Template:Kyoto/Notebook/Sep 17
From 2013.igem.org
Sep 17
Sep 18
Restriction Enzyme Digestion
| 9/18Plux | SpeI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 3.2µL | 1µL | 1µL | 3µL | 3µL | 18.8µL | 30µL |
| NC | 1.7µL | 0µL | 0µL | 1µL | 1µL | 6.3µL | 10µL |
| 9/18Pcon-pT181antisense-spinach-DT② | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 7.5µL | 1µL | 1µL | 3µL | 3µL | 14.5µL | 30µL |
| NC | 1.4icro;L | 0µL | 0µL | 1µL | 1µL | 6.6µL | 10µL |
| 9/16 Plac② | SpeI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 13.8µL | 1µL | 1µL | 3µL | 3µL | 14.5µL | 30µL |
| NC | 0.7µL | 0µL | 0µL | 1µL | 1µL | 7.3µL | 10µL |
| 9/16 Spinach-DT | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 17.6µL | 1µL | 1µL | 3µL | 3µL | 4.4µL | 30µL |
| NC | 0.9µL | 0µL | 0µL | 1µL | 1µL | 7.1µL | 10µL |
| 9/8 Pcon J23100 | SpeI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 5.7µL | 1µL | 1µL | 3µL | 3µL | 16.3µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1µL | 1µL | 7.7µL | 10µL |
| 9/16 RBS-lacZα-DT | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 8.4µL | 1µL | 1µL | 3µL | 3µL | 13.6µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/11 Pcon-pT181antisense-spinach-DT①: | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 7.2µL | 1µL | 1µL | 3µL | 3µL | 14.8µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/6 apt12-1R-DT | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 7.9µL | 1µL | 1µL | 3µL | 3µL | 14.1µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/3 pT181antisense | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 6.3µL | 1µL | 1µL | 3µL | 3µL | 15.7µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1µL | 1µL | 7.7µL | 10µL |
| 9/14 pSB4K5 | EcoRI | SpeI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 7.3µL | 1µL | 1µL | 3µL | 3µL | 14.7µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 8/31 Plux-RBS-GFP-DT | EcoRI | SpeI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 9.2µL | 1µL | 1µL | 3µL | 3µL | 12.8µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 1µL | 1µL | 7.5µL | 10µL |
| 9/12 RBS-lysis3-DT | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 5.2µL | 1µL | 1µL | 3µL | 3µL | 16.8µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/18 Pcon-pT181attenuator | EcoRI | SpeI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 14.3µL | 1µL | 1µL | 3µL | 3µL | 7.7µL | 30µL |
| NC | 0.7µL | 0µL | 0µL | 1µL | 1µL | 7.3µL | 10µL |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | Plae② | SpeI | PstI |
| 2 | RBS-lysis3-DT | XbaI | PstI |
| 3 | Plux-RBS-GFP-DT | EcoRI | SpeI |
| 4 | PSB4K5 | EcoRI | SpeI |
| 5 | Pcon | SpeI | PstI |
| 6 | Plux | SpeI | PstI |
| 7 | 1kbp ladder | -- | -- |
| 8 | Plae② | -- | -- |
| 9 | RBS-lysis3-DT | -- | -- |
| 10 | Plux-RBS-GFP-DT | -- | -- |
| 11 | PSB4K5 | -- | -- |
| 12 | Pcon | -- | -- |
