Team:Tsinghua-E/Parts
From 2013.igem.org
| Line 9: | Line 9: | ||
<div style="position:absolute;top:176px;left:56px;width:105px;height:186px;z-index:99;"> | <div style="position:absolute;top:176px;left:56px;width:105px;height:186px;z-index:99;"> | ||
<ul class="leftdh"> | <ul class="leftdh"> | ||
| - | <li><a href=" | + | <li><a href="#" id="partslink">Part1</a></li> |
| - | <li><a href=" | + | <li><a href="#" id="partslink">Part2</a></li> |
| - | <li><a href=" | + | <li><a href="#" id="partslink">Part3</a></li> |
| - | <li><a href=" | + | <li><a href="#" id="partslink">Part4</a></li> |
</ul> | </ul> | ||
</div> | </div> | ||
Revision as of 15:57, 22 September 2013



2013 iGEM Parts by Tsinghua-E
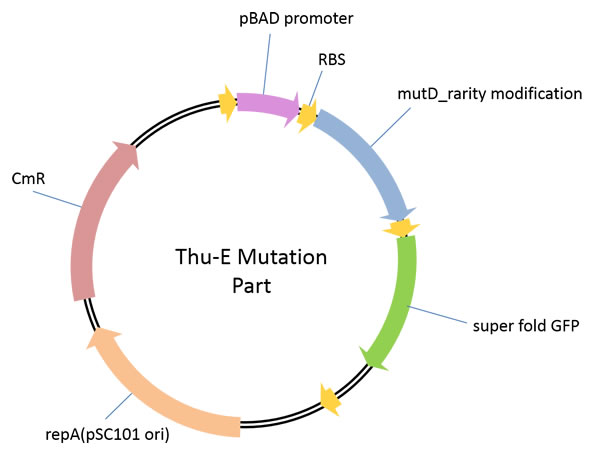
1、iGEM mut part ([http://parts.igem.org/wiki/index.php?title=Part:BBa_K1025003 BBa_K1025003])This part is used for genome level in vivo high-diversity library construction.In this vector, highly error-prone dnaQ mutant, mutD was cloned downstream of araBAD promoter so that we can control the mutation intensity of the target genome by the concentration of araBAD promoter’s inducer, L-arabinose in a strict manner. And the GFP downstream mutD was to test whether the mutD can be translated.
2、iGEM trp sensor performance test ([http://parts.igem.org/wiki/index.php?title=Part:BBa_K1025005 BBa_K1025005])
This part is used for performance test of our novel tryptophan sensor.The mechanism of this biosensor has been described in detail by Gong F. and Yanofsky C. It was derived from one regulation sequence upstream of tryptophanase(tnaA) operon in wild type E.coli. This sequence codes one 24-residue nascent peptide. Following this nascent peptide sequence stands one transcription termination factor (Rho) recognition sites. When certain amount of tryptophan exists, it is recognized by the nascent peptide. This leads to the hindering of TnaC-peptidyl-tRNAPro from being cleaved from ribosome. This peptide-mRNA-ribosome complex blocks Rho factor’s access to its binding site which is just adjacent to termination codon of nascent peptide so that initiate the transcription of downstream sequence. As far as we know, this novel mechanism has not been utilized before as tryptophan sensor. Hence, as a proof of principle, we cloned beta-lactamase gene lacZ downstream of wild type nascent peptide and Rho interaction sequence.The assembly was cloned between the NcoI and BamHI sites of pTrc99A vector.
