Template:Kyoto/Notebook/Sep 26
From 2013.igem.org
(Difference between revisions)
(→Electrophoresis) |
(→Colony PCR) |
||
| Line 427: | Line 427: | ||
===Colony PCR=== | ===Colony PCR=== | ||
| - | |||
<div class="experiment"> | <div class="experiment"> | ||
<span class="author">Nakamoto Kozima</span> | <span class="author">Nakamoto Kozima</span> | ||
| Line 444: | Line 443: | ||
| 94°C|| 94°C|| 55°C|| 65°C||-- | | 94°C|| 94°C|| 55°C|| 65°C||-- | ||
|- | |- | ||
| - | |5min||30sec ||30sec ||54sec || | + | |5min||30sec ||30sec ||54sec ||30 |
|} | |} | ||
</div> | </div> | ||
| - | |||
===Electrophoresis=== | ===Electrophoresis=== | ||
Latest revision as of 18:26, 27 September 2013
Sep 26
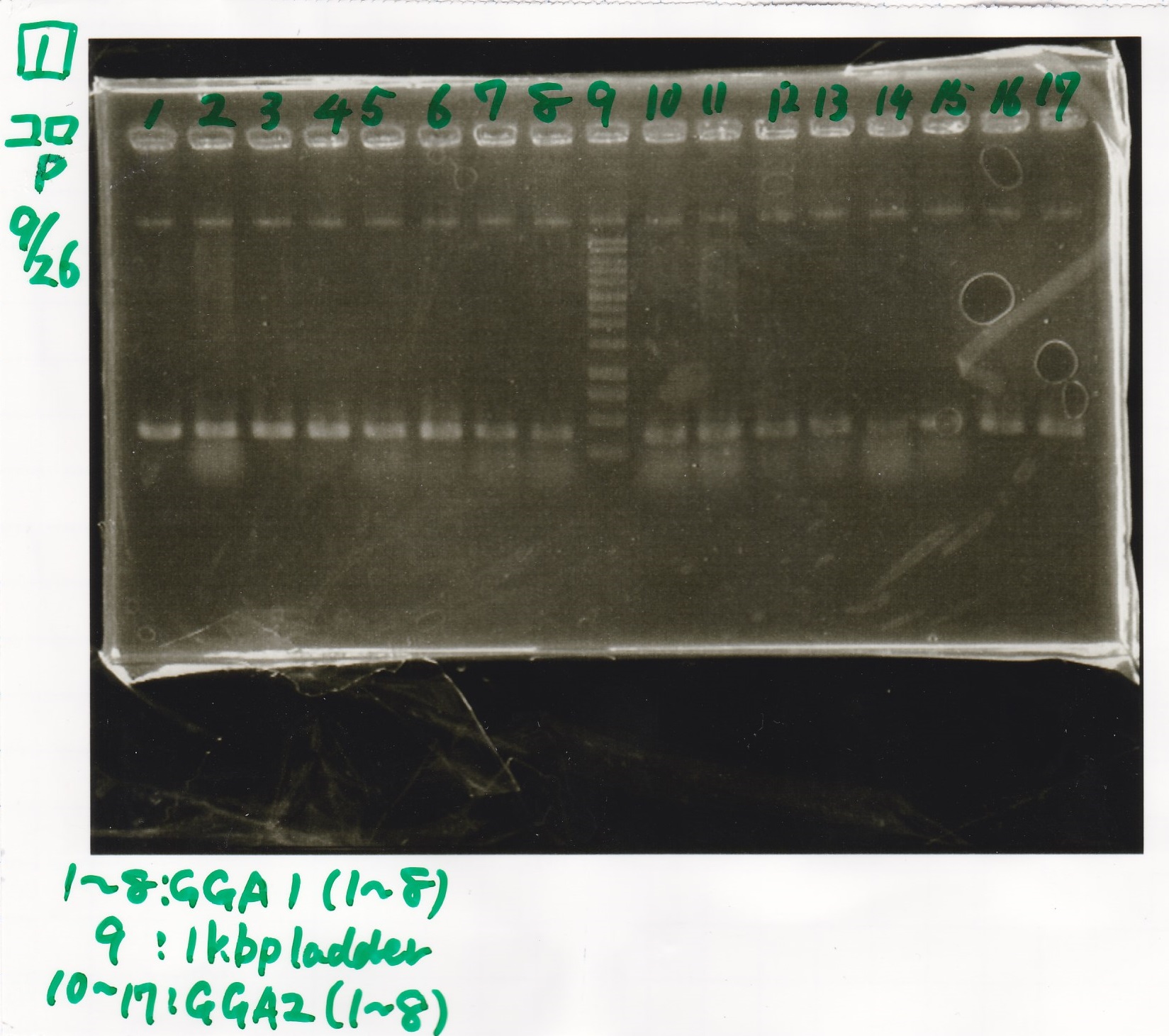
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA1-1 | -- | -- |
| 2 | GGA1-2 | -- | -- |
| 3 | GGA1-3 | -- | -- |
| 4 | GGA1-4 | -- | -- |
| 5 | GGA1-5 | -- | -- |
| 6 | GGA1-6 | -- | -- |
| 7 | GGA1-7 | -- | -- |
| 8 | GGA1-8 | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | GGA2-1 | -- | -- |
| 11 | GGA2-2 | -- | -- |
| 12 | GGA2-3 | -- | -- |
| 13 | GGA2-4 | -- | -- |
| 14 | GGA2-5 | -- | -- |
| 15 | GGA2-6 | -- | -- |
| 16 | GGA2-7 | -- | -- |
| 17 | GGA2-8 | -- | -- |
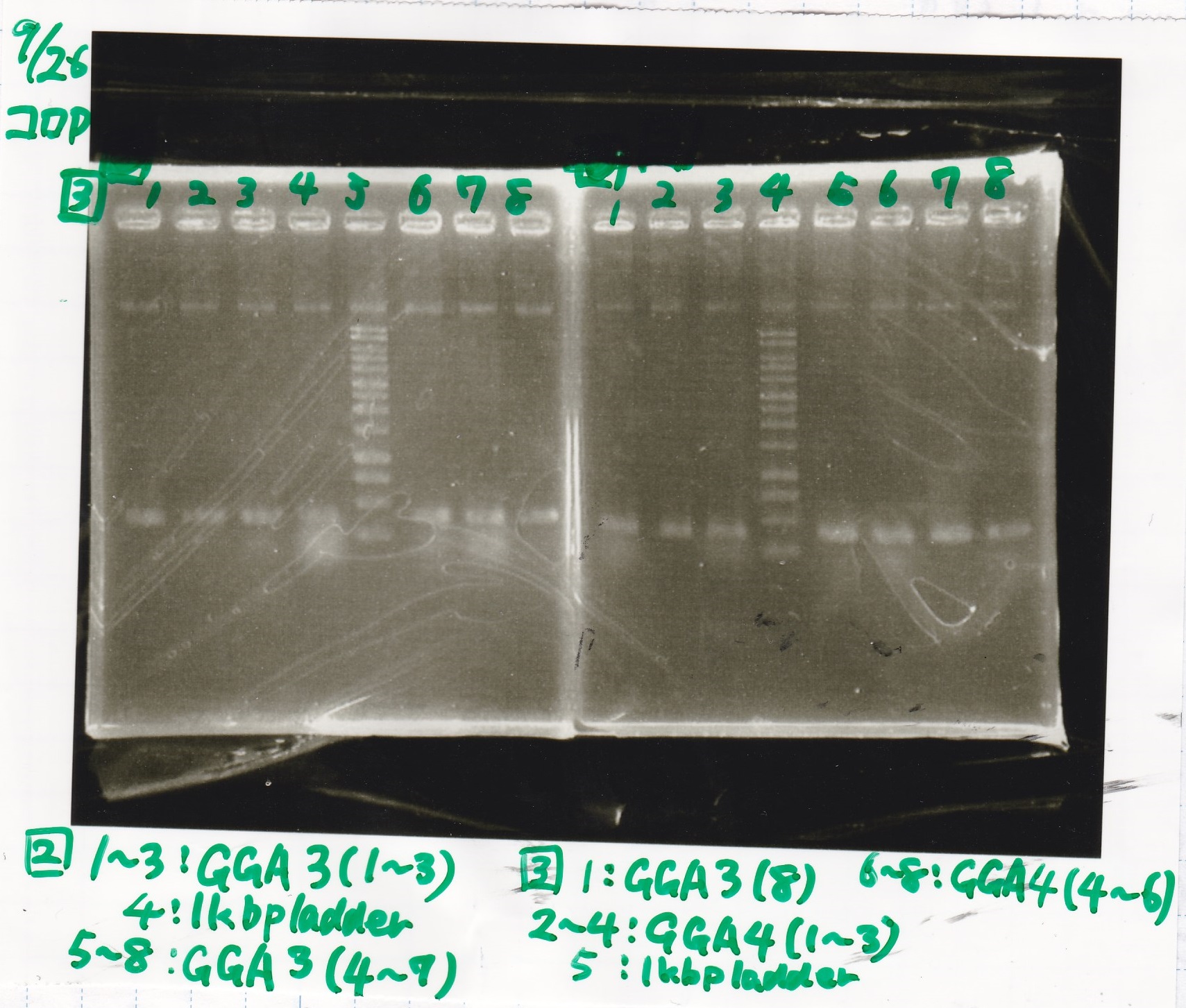
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA3-1 | -- | -- |
| 2 | GGA3-2 | -- | -- |
| 3 | GGA3-3 | -- | -- |
| 4 | 1kbp ladder | -- | -- |
