Team:Imperial College/mainresults
From 2013.igem.org
| Line 8: | Line 8: | ||
<div id="TabbedPanels1" class="TabbedPanels"> | <div id="TabbedPanels1" class="TabbedPanels"> | ||
<ul class="TabbedPanelsTabGroup"> | <ul class="TabbedPanelsTabGroup"> | ||
| - | <li class="TabbedPanelsTab" tabindex="0">< | + | <li class="TabbedPanelsTab" tabindex="0"><h5>Resource-full Waste</h5></li> |
| - | <li class="TabbedPanelsTab" tabindex="0">< | + | <li class="TabbedPanelsTab" tabindex="0"><h5>Plastic Fantastic</h5></li> |
</ul> | </ul> | ||
| Line 17: | Line 17: | ||
<div id="CollapsiblePanelM11" class="CollapsiblePanel"> | <div id="CollapsiblePanelM11" class="CollapsiblePanel"> | ||
| - | <div class="CollapsiblePanelTab" tabindex="0">< | + | <div class="CollapsiblePanelTab" tabindex="0"><h5>We made P3HB bioplastic </html><font size="1">▼</font size="1"><html></h5></div> |
<div class="CollapsiblePanelContent"> | <div class="CollapsiblePanelContent"> | ||
| Line 34: | Line 34: | ||
<div id="CollapsiblePanelM12" class="CollapsiblePanel"> | <div id="CollapsiblePanelM12" class="CollapsiblePanel"> | ||
| - | <div class="CollapsiblePanelTab" tabindex="0">< | + | <div class="CollapsiblePanelTab" tabindex="0"><h5>Modelling </html><font size="1">▼</font size="1"><html></h5></div> |
<div class="CollapsiblePanelContent"> | <div class="CollapsiblePanelContent"> | ||
Revision as of 17:01, 4 October 2013
Main Results
Resource-full Waste
Plastic Fantastic
We made P3HB bioplastic ▼
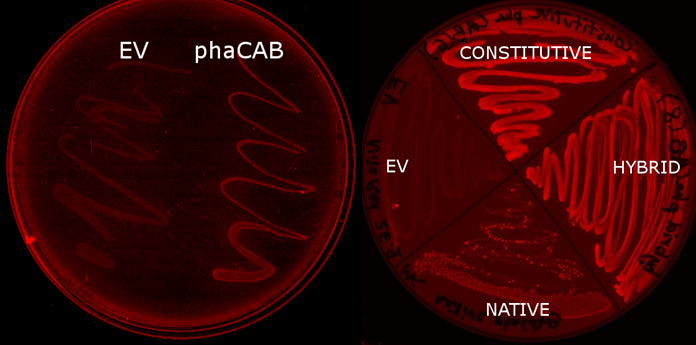
We used E. coli that expresses PhaCAB enzymes to make P3HB bioplastic. This is tested on nile red plates which stains P3HB.
Modelling ▼
Using model simulation results, we predict that the concentration of PhaCAB enzymes
We optimised bioplastic production ▼
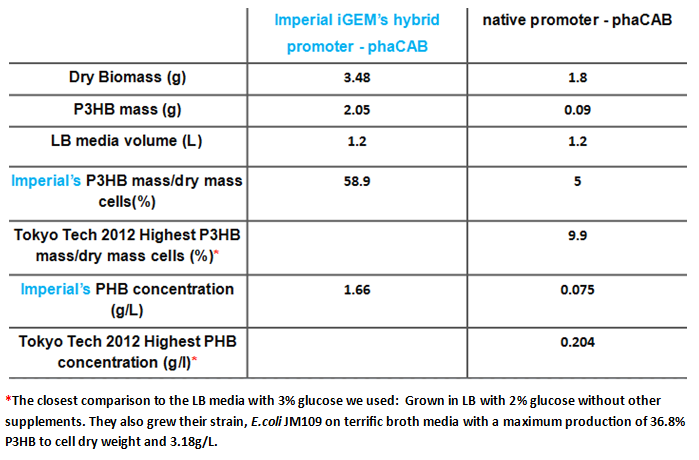
Since our model predicts that the concentration of PhaB enzymes is the rate limiting step in P3HB production, we designed a [http://parts.igem.org/Part:BBa_K1149051 hybrid promoter] consists of the J23104 constitutive promoter and the native promoter to optimise gene expression. Our results show that we have successfully produced 10-fold more P3HB bioplastic compared with the native promoter.
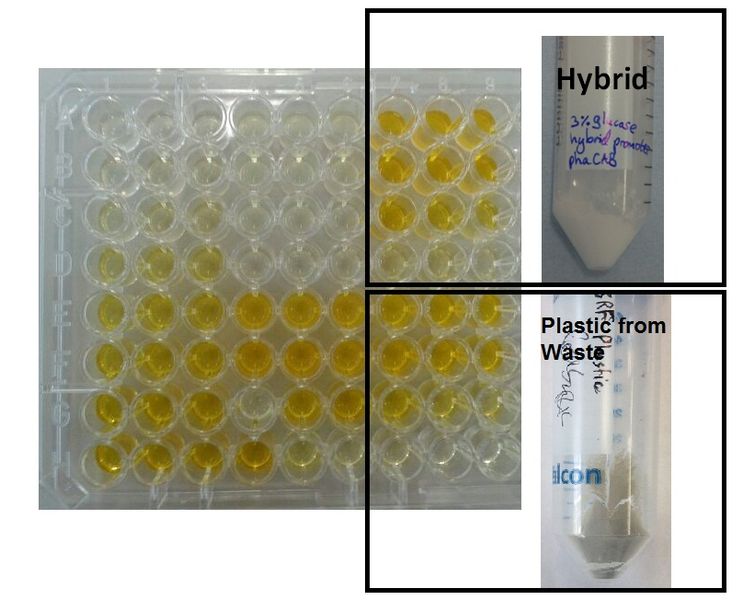
We made bioplastic from mixed waste ▼
One of the objectives of Module 1 is to produce P3HB bioplastic from waste. By comparing degradation product 3HB of P3HB bought from Sigma, produced from glucose and produced from the waste, we found there is no significant differences in 3HB concentration between these samples.

E. coli grows in P3HB and 3HB ▼
E. coli feeds on 3HB ▼
E. coli degrades P3HB ▼
Modelling P3HB degradation ▼
PLA degraded by E. coli ▼
 "
"