| 13 | Plux | -- | -- |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 100bp ladder | -- | -- |
| 2 | Pcon pT181antisense spinach-DT② | XbaI | PstI |
| 3 | Spinach-DT | XbaI | PstI |
| 4 | RBS-lacZα-DT | XbaI | PstI |
| 5 | Pcon pT181antisense-Spinach-DT① | XbaI | PstI |
| 6 | aptamer12-1R-DT | XbaI | PstI |
| 7 | pT181attenuator | XbaI | PstI |
| 8 | Pcon attenuator | EcoRI | SpeI |
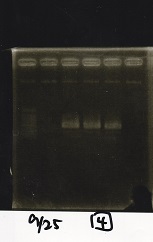
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 100bp ladder | -- | -- |
| 2 | Pcon pT181antisense spinach-DT② | XbaI | PstI |
| 3 | Spinach-DT | XbaI | PstI |
| 4 | RBS-lacZα-DT | XbaI | PstI |
| 5 | Pcon pT181antisense-Spinach-DT① | XbaI | PstI |
| 6 | apt12-1R-DT | XbaI | PstI |
| 7 | pT181attenuator | XbaI | PstI |
| 8 | Pcon attenuator | EcoRI | SpeI |
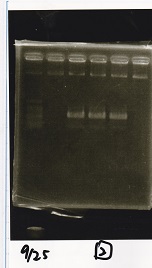
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 100bp ladder | -- | -- |
| 2 | Pcon pT181antisense spinach-DT② | -- | -- |
| 3 | Spinach-DT | -- | -- |
| 4 | RBS-lacZα-DT | -- | -- |
| 5 | Pcon pT181antisense-Spinach-DT① | -- | -- |
| 6 | apt12-1R-DT | -- | -- |
| 7 | pT181attenuator | -- | -- |
| 8 | Pcon attenuator | -- | -- |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kbp ldder | -- |
| 2 | Plac② | SpeI+PstI |
| 3 | Plac② | SpeI+PstI |
| 4 | -- | -- |
| 5 | RBS-lysis3-DT | XbaI+PstI |
| 6 | RBS-lysis3-DT | XbaI+PstI |
| 7 | -- | -- |
| 8 | Plux-RBS-GFP-DT | EcoRI+SpeI |
| 9 | Plux-RBS-GFP-DT | EcoRi+SpeI |
| 10 | -- | -- |
| 11 | PSB4KS | EcoRI+SpeI |
| 12 | PSB4KS | EcoRI+SpeI |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kbp ldder | -- |
| 2 | Pcon | SpeI+PstI |
| 3 | Pcon | SpeI+PstI |
| 4 | -- | -- |
| 5 | Plux | SpeI+PStI |
| 6 | Plux | SpeI+PStI |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | Pcon PT181 antisense spinach-DT | XbaI+PstI |
| 2 | Pcon PT181 antisense Spinach-DT | XbaI+PstI |
| 3 | 100bp ladder | -- |
| 4 | Pcon PT181 antisense Spinach-DT | XbaI+PstI |
| 5 | Pcon PT181 antisense Spinach-DT | XbaI+PstI |
| 6 | 1kbp ladder | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2 | apt12-1R-DT | XbaI+PstI |
| 3 | apt12-1R-DT | XbaI+PstI |
| 4 | -- | -- |
| 5 | PT181 attenuator | XbaI+PstI |
| 6 | PT181 attenuator | XbaI+PstI |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2 | Pcoa attenuator | EcoRI+PstI |
| 3 | Pcoa attenuator | EcoRI+PstI |
| 4 | -- | -- |
| 5 | RBS-lacZα-DT | XbaI+PstI |
| 6 | RBS-lacZα-DT | XbaI+PstI |
Sep 19
Sep 20
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/18 Plac (SpeI & PstI) | 3.0 µL | 9/18 RBS-lysis3-DT (XbaI & PstI) | 12.4 µL | 3.5 µL |
| experiment | 9/12 Ptet-pT181 antisense | 1.0 µL | 9/18 spinach-DT | 11.9 µL | 3.5 µL |
| experiment | 9/13 Pcon-RBS-luxR-DT | 2.1 µL | 9/10 Plux-RBS-GFP-DT | 5.6 µL | 3.5 µL |
| experiment | 9/13 Pcon-pT181 attenuator (SpeI & PstI) | 2.0 µL | 9/18 aptamer 12-1R-DT (XbaI & PstI) | 6.3 µL | 3.5 µL |