| 5 | GGA3-4 | -- | -- |
| 6 | GGA3-5 | -- | -- |
| 7 | GGA3-6 | -- | -- |
| 8 | GGA3-7 | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA3-8 | -- | -- |
| 2 | GGA4-1 | -- | -- |
| 3 | GGA4-2 | -- | -- |
| 4 | GGA4-3 | -- | -- |
| 5 | 1kbp ladder | -- | -- |
| 6 | GGA4-4 | -- | -- |
| 7 | GGA4-5 | -- | -- |
| 8 | GGA4-6 | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA4-7 | -- | -- |
| 2 | GGA4-8 | -- | -- |
| 3 | 1kbp ladder | -- | -- |
| 4 | GGA7-1 | -- | -- |
| 5 | GGA16-1 | -- | -- |
| 6 | NC(RBS-GFP-DT) | -- | -- |
Ligasion(Golden Gate Assenbly)
| state | Vector | Inserter1 | Inserter2 | Inserter3 | Inserter4 | Inserter5 | (NEB)T4 ligase | (NEB)10*T4 ligase | MilliQ | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| experiment | RBS-GFP-DT | 1.0 µL | Pcon-pT181 antisense(E-2A) | 0.6µL | DT(2-1) | 0.5µL | Pcon-pT181 attenuator(1-SA) | 2.1 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | pSB1C3(EcoRI&SpeI) | 1.8 µL | Pcon-pT181 antisense(E-2A) | 0.9 µL | DT(2-1) | 0.5 µL | Pcon-GFP-DT(1-SA) | 1.5 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Ptet(E-0) | 0.7µL | Pcon pT181 attenuator(0-1A) | 0.3µL | tet aptamer(1-3) | 0.3 µL | Pcon-tetR-DT(3-2A) | 1.1 µL | Ptet(2-5) | 0.3 µL | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Ptet(E-0) | 0.7 µL | Pcon-pT181 attenuator-DT(0-1A) | 1.0 µL | Pcon-tetR-DT(1-2A) | 1.0 µL | Ptet(2-5) | 0.3µL | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Pcon-pT181 