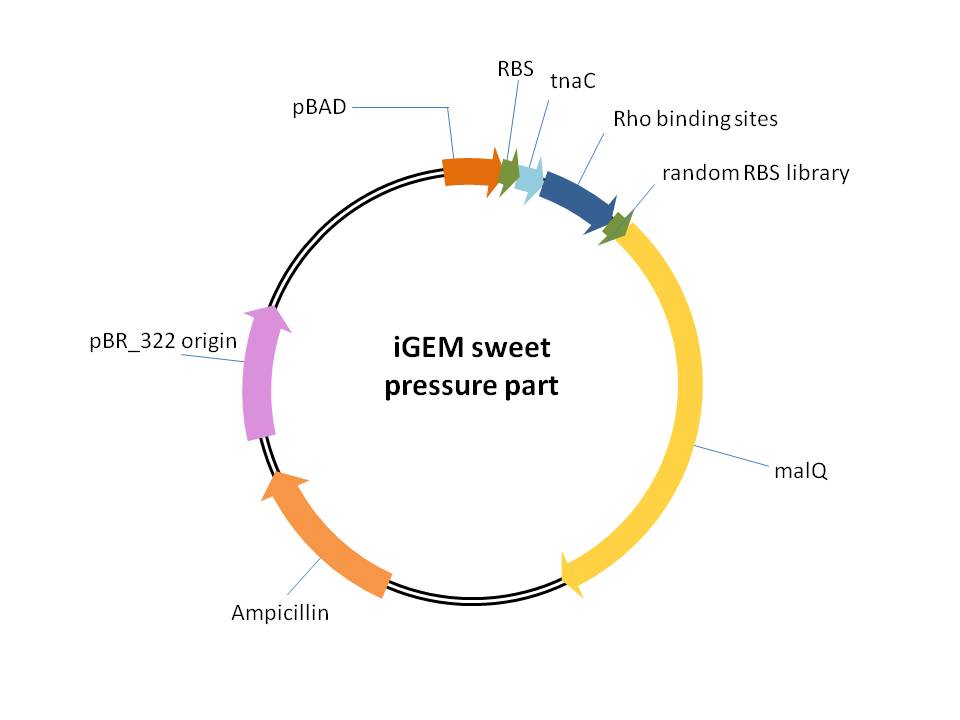
3、iGEM sweet pressure part([http://parts.igem.org/wiki/index.php?title=Part:BBa_K1025008 BBa_K1025008] and [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1025009 BBa_K1025009] )
This part is used for providing of selection pressure for the evolution and enrichment of tryptophan overproduction microorganism phenotype.This selection pressure was based on the tryptophan dependent maltose hydrolase expression which functioned in a maltose-sole carbon source culture condition. It is achieved by cloning E.coli maltose hydrolase gene (malQ) downstream of our previously constructed tryptophan biosensor which is controlled by the strict araBAD promoter in one malQ single deletion strain E.coli JW3379.
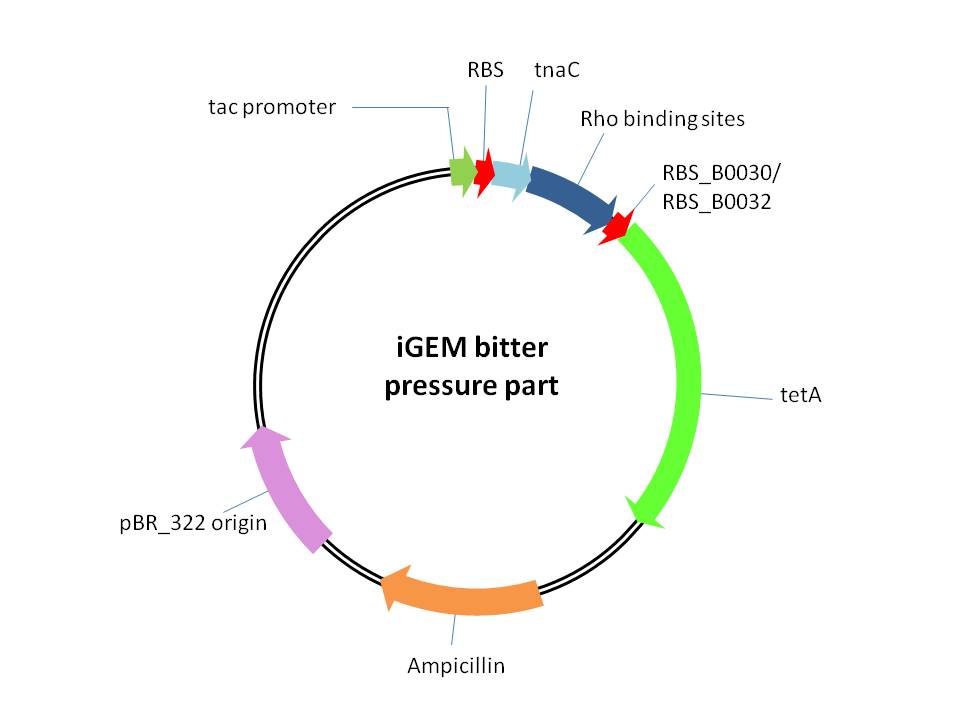
4、iGEM bitter pressure part([http://parts.igem.org/wiki/index.php?title=Part:BBa_K1025006 BBa_K1025006] and [http://parts.igem.org/wiki/index.php?title=Part:BBa_K1025007 BBa_K1025007] )
This part is used for providing of selection pressure for the evolution and enrichment of tryptophan overproduction microorganism phenotype.This selection pressure was based on the tryptophan dependent tetracycline antiporter expression which functioned in tetracycline culture condition. It is achieved by cloning E.coli tetracycline antiporter gene (tetA) downstream of our previously constructed tryptophan biosensor which is controlled by tac promoter between NcoI and BamHI restriction sites in pTrc99A vector. We constructed three bitter pressure part by utilizing three different RBS upstream of tetA gene.
Summary: <groupparts>iGEM013 Tsinghua-E</groupparts>
 "
"