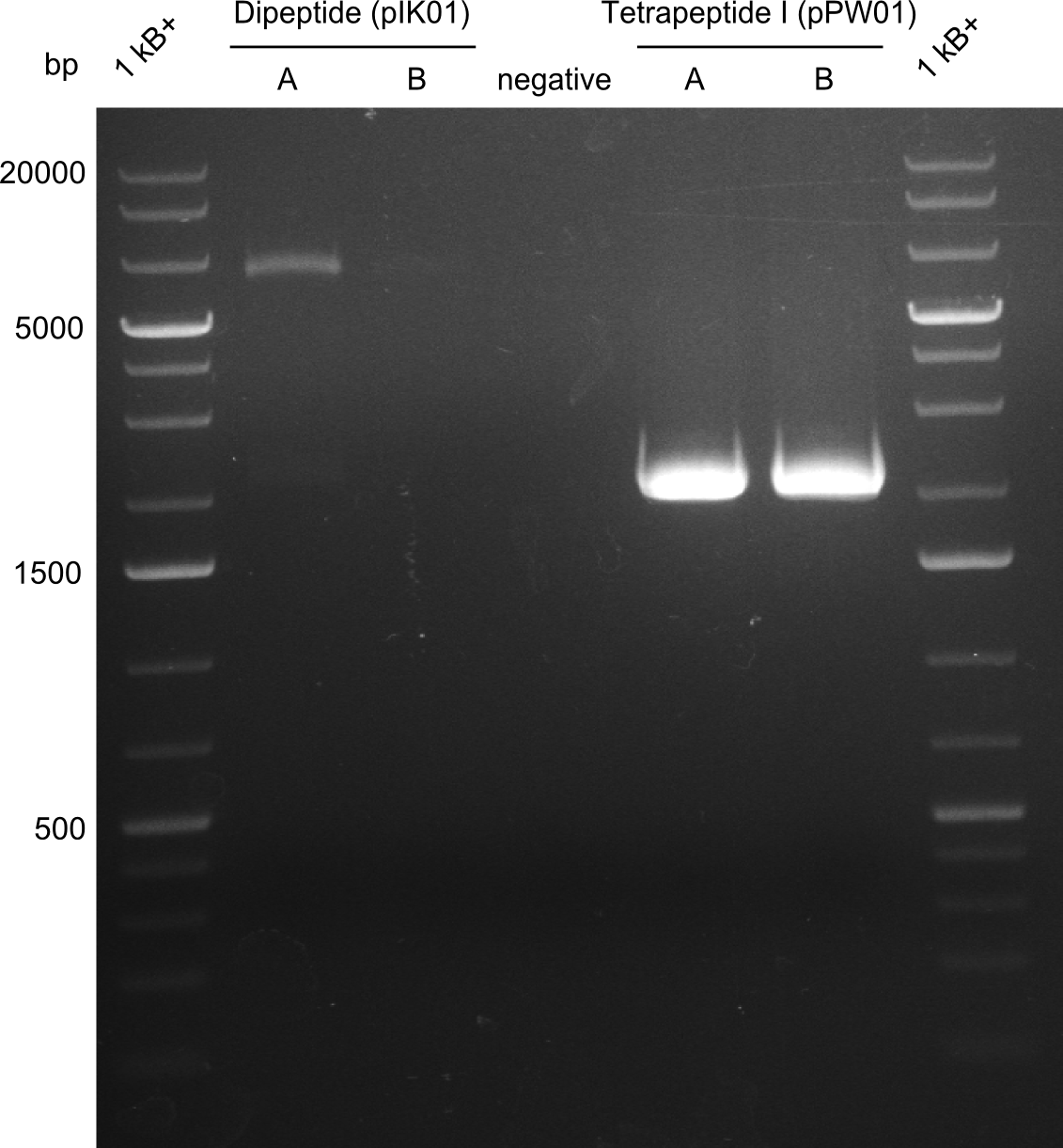
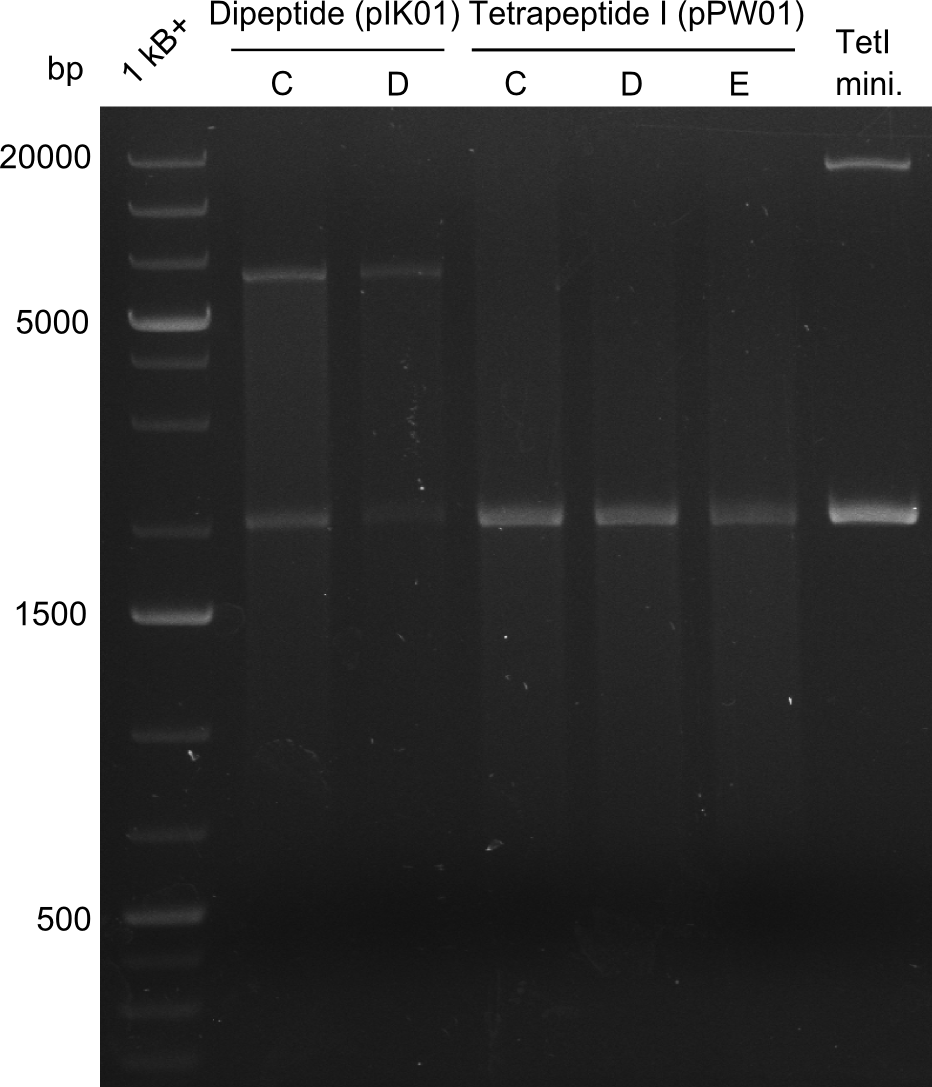
After a first, negative PstI-EcoRI-restriction digest from minipreps of BAPI-cells that were transformed with the validated plasmid for the Tetrapeptide-I-NRPS (figure XY), further colonies from the same plate were picked and tested, following the same procedure. This time the plasmid that was validated by sequencing and used for transformation was added as a positive contol (figure XY). However, the expected bands were not visible on the gel. Hence, the transformation was done once again, following the protocol described in the section above. | After a first, negative PstI-EcoRI-restriction digest from minipreps of BAPI-cells that were transformed with the validated plasmid for the Tetrapeptide-I-NRPS (figure XY), further colonies from the same plate were picked and tested, following the same procedure. This time the plasmid that was validated by sequencing and used for transformation was added as a positive contol (figure XY). However, the expected bands were not visible on the gel. Hence, the transformation was done once again, following the protocol described in the section above. | ||
Latest revision as of 16:28, 4 October 2013
Contents |
Validation of constructs by sequencing
- The sequences of for our NRPSs arrived: The alignments for the Di-, TriI-, TriII- and TetII - constructs and the new TriI - constructs.