attenuator-tetR-DT(E-1A) | 1.0 µL | Pcon antisense-spinach-DT(1-2A) | 0.7µL | Pcon-pT 181 attenuator(2-SA) | 2.1 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Pcon antisense(E-1A) | 0.5 µL | spinach-DT(1-2) | 1.0 µL | Pcon-GFP-DT(1-2) | 0.8 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | pSB1C3(EcoRI&SpeI) | 1.8 µL | Pcon-pT181 antisense(E-1A) | 0.8µL | spinach-DT(1-2) | 1.0µL | Pcon-GFP-DT(2-SA) | 0.8 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | pSB1C3(EcoRI&SpeI) | 1.8 µL | Pcon-tet aptamer-DT(E-1A) | 3.0 µL | Ptet-pT181 antisense(1-3) | 1.5 µL | spinach-DT(3-2) | 1.5 µL | Pcon-GFP-DT(2-5A) | 0.8 µL | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-GFP-DT | Amp |
| RBS-GFP-DT | CP |
| Pcon-pT181 attenuator-DT | Amp |
| Pcon-tet aptamer-DT | Amp |
| Pcon-spinach-DT | Amp |
| Pcon-pT181 antisense-spinach-DT | Amp |
| spinach-DT | CP |
BSAI Digestion
| DNA | MilliQ | BSAI-HF | 10*cut smart | total | |
|---|---|---|---|---|---|
| Pcon | 7 µL | 10.7 µL | 0.3 µL | 2 µL | 20 µL |
| RBS-luxR-DT | 10 µL | 7 µL | 0.3 µL | 2 µL | 20 µL |
Colony PCR
| Sample | base pair |
|---|---|
| GGA-1 (9~16) | 3177 |
| GGA-2 (9~16) | 3189 |
| GGA-3 (9~21) | 2989 |
| GGA-4 (9~16) | 2939 |
| GGA-6 (1) | 3359 |
| GGA-16(2~4) | 2939 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 3min30s | 30cycles |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-1 (9) | -- | -- |