| experiment | 9/18 Plac ②(SpeI & PstI) | 3.0 µL | 9/18 aptamer 12-1R-DT(XbaI & PstI) | 4.5µL | x3.5 µL |
Transformation
| Name | Sample | Competent Cells | Plate |
|---|---|---|---|
| 9/17 pSB1C3-Pcon-pT181 attenuator-DT | 2 µL | 20 µL | CP |
| 9/17 pSB1C3-Pcon-aptamer 12_1R-DT | 2 µL | 20 µL | CP |
| 9/17 pSB1C3-Pcon-antisense-spinach-DT | 2 µL | 20 µL | CP |
| 9/17 pSB1C3-Pcon-RBS-tetR-DT x2 | 2 µL | 20 µL | CP |
| 9/14 pSB1C3-PkaiBC | 2 µL | 20 µL | CP |
| 9/14 pSB1C3-RpaB | 2 µL | 20 µL | CP |
| 9/16 pSB1C3-pT181 attenuator(EcoRI&SpeI) | 2 µL | 20 µL | CP |
| 9/17 pSB1C3-pT181 attenuator(XbaI&PstI) | 2 µL | 20 µL | CP |
| 9/17 pSB1C3-Pcon-spinach-DT | 2 µL | 20 µL | CP |
| 9/14 RBS-GFP-DT-PkaiBC | 2 µL | 20 µL | CP |
| Plux-RBS-lysis3-DT | 2 µL | 20 µL | CP |
| 9/20 Ptet-pT181 antisense-spinach-DT | 2 µL | 20 µL | CP |
| 9/20 Pcon-spinach-DT(pSB4K5) | 2 µL | 20 µL | Kan |
| 9/20 Pcon-pT181 antisense-spinach-DT(pSB4K5) | 2 µL | 20 µL | Kan |
| 9/20 KaiABC | 2 µL | 20 µL | Kan |
| 9/20 KaiABC | 2 µL | 20 µL | Kan |
| 9/20 Plac-RBS-lysis3-DT | 2 µL | 20 µL | Amp |
| 9/20 Plac-aptamer 12_1R | 2 µL | 20 µL | Amp |
| 9/20 Plux-RBS-GFP-DT-Pcon-RBS-luxR-DT | 2 µL | 20 µL | Amp |
| Plac-aptamer12_1R-DT | 2 µL | 20 µL | Amp |
| 9/20 Plac-aptamer12_1R-DT | 2 µL | 20 µL | Amp |
| 9/20Pcon-pT181attenuator-aptamer-DT | 2 µL | 20 µL | Amp |
| 9/17 Plac-pT181-attenuatorx2 | 2 µL | 20 µL | Amp |
| 9/15 Pcon-RBS-lacZα-DT | 2 µL | 20 µL | Amp |
| 9/14 Pcon-pT181attenuator-RBS-lacZα-DT | 2 µL | 20 µL | Amp |
| PUC19 | 2 µL | 20 µL | Amp |
PCR
| Sample | base pair |
|---|---|
| pT181 attenuator(pSB1C3) | -- |
| Pcon-RBS-lacZα-DT | -- |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94 °C | 94 °C | 55 °C | 68 °C | -- |
| 5 min | 30 s | 30 s | 3 min | 30 cycles |
Electrophoresis
| Lane | Sample | Enzyme |
|---|---|---|
| 1 | Pcon-RBS-lacZα-DT | -- |
| 2 | 1kb ladder | -- |
| 3 | pSB1C3-pT181attenuator | -- |
Sep 21
Electrophoresis
Liquid Culture
| Sample | medium |
|---|---|
| 9/2 spinach-DT-2 | -- |
| 8/16 Plux-4 | -- |
| 8/27 Plux-RBS-GFP-DT-1 | -- |
| 9/5 Pcon-pT181 attenuator-1 | -- |
Master Plate
| Number | Use LB plate |
|---|---|
| 1 | 8/27 Pλ-luxI-1(+Amp) |
| 2 | 9/10 Plac(BBa-R0011)-3(+Amp) |
| 3 | 9/5 Pcon-RBS-tetR-DT-1(+Amp) |
| 4 | 9/5 Pcon-pT181 antisense-1(+Amp) |
| 5 | 9/15 Pcon-pT181 attenuator(+Amp) |
| 6 | 9/- Pcon-apt12_1R-DT-1(+Amp) |
| 7 | 9/5 Pcon-spinach-DT-3(+Amp) |
| 8 | 8/27 Pcon-RBS-GFP-DT-Pcon-RBS-luxR-DT(+Amp) |
| Number | Use LB plate |
|---|---|
| 1 | 9/4 pT181 antisense(pSB1C3)-1(+CP) |
| 2 | 9/11 Fusion1 antisense(pSB1C3)-4(+CP) |
| 3 | 9/11 Fusion6 antisense(pSB1C3)-1(+CP) |
| 4 | 9/4 pT181 attenuator(pSB1C3)-1(+CP) |
| 5 | 9/10 Fusion1 attenuator(pSB1C3)-1(+CP) |
| 6 | 9/11 Fusion3n2 attenuator(pSB1C3)-1(+CP) |
| 7 | 9/4 aptamer12_1R(pSB1C3)-1(+CP) |
| 8 | 9/11 aptamer12_P(pSB1C3)-3(+CP) |
| 9 | 9/11 aptamer12_1M(pSB1C3)-1(+CP) |
</div>