- We also have positive Alignments, you can find them here
- So far, we have positive alignments for:
- Tripeptide I
- Tetrapeptide II
- one direction for Tripeptide II (the other direction - with IK13 - is not yet finished)
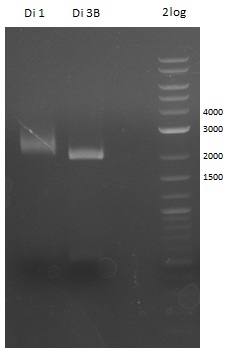
Restriction Digest of Dipeptide-NRPS from Minipreps
Enzymes used
- PstI
- EcoRI
- expected fragments: ~6000 & ~2000 bp
Protocol
| what | µl |
|---|---|
| DNA | 6 |
| Cutsmart Mix | 2 |
| EcoRI | 0.75 |
| PstI | 0.75 |
| ddH20 | 10.5 |
Results
- the obtained band does not match the expected size
- => optimize Gibson Assembly for Dipeptide
Gibson Assembly of missing short-peptide NRPS
Protocols
- According to the problems detected by sequencing, the following changes were performed
- Dipeptide: lower backbone-to-insert-ratio (less backbone used), as most cells were transformed with religated backbone-constructs
- Tripeptide II: lower backbone-to-insert-ratio, as religations or false ligations occured
- Tetrapeptide I: fragment 12 was given in excess, as it was missing in the sequenced plasmid
- The protocols used were the following:
Dipeptide
| fragment | concentration [ng/µl] | volume for gibson assembly [µL] |
|---|---|---|
| 1 | 30 | 0.80 |
| 2 | 16 | 4.81 |
| 3 | 35 | 4.39 |
Tripeptide II
| fragment | concentration [ng/µl] | volume for gibson assembly [µL] |
|---|---|---|
| 4 | 32 | 0.38 |
| 7 | 18 | 1.08 |
| 8 | 8 | 8.54 |
Tetrapeptide I
| fragment | concentration [ng/µl] | volume for gibson assembly [µL] |
|---|---|---|
| 9 | 30 | 0.35 |
| 10 | 40 | 0.42 |
| 11 | 10 | 4.20 |
| 12 | 10 | 3.36 |
| 13 | 20 | 1.68 |
Tetrapeptide II
| fragment | concentration [ng/µl] | volume for gibson assembly [µL] |
|---|---|---|
| 9 | 30 | 0.25 |
| 10 | 40 | 0.61 |
| 11 | 10 | 6.09 |
| 14 | 40 | 0.61 |
| 15 | 20 | 2.44 |
Transformation
- DH10β cells were transformed with electroporation:
- Dipeptide: purified DNA (15µl Gibson Mix purified with Isopropanol and EtOH, eluted in 10µl water)
- Dipeptide: raw Gibson mix (5µl Gibson Mix diluted in 10µl water)
- Tripeptide II: purified DNA (15µl Gibson Mix purified with Isopropanol and EtOH, eluted in 10µl water)
- Tripeptide II: raw Gibson mix (5µl Gibson Mix diluted in 10µl water)
- Tetrapeptide I: purified DNA (15µl Gibson Mix purified with Isopropanol and EtOH, eluted in 10µl water)
- Tetrapeptide I: raw Gibson mix (5µl Gibson Mix diluted in 10µl water)
- Tetrapeptide II: 1 µl of 5 µl raw Gibson Mix diluted in 10 µl water
- Tetrapeptide II: 14 µl of 5 µl raw Gibson Mix diluted in 10 µl water
- negative control: 10 µl water
Results
- white colonies were picked and grown overnight in 2x YT-medium with Chloramphenicol
Verification of correct constructs
Procedure
- 2 ml of overnight culture were used for miniprep and obtained DNA was quantified on Nano-drop