| 2 | GGA-1 (10) | -- | -- |
| 3 | GGA-1 (11) | -- | -- |
| 4 | GGA-1 (12) | -- | -- |
| 5 | GGA-1 (13) | -- | -- |
| 6 | GGA-1 (14) | -- | -- |
| 7 | GGA-1 (15) | -- | -- |
| 8 | GGA-1 (16) | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | GGA-2 (9) | -- | -- |
| 11 | GGA-2 (10) | -- | -- |
| 12 | GGA-2 (11) | -- | -- |
| 13 | GGA-2 (12) | -- | -- |
| 14 | GGA-2 (13) | -- | -- |
| 15 | GGA-2 (14) | -- | -- |
| 16 | GGA-2 (15) | -- | -- |
| 17 | GGA-2 (16) | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-4 (9) | -- | -- |
| 2 | GGA-4 (10) | -- | -- |
| 3 | GGA-4 (11) | -- | -- |
| 4 | GGA-4 (12) | -- | -- |
| 5 | GGA-4 (13) | -- | -- |
| 6 | GGA-4 (14) | -- | -- |
| 7 | GGA-4 (15) | -- | -- |
| 8 | GGA-4 (16) | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | GGA-6 (1) | -- | -- |
| 11 | GGA-16 (2) | -- | -- |
| 12 | GGA-16 (3) | -- | -- |
| 13 | GGA-16 (4) | -- | -- |
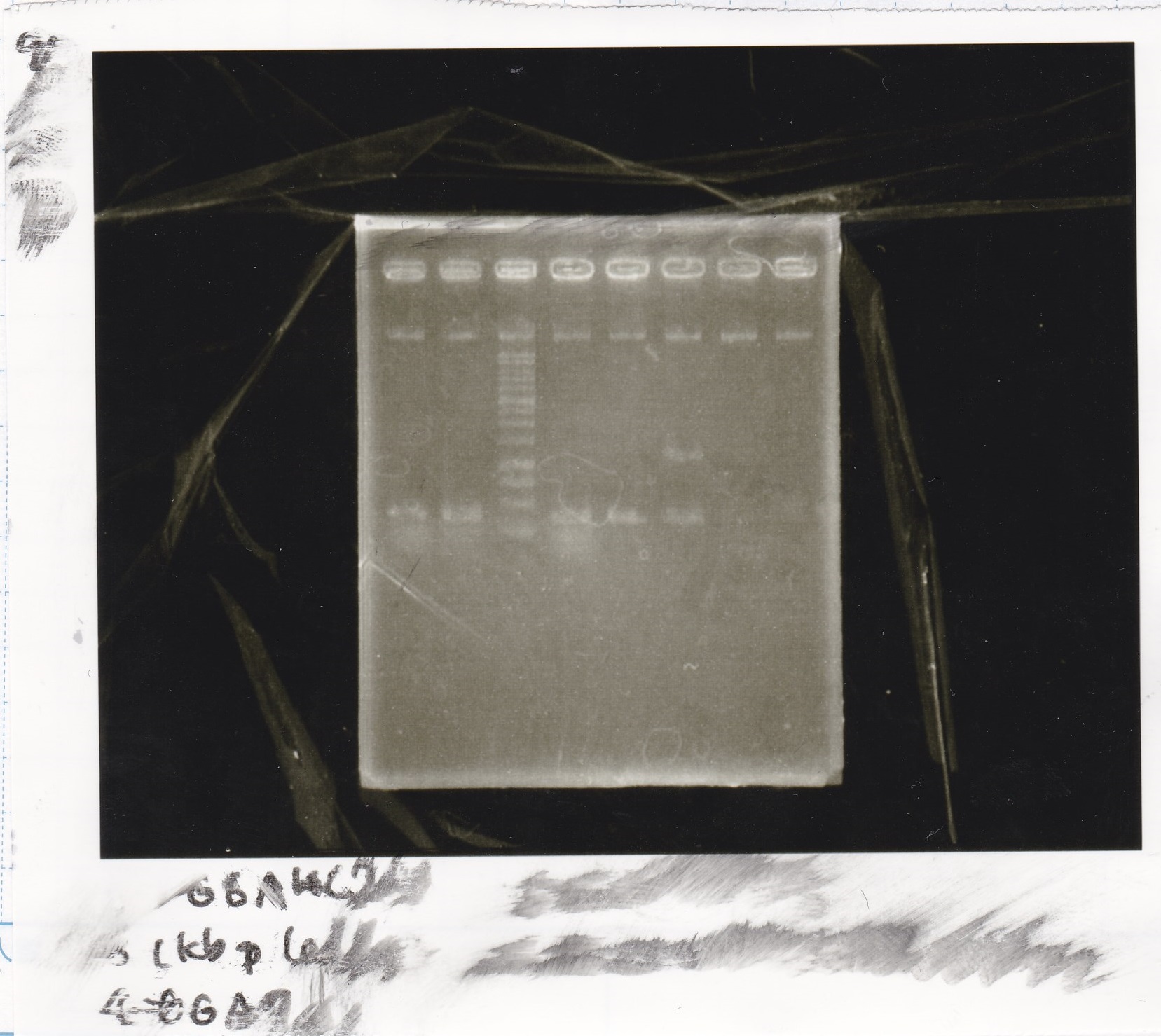
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-3 (9) | -- | -- |
| 2 | GGA-3 (10) | -- | -- |
| 3 | GGA-3 (11) | -- | -- |
| 4 | GGA-3 (12) | -- | -- |
| 5 | 1kbp ladder | -- | -- |
| 6 | GGA-3 (13) | -- | -- |
| 7 | GGA-3 (15) | -- | -- |
| 8 | GGA-3 (14) | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-3 (17) | -- | -- |
| 2 | GGA-3 (18) | -- | -- |
| 3 | GGA-3 (19) | -- | -- |
| 4 | 1kbp ladder | -- | -- |
| 5 | GGA-3 (20) | -- | -- |
| 6 | GGA-3 (21) | -- | -- |