| Number | Use LB plate |
|---|---|
| 1 | 8/29 Pbad/araC-RBS-RFP-DT-1(+CP) |
| 2 | 9/4 RBS-lysis3-DT-3(+CP) |
| 3 | 9/10 Pcon-pT181 attenuator-DT-1(+CP) |
| 4 | 9/5 Ptet-pT181 antisense-1(+CP) |
| 5 | 9/4aptamer 12_1R-DT-1(+CP) |
| 6 | 9/5 Ptet-RBS-lacZα-DT-4(+CP) |
| 7 | 8/10 RBS-lacI-DT-2(+CP) |
| 8 | 8/29 Plux-RBS-GFP-DT-1(+CP) |
</div>
Restriction Enzyme Digestion
| 8/17 RBS-DT | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cuts(XbaI&PstI) | 8.1 | 0µL | 0µL | 1µL | 1µL | 3µL | 3µL | 13.9µL | 30µL |
| NC | 0.4 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 8/17 RBS-DT | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cut(EcoRI&XbaI) | 16.2 | 1µL | 1µL | 0µL | 0µL | 3µL | 3µL | 5.8µL | 30µL |
| 9/15 Pcon-pT181attenuator | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cuts(EcoRI&PstI) | 12 | 1µL | 0µL | 0µL | 1µL | 3µL | 3µL | 10µL | 30µL |
| NC | 0.6 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.4µL | 10µL |
| 8/20 Pcon=RBS-GFP-DT-1 | EcoRI | SpeI | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cut(EcoRI&XbaI) | 6 | 1µL | 0µL | 1µL | 0µL | 3µL | 3µL | 0µL | 14µL |
| NC | 0.3 | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.3µL | 10µL |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | RBS-GFP-DT | XbaI | PstI |
| 2 | RBS-GFP-DT | -- | -- |
| 3 | RBS-GFP-DT | EcoRI | XbaI |
| 4 | 1kbp ladder | -- | -- |
| 5 | Pcon-pT181attenuator | SpeI | PstI |
| 6 | Pcon-pT181attenuator | -- | -- |
| 7 | Pcon-RBs-GFP-DT | EcoRI | XbaI |
| 8 | Pcon-RBs-GFP-DT | -- | -- |
PCR
| Sample | base pair |
|---|---|
| Pcon-apt12_1R-DT(E-1A) | 50 |
| Pcon-spinach-DT(E-1A) | 402 |
| Pcon-pT181 attenuator(E-1A) | 400 |
| DT(1-3) | 65 |
| tetR aptamer 12_1R-DT(1-3) | 283 |
| Pcon-pT181 attenuator(1-SA) | 303 |
| spinach-DT(2-1) | 338 |
| Pcon-pT181 attenuator(E-0) | 403 |
| Pcon-pT181 attenuator(2-SA) | 403 |
| spinach-DT(3-2) | 395 |
| Ptet-PT181 anisense | 400 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 98°C | 57°C | 68°C | -- |
| 2min | 10s | 30s | 35s | 30cycles |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kb ladder | -- |
| 3 | RBS-GFP-DT | XbaI&PstI |
| 4 | RBS-GFP-DT | XbaI&PstI |
| 5 | RBS-GFP-DT | XbaI&PstI |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| RBS-GFP-DT(XbaI&PstI) | 14.8 | 1.74 | 0.47 |
| RBS-GFP-DT(EcoRI&SpeI) | 88.4 | 1.85 | 1.49 |
| Pcon-RBS-GFP-DT(EcoRI&XbaI) | 47.5 | 1.82 | 1.12 |
Sep 22
Colony PCR
| Sample | base pair |
|---|---|
| 9/20 Plac-PT181 attenuator(1~4) | 664 |
| 9/20 Plac-PT181 antisense(1~6) | 369 |
| 9/20 Pcon-PT181 attenuator-aptamer-DT(1~6) | 859 |
| 9/20 Pcon-PT181 antisense-spinach-DT(1~8) | 723 |
Electrophoresis
| Lane | Sample |
|---|---|
| 1 | Pcon-apt12_1R-DT(E-1A) |
| 2 | Pcon-spinach-DT(E-1A) |
| 3 | Pcon-pT181 attenuator(E-1A) |
| 4 | DT(1-3) |
| 5 | tetR aptamer 12_1R-DT(1-3) |
| 6 | 100bp ladder |
| 7 | Pcon-pT181 attenuator(1-SA) |