- Restriction digests with Not I
| what | µl |
|---|---|
| DNA | 600ng, alternatively 4 µl |
| Cutsmart Mix | 2 |
| Enzyme | 0.5 |
| ddH20 | ad 20 µl |
- Sequencing of positive samples
Results
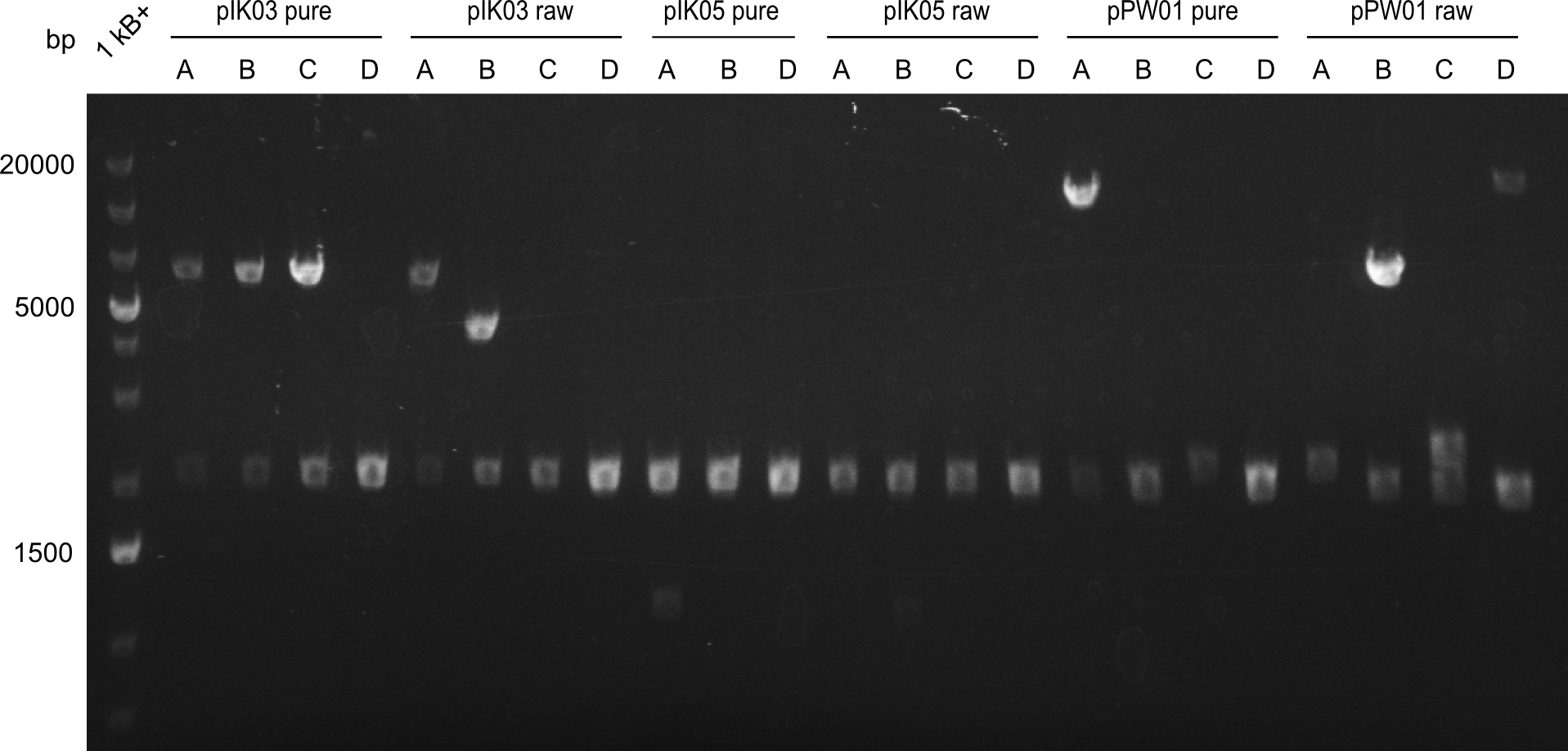
Heidelberg pPW02 DH10β ACD.png
Restriction digest with PstI & EcoRI. All lanes: Tetrapeptide-II-NRPS |
Heidelberg pPW02 DH10β B.png
Restriction digest with PstI & EcoRI. All lanes: Tetrapeptide-II-NRPS |
- 2 samples for the Dipeptide-NRPS (lane 3 and lane 5) and 2 samples for the Tetrapeptide I-construct (lane 16 and lane 23) were sent to sequencing.
- Positive alignments -> linkout
- All colonies that were transformed with Tripeptide II were negative, hence three additional colonies were picked
Tetrapeptide II was abandoned
Transformation of BAP I cells
Protocol
- thaw chemically competent BAP I cells on ice
- add ~50-100 ng of DNA
- let incubate for 20-30 minutes
- heat-shock at 42°C for 40 seconds
- let cool down on ice for 2 minutes
- add 1 ml 2x YT-medium
- let incubate for 60 minutes at 37°C
- centrifuge at 6000 rpm for 2:30 minutes
- decant medium and resuspend cells in remaining medium
- plate cells on agar plates with Chloramphenicol
Tripeptide I
Results
As the first digest with NotI (figure XY) did not give any positive results - in fact the bands lead to the conclusion that the digest did not work, as one can see the bands that are characteristic for coiled-coil, knicked and linearized plasmid - further colonies were picked and grown as an overnight-culture. Miniprep and restriction digest with NotI on the next day lead to a positive result (figure XY). In this case, the validated plasmid that was used for transformation was used as a positive control. We use Glycerol stocks of both BAP I cultures that were transformed with the NRPS for Tripeptide I for conservation.
Dipeptide
Results
A first restriction digest with PstI & EcoRI only lead to faint positive bands (figure XY), hence further colonies were picked and grown as an overnight culture. After miniprep and restriction digest - again with PstI & EcoRI - a clearer validation was obtained (figure XY). We created Glycerol Stocks of the cells containing the validated plasmid for the Dipeptide-NRPS.
Tetrapeptide I
After a first, negative PstI-EcoRI-restriction digest from minipreps of BAPI-cells that were transformed with the validated plasmid for the Tetrapeptide-I-NRPS (figure XY), further colonies from the same plate were picked and tested, following the same procedure. This time the plasmid that was validated by sequencing and used for transformation was added as a positive contol (figure XY). However, the expected bands were not visible on the gel. Hence, the transformation was done once again, following the protocol described in the section above.
 "
"