| 7 | GGA-3 (16) | -- | -- |
Electrophoresis
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-GFP-DT | -- |
| RBS-GFP-DT | -- |
| Pcon-tet aptamer-DT | -- |
| Pcon-spinach-DT | -- |
| Pcon-pT181 antisense-spinach-DT | -- |
| spinach-DT | -- |
Colony PCR
| Sample | base pair |
|---|---|
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-1 | -- |
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-2 | -- |
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-3 | -- |
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-4 | -- |
| 9/24 Ptet-RBS-GFP-DT-1 | -- |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 1min40s | 30cycles |
Ligasion(Golden Gate Assenbly)
| state | Vector | Inserter1 | Inserter2 | Inserter3 | Inserter4 | Inserter5 |
|---|---|---|---|---|---|---|
| experiment | GGA2 | Pcon-attenuator(E-1A) | Pcon-tetR-DT(1-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) | |
| experiment | GGA3 | Pcon-tetap-DT(E-1A) | Pcon-tetR-DT(1-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) | |
| experiment | GGA4 | Pcon-spi-DT(E-1A) | Pcon-tetR-DT(1-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) | |
| experiment | GGA5 | Pcon-antisense(E-1A) | DT(1-3) | Pcon-tetR-DT(3-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) |
| experiment | GGA6 | Pcon-attenuator(E-1A) | tetap-DT(1-3) | Pcon-tetR-DT(3-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) |