| 8 | spinach-DT(2-1) |
| 9 | Pcon-pT181 attenuator(E-0) |
| 10 | Pcon-pT181 attenuator(2-SA) |
| 11 | spinach-DT(3-2) |
| 12 | Ptet-PT181 anisense |
Miniprep
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 9/21 spinach-DT | 197.9 | 1.67 | 1.54 |
| 9/21 Plux | 125.9 | 1.71 | 1.27 |
| 9/21 Plux-RBS-GFP-DT-(1) | 168.0 | 1.62 | 1.73 |
| 9/21 Pcon-PT181 attenuator-(1) | 368.3 | 1.57 | 1.62 |
| 9/21 RBS-GFP-DT | 532.8 | 1.75 | 1.78 |
| 9/21 Pcon-RBS-GFP-DT | 845.2 | 2.02 | 1.70 |
Colony PCR
| Sample | base pair |
|---|---|
| 9/20 Plux-RBS-GFP-DT-Pcon-RBS-luxR-DT-(1~2) | 283 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 2min10s | 30cycles |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2~4 | Pcon-aptamer12-1R-DT(E-1A) | -- |
| 6~8 | Pcon-spinach-DT(E-1A) | -- |
| 10~12 | Pcon-attenuator PT181(E-1A) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2~4 | DT(1-3) | -- |
| 6~8 | aptamer12-1RDT(1-3) | -- |
| 10~12 | Pcon-PT181 attenuator(1-5A) | -- |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-aptamer12-1R-DT(E-1A) | 31.2 | 1.87 | 0.30 |
| Pcon-spinach-DT(E-1A) | 25.6 | 1.88 | 0.99 |
| Pcon-attenuator PT181(E-1A) | 25.3 | 1.99 | 0.86 |
| DT(1-3) | 22.1 | 1.82 | 0.20 |
| aptamer12-1RDT(1-3) | 33.5 | 1.99 | 0.05 |
| Pcon-PT181 attenuator(1-5A) | 36.9 | 1.12 | 0.96 |
Restriction Enzyme Digestion
| 9/16 Plac | XbaI | SpeI | PstI | BufferB | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|
| 2cuts | 12.5µL | 0µL | 1µL | 1µL | 3µL | 3µL | 9.5µL | 30µL |
| NC | 0.8µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.4µL | 10µL |
| 9/22 Plux | XbaI | SpeI | PstI | BufferB | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|
| 2cuts | 16µL | 0µL | 1µL | 1µL | 3µL | 3µL | 14µL | 30µL |
| NC | 0.8µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.2µL | 10µL |
| 9/20 RBS-tetR-2R-DT | XbaI | SpeI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|
| 2cuts | 8.5µL | 1µL | 0µL | 1µL | 3µL | 3µL | 13.5µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2~4 | Pcon-attenuator PT181(E-0) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2~4 | Pcon-attenuator PT181(E-0) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 2 | 100bp ladder | -- |
| 4~6 | Pcon-attenuator PT181(2-SA) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 3~5 | spinach-DT(3-2) | -- |
| 8~10 | Ptet-antisense PT181(1-3) | -- |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-attenuator PT181(E-0) | 2.4 | 1.80 | 0.17 |
| Pcon-attenuator PT181(2-SA) | 10.8 | 1.85 | 0.54 |
| spinach-DT(3-2) | 36.7 | 1.80 | 0.75 |
| Ptet-antisense PT181(1-3) | 16.8 | 1.74 | 0.78 |
Colony PCR
| Sample | base pair |
|---|---|
| 9/20 Pcon-PT181 attenuator-aptamer12-1R-DT-(7~14) | 859 |
Colony PCR
| Sample | base pair |
|---|---|
| PconPT181antisense-spinach-DT(9~15) | 723 |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/13 Pcon-PT181 attenuator (SpeI & PstI) | 1.7 µL | 9/18 aptamer-1R-DT (XbaI & PstI) | 5.4µL | 3.5µL |
