Team:Clemson/Notebook
From 2013.igem.org
Notebook
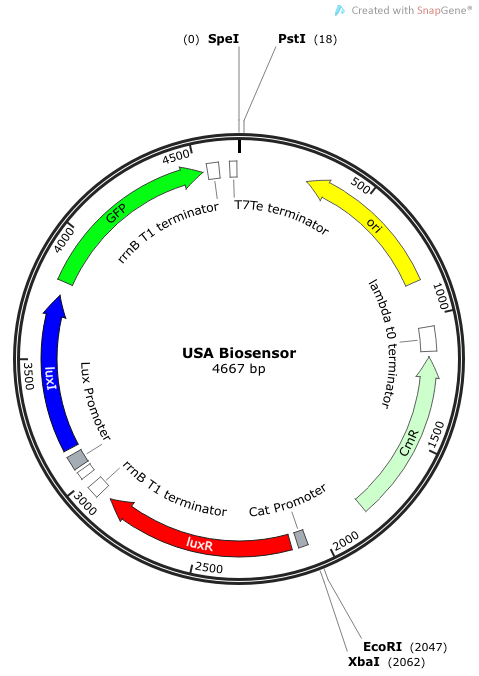
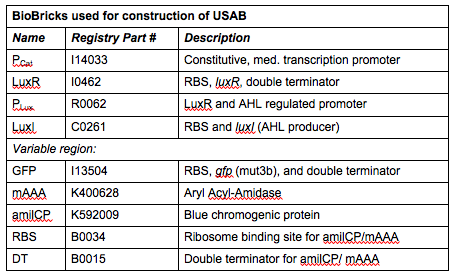
Rationale: I14033 was used because the medium transcription rate would be less energetically burdensome on the cell, and this seemed reasonable for a transcriptional regulator. The purpose of having I0462, R0062, and C0261 was to create a positive feedback loop. I0462 would be activated by an outside source of AHLs thereby activating R0062. R0062 would then drive transcription of C0261 to express LuxI, which is responsible for synthesizing more AHLs to drive I0462. This positive feedback loop is what makes the sensor self-amplifying, thus making a very minute signal from a pathogen become more detectable in smaller concentrations. Detection is made possible by GFP or the colorimetric proteins amilCP and mAAA.
Note: We also constructed prototype operons composed of the BioBrick parts R0010 (lac promoter) and I13504 (RBS-GFP-DT), or B0034-K400628-B0015 (mAAA version), or B0034-K592009-B0015 (amilCP version). We qualitatively determined that ligation of R0010 and I13504 and the subsequent transformation of R0010 + I13504 into E. coli was functional due to green florescence under UV light. We were never able to see blue color formation from amilCP or red color formation from mAAA. There may have been a mistake in our process for amilCP (e,g, missing RBS perhaps) to account for our results. See our “Experience” page for mAAA for more info on that.
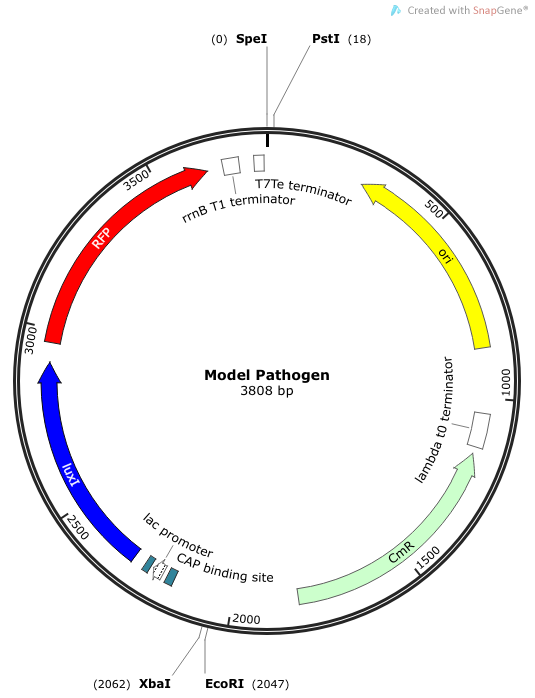
Phase II: Construction of “Model Pathogen”
We needed a bacterium that would produce an initial AHL signal to stimulate the USAB. Eventually a similar construct would be delivered via phage to specific pathogens, but for now we needed an easy way to test the self-amplifiying ability of our USAB.
 "
"