| experiment | GGA7 | Pcon-tetR-DT(E-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 37°C | 16°C | 50°C | 80°C | -- |
| 3min | 4min | 5min | 5min | 25cycles |
RNA Extraction
| Sample |
|---|
| Pcon-Spinach-DT |
| Pcon-tetRaptamer-DT |
| Pcon-attenuator-DT |
| Spinach-DT |
| Pcon-antisense-Spinach-DT |
Liquid Culture
| Sample |
|---|
| Pcon-attemuater-DT |
| Pcon-aptamer-DT |
| Pcon-spinach-DT |
| Pcon-RBS-tetR-DT |
| Pcon-antisense-spinach-DT(Master 1) |
| Pcon-antisense |
Colony PCR
| Sample | base pair |
|---|---|
| 9/25 9/19pSB1C3&9/3PT181 attenuator(1~3) | 601bp |
| 9/25 aptamer12_1R-DT&9/24Pcon-pT181 attenuator(1~4) | 859bp |
| Pcon-attenuator-aptamer-DT(1) | 859bp |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 65°C | -- |
| 5min | 30sec | 30sec | 54sec | 30 |
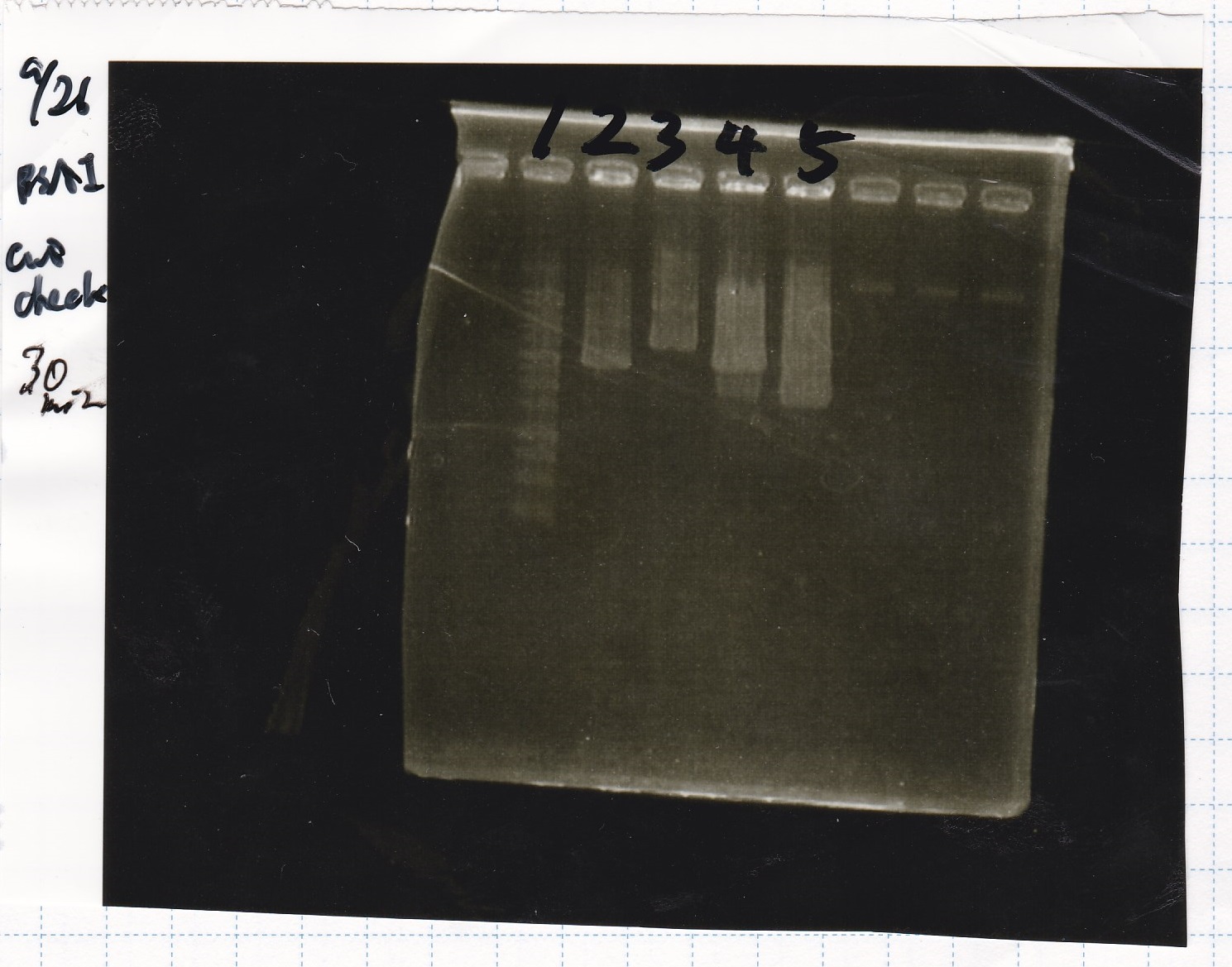
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kbp ladder | -- | -- |
| 2 | 9/25 9/23 1C3&9/3attenuator | -- | -- |
| 3 | 9/25 9/23 1C3&9/3attenuator | -- | -- |
| 4 | 9/25 9/23 1C3&9/3attenuator | -- | -- |
| 5 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
| 6 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
| 7 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
| 8 | 9/24 Pcon attenuator-aptamer-DT(1) | -- | -- |
| 9 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
Plating
| Sample | Use plate |
|---|---|
| 9/26 GGA2 | LB(CP) |
| 9/26 GGA3 | LB(CP) |
| 9/26 GGA4 | LB(CP) |