| experiment | 9/12 Plac(SpeI & PstI) | 2.4 µL | 9/18 aptamer-1R-DT (XbaI & PstI) | 6.1µL | 3.5 µL |
| experiment | 9/13 Ptet(SpeI & PstI) | 1.7 µL | 9/10 Plux-RBS-GFP-DT | 7.0 µL | 4.4 µL |
| experiment | 9/21 RBS-GFP-DT (EcoRI & XbaI) | 0.6 µL | 9/18 aptamer 12-1R-DT (XbaI & PstI) | 5.6µL | 3.1 µL |
| experiment | 9/13 PSB1C3(XbaI & PstI) | 1.9 µL | 9/16 PT181 attenuator(XbaI & PstI) | 4.6µL | 3.3 µL |
Liquid Culture
| Sample | medium |
|---|---|
| 9/20 Plac-antisense-1 | -- |
| 9/20 Plac-attenuator-1 | -- |
| 9/20 Plac-antisense-1 | -- |
| 9/20 Plac-attenuator-1 | -- |
digestion
| 9/8 Pcon-PT181 antisense | EcoRI | SpeI | BufferM | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 8.3µL | 1µL | 1µL | 3µL | 16.7µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 8.3µL | 10µL |
| 9/7 Pcon-RBS-tetR-DT | EcoRI | XbaI | SpeI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cut | 14µL | 1µL | 1µL | 0µL | 0µL | 3µL | 3µL | 18.2µL | 30µL |
| NC | 0.7µL | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.3µL | 10µL |
| 9/22 RBS-GFP-DT | EcoRI | XbaI | SpeI | PstI | BufferD | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|---|---|
| 2cuts | 3.8µL | 1µL | 1µL | 0µL | 0µL | 3µL | 3µL | 18.2µL | 30µL |
| NC | 0.2µL | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.8µL | 10µL |
| 9/6 aptamer12-1R-DT | EcoRI | XbaI | SpeI | PstI | BufferD | BSA | MilliQ | |
|---|---|---|---|---|---|---|---|---|
| 2cut | 8µL | 1µL | 1µL | 0µL | 0µL | 3µL | 3µL | 16µL |
| NC | 0.4µL | 0µL | 0µL | 0µL | 0µL | 1µL | 1µL | 7.6µL |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kbp ladder | -- |
| 3~5 | Pcon-PT181 antisense (EcoRI&SpaI) | -- |
Transformation
| Name | Sample | Competent Cells | Plate |
|---|---|---|---|
| 9/20 Plac-aptamer12-1R-DT(XbaI&PstI) | 2 µL | 20 µL | Amp |
| 9/20 pT181 attenuator(1C3)(EcoRI&SpeI) | 2 µL | 20 µL | CP |
| 9/17 pT181 attenuator(1C3)(XbaI&PstI) | 2 µL | 20 µL | CP |
| 9/20 Pcon-pT181 attenuator(SpaI&PstI)+aptamer12-1R-DT(XbaI&PstI) | 2 µL | 20 µL | Amp |
| kaiABC | 2 µL | 20 µL | Amp |
| 9/22 pSB1C3-RpaB(XbaI&PstI) | 2 µL | 20 µL | Amp |
| 9/22 pSB1C3-pT181 attenuator(XbaI&PstI) | 2 µL | 20 µL | Amp |
| 9/22 Ptet(SpaI&PstI)+RBS-GFP-DT(XbaI&PstI) | 2 µL | 20 µL | CP |
| 9/22 RBS-GFP-DT+Pcon-attenuator pT181 (EcoRI&SpeI) | 2 µL | 20 µL | CP |
| 9/22 pT181 attenuator(XbaI&PstI)(1C3) | 2 µL | 20 µL | CP |
PCR
| Sample | base pair |
|---|---|
| Pcon-RBS-tetR-DT(1-2(A)) | -- |
| Pcon-RBS-tetR-DT(3-2(A)) | -- |
| Pcon-RBS-GFP-DT(1-S(A)) | -- |
| Pcon-RBS-GFP-DT(2-S(A)) | -- |
| Pcon-tetR-DT(E-2(A)) | -- |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94 °C | 94 °C | 57 °C | 68 °C | -- |
| 2 min | 10 s | 30 s | 1min5s | 30 cycles |
Electrophoresis
| Lane | Sample |
|---|---|
| 1 | 1kbp ladder |
| 2 | Pcon-RBS-tetR-DT(1-2Ⓐ) |
| 3 | Pcon-RBS-tetR-DT(3-2Ⓐ) |