| 9/26 GGA5 | LB(CP) |
| 9/26 GGA6 | LB(CP) |
| 9/26 GGA7 | LB(CP) |
| 9/26 GGA11 | LB(CP) |
| 9/26 GGA312 | LB(CP) |
| 9/26 GGA13 | LB(CP) |
| 9/26 GGA14 | LB(CP) |
| 9/26 GGA15 | LB(CP) |
| 9/26 GGA20 | LB(CP) |
| 9/26 GGA23 | LB(CP) |
| 9/26 GGA24 | LB(CP) |
| 9/26 Ptet+RBS-GFP-DT | LB(CP) |
incubate 37°C
Restriction Enzyme Digestion
| 9/24 pSB1C3 | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 11µL | 1µL | 1µL | 3µL | 14µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 1µL | 8.5µL | 10µL |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 8.7µL | 1µL | 1µL | 3µL | 16.3µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 8.6µL | 10µL |
| 9/24 pT181attenuator | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 3.1µL | 1µL | 1µL | 3µL | 21.9µL | 30µL |
| NC | 0.2µL | 0µL | 0µL | 1µL | 8.8µL | 10µL |
| 8/17 RBS-GFP-DT | EcoRI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 1cut | 16.4µL | 1µL | 3µL | 3µL | 6.6µL | 30µL |
| NC | 0.4µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/17 pSB4K5 | EcoRI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 8.5µL | 1µL | 1µL | 3µL | 3µL | 13.5µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/14 pSB4K5 | EcoRI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 7.3µL | 1µL | 1µL | 3µL | 3µL | 14.7µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/22 Pcon-pT181attenuator-DT | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 5.4µL | 1µL | 1µL | 3µL | 19.6µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1µL | 8.7µL | 10µL |
| 8/7 RBS-GFP-DT 1 | XbaI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 8.1µL | 1µL | 1µL | 3µL | 3µL | 13.9µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/26 RBS-GFP-DT 1 | XbaI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 1cut | 8.1µL | 1µL | 3µL | 3µL | 14.9µL | 30µL |
| 9/16 Pλ-luxI | EcoRI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 14.7µL | 1µL | 1µL | 3µL | 3µL | 7.3µL | 30µL |
| NC | 0.7µL | 0µL | 0µL | 1µL | 1µL | 7.3µL | 10µL |
| 9/21 Ptet | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 14.2µL | 1µL | 1µL | 3µL | 10.8µL | 30µL |
| NC | 0.7µL | 0µL | 0µL | 1µL | 8.3µL | 10µL |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT | XbaI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 8.7µL | 1µL | 1µL | 3µL | 3µL | 13.3µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
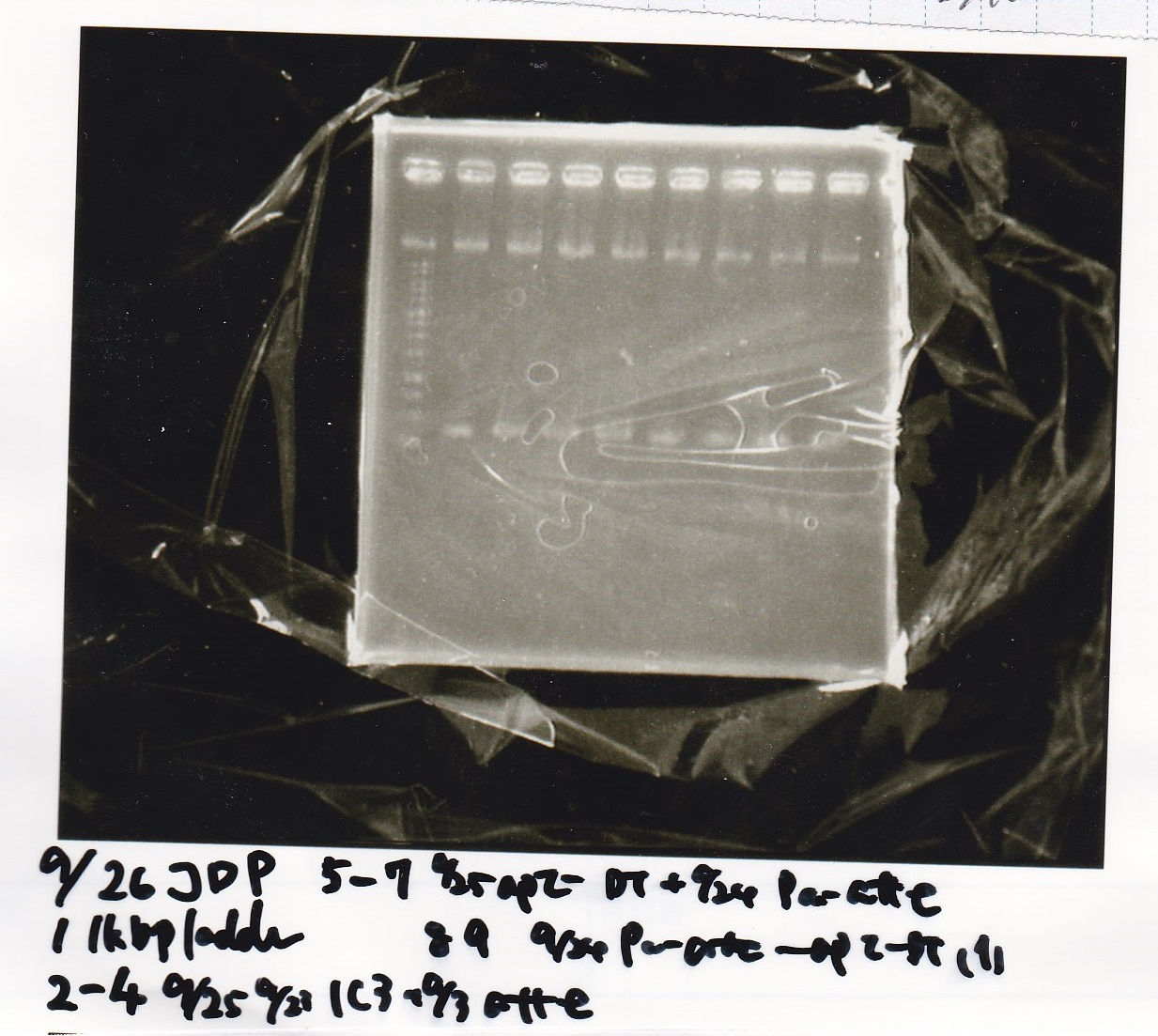
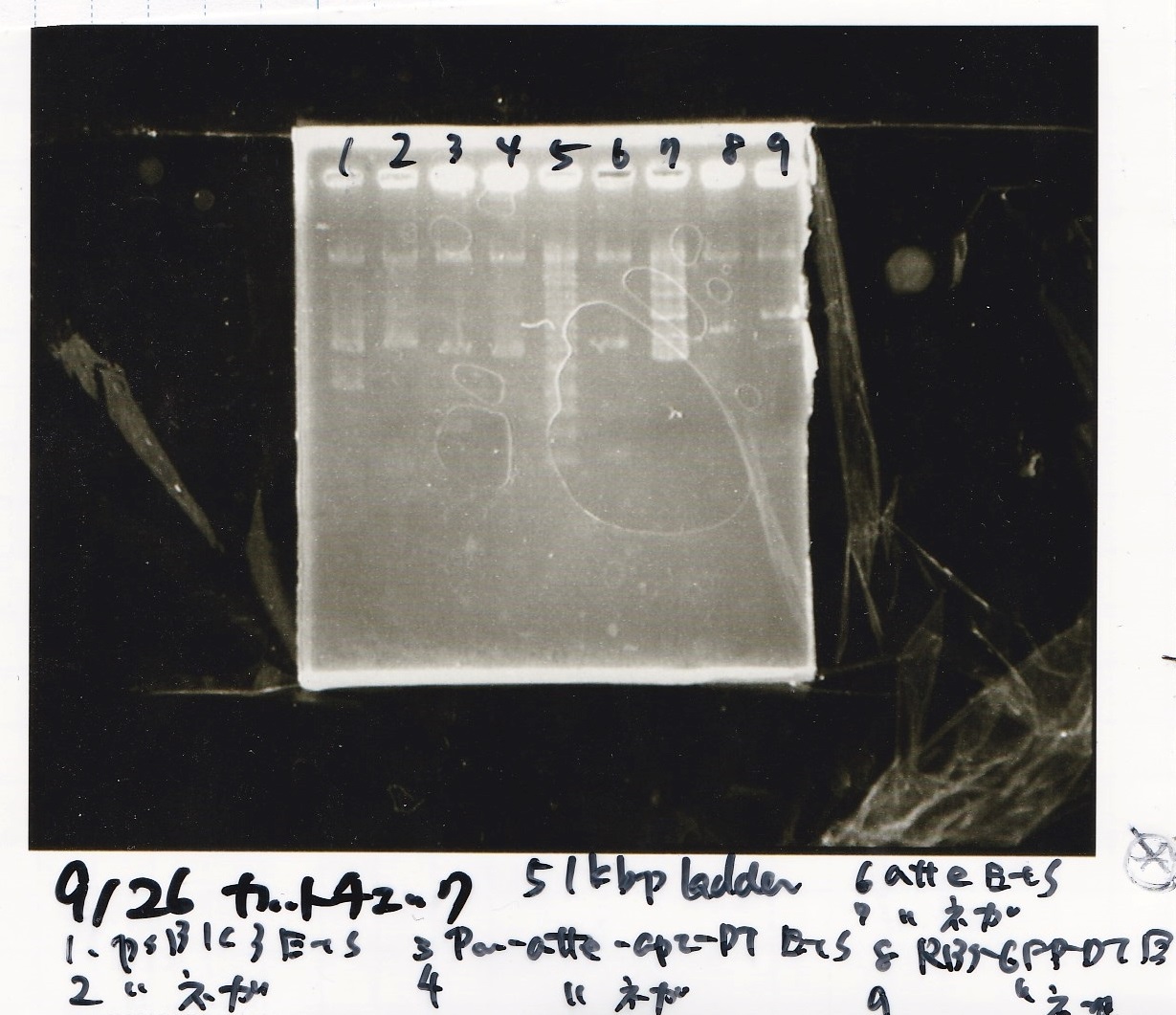
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | pSB1C3 | EcoRI | SpeI |
| 2 | pSB1C3 | -- | -- |
| 3 | Pcon-pT181attenuator-aptamer12-1R-DT | EcoRI | SpeI |
| 4 | Pcon-pT181attenuator-aptamer12-1R-DT | -- | -- |
| 5 | 1kb ladder | -- | -- |
| 6 | pT181attenuator | EcoRI | SpeI |
| 7 | pT181attenuator | -- | -- |
| 8 | RBS-GFP-DT | EcoRI | -- |
| 9 | RBS-GFP-DT | -- | -- |
 "
"