| 4 | Pcon-RBS-GFP-DT(1-SⒶ) |
| 5 | Pcon-RBS-GFP-DT(2-SⒶ) |
| 6 | Pcon-tetR-DT(E-2Ⓐ) |
Sep 23
Transformation
| Name | Sample | Competent Cells | Plate |
|---|---|---|---|
| 9/14 pSB1C3+PkaiBC | 2 µL | 20 µL | -- |
| 9/14 pSB1C3+RpaB | 2 µL | 20 µL | -- |
| 9/14 pkaiBC+Rbs-GFP-DT | 2 µL | 20 µL | -- |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kbp ladder | |
| 3-4 | Pcon-RBS-tetR-PT |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-PT181 anti EcoRI+SpeI | 46.7 | 1.06 | 0.93 |
| Pcon-RBS-tetR-DT EcoRI+XbaI | 119.3 | 0.99 | |
| RBS-8FP-DT EcoRI+XbaI | 87.7 | 1.00 | 0.98 |
| ap42_1R-DT EcoRI+XbaI | 68.8 | 1.32 | 1.03 |
| Pcon-RBS-tetR-DT 1-2 A | 203.0 | 1.66 | 1.79 |
| Pcon-RBS-tetR-DT 3-2 A | 67.7 | 2.64 | 1.28 |
| Pcon-RBS-GFP-DT 1-S A | 185.6 | 1.88 | 2.04 |
| Pcon-RBS-GFP-DT 2-S A | 298.6 | 1.68 | 1.55 |
| Pcon-tetR-DT E-2 A | 202.4 | 1.57 | 1.66 |
PCR
| genome DNA | KOD plus | 10x buffer | dNTP | MgSO4 | SasA_fwd primer | SasA_rev primer | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| 9/8 Pcon-pT181 anti | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 15.5 | 25 |
| 8/20 Ptet(1) | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 15.5 | 25 |
| 9/16 Plac 2 | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 15.5 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94 °C | 98 °C | 57 °C | 68 °C | -- |
| 2min | 10sec | 30sec | 20sec | 30cycles |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/21 Pcon-RBS-GFP-DT EcoRI+XbaI | 47.5 µg/mL | 9/15 Pcon-RBS-tetR-DT EcoRI+SpeI | 59 µg/mL | 2.3 µg/mL |
| experiment | 9/21 Pcon-RBS-GFP-DT EcoRI+XbaI | 47.5 µg/mL | 9/15 Pcon-apt12_1R-DT EcoRI+SpeI | 17 µg/mL | 2.4 µg/mL |
| experiment | 9/21 Pcon-RBS-GFRDT EcoRI+XbaI | 2.1 µg/mL | 9/15 Pcon-DT181 attenuator-DT | 21 µg/mL | 3.0 µg/mL |
| experiment | 9/22 apt 12_1R-DT EcoRI+XbaI | 67.8 µg/mL | 9/18 Pcon-DT181 attenuator EcoRI+SpeI | 5.7 µg/mL | 3.5 µg/mL |
| experiment | 9/22RBS-GFP-DT EcoRI+XbaI | 87.7 µg/mL | 9/18 Pcon-DT18 attenuator EcoRI+SpeI | 6.7 µg/mL | 3.5 µg/mL |
| experiment | 9/13 DT EcoRI+XbaI | 25.4 µg/mL | 9/22 Pcon-DT181 antisense EcoRI+SpeI | 46.8 µg/mL | 2.3 µg/mL |
PCR
| genome DNA | KOD plus | 10x buffer | dNTP | MgSO4 | SasA_fwd primer | SasA_rev primer | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| Pcon-attenuator-DT (E-1A) | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 77.5 | 25 |
| Pcon-attenuator-DT (2-1A) | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 77.5 | 25 |
| Pcon-antisense-spinach-DT (1-2A) | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 77.5 | 25 |
| Pcon-antisense-spinach-DT (E-1A) | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 77.5 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94 °C | 98 °C | 57 °C | 68 °C | -- |
| 2min | 10s | 30s | 45s | 30cycle |
Miniprep
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 9/22 Plac-DT181 attenuator-1 | 204.0 | 2.00 | 1.95 |
| 9/22 Plac-DT181 attenuator-2 | 195.3 | 1.84 | 1.62 |
| 9/22 Plac-DT181 antisense-1 | 188.2 | 1.87 | 2.15 |
| 9/22 Plac-DT181 antisense-2 | 165.0 | 1.95 | 2.66 |
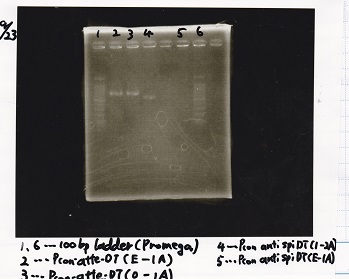
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1,6 | 100bp ladder(Promega) | -- | -- |
| 2 | Pcon-attenuator-DT (E-1A) | -- | -- |
| 3 | Pcon-attenuator-DT (2-1A) | -- | -- |
| 4 | Pcon-antisense-spinach-DT (1-2A) | -- | -- |
| 5 | Pcon-antisense-spinach-DT (E-1A) | -- | -- |
Gel Extraction
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-pT181 attenuator-DT(E-1A) | 166.7 | 1.51 | 0.99 |
| Pcon-pT181 attenuator-DT(2-1A) | 122.1 | 1.75 | 1.20 |
| Pcon-pT181 antisense-spinach-DT(1-2A) | 99.1 | 1.86 | 1.81 |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 100bp ladder | -- | -- |
| 2 | 9/8 Pcon-pT181 antisense(E-1A) | -- | -- |
| 3 | 8/20 Ptet(1)(2-S) | -- | -- |
| 4 | 9/16 Plac2(E-1A) | -- | -- |
Transformation
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| pSB1C3 | 2µL | 20µL | 22µL |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/19 Plac SpeI+PstI | 24.6 µg/mL | 9/18 ape12_1R-DT XbaI+PstI | 4.5 µg/mL | 2 µg/mL |
| experiment | 9/21 RBS-GFP-DT EcoRI+XbaI | 88.4 µg/mL | 9/15 Pcon-pT181 attenuator EcoRI+SpeI | 21 µg/mL | 2 µg/mL |
| experiment | 9/13 PSB1C3 XbaI+PstI | 26.0 µg/mL | 9/16 pT181 attenuator XbaI+PstI | 8.2 µg/mL | 2 µg/mL |
Colony PCR
| Sample | base pair |
|---|---|
| 9/22 Pcon-PT181 attenuator12-1R-DT(1~8) | -- |
| 9/22 Pcon-PT181 attenuator(1~4) | -- |
| 9/20 Plac+aptamer12-1R-DT(1~4) | -- |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 54s | 30cycle |
Sep 24
Electrophoresis
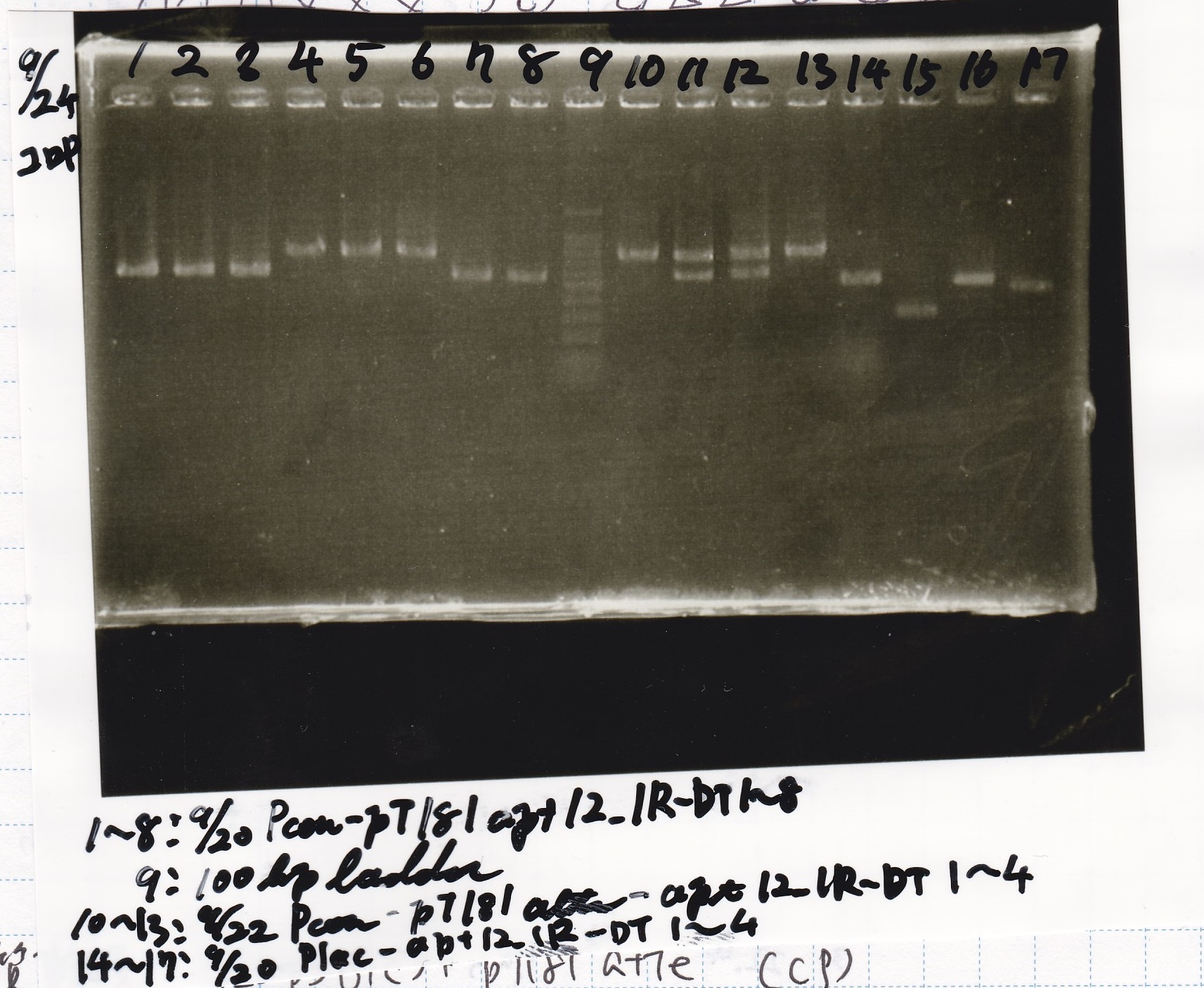
| Lane | Sample |
|---|---|
| 1 | 9/20 Pcon-pT181aptamer12-1R-DT 1 |
| 2 | 9/20 Pcon-pT181aptamer12-1R-DT 2 |
| 3 | 9/20 Pcon-pT181aptamer12-1R-DT 3 |
| 4 | 9/20 Pcon-pT181aptamer12-1R-DT 4 |
| 5 | 9/20 Pcon-pT181aptamer12-1R-DT 5 |
| 6 | 9/20 Pcon-pT181aptamer12-1R-DT 6 |
| 7 | 9/20 Pcon-pT181aptamer12-1R-DT 7 |
| 8 | 9/20 Pcon-pT181aptamer12-1R-DT 8 |
| 9 | 100bp ladder |
| 10 | 9/22 Pcon-pT181attenuator-aptamer12-1R-DT 1 |
| 11 | 9/22 Pcon-pT181attenuator-aptamer12-1R-DT 2 |
| 12 | 9/22 Pcon-pT181attenuator-aptamer12-1R-DT 3 |
| 13 | 9/22 Pcon-pT181attenuator-aptamer12-1R-DT 4 |
| 14 | 9/20 Plac-aptamer12-1R-DT 1 |
| 15 | 9/20 Plac-aptamer12-1R-DT 2 |
| 16 | 9/20 Plac-aptamer12-1R-DT 3 |
| 17 | 9/20 Plac-aptamer12-1R-DT 4 |
Transformation
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| 9/23 pSB1C3-pT181attenuator | 2 µL | 20 µL | 22 µL | LB (+CP) |
| RBS-GFP-DT+Pcon-pT181attenuator | 2 µL | 20 µL | 22 µL | LB (+CP) |
| 9/23 Plac-+spinach-DT | 2 µL | 20 µL | 22 µL | LB (+Amp) |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT | 2 µL | 20 µL | 22 µL | LB (+Amp) |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT | 2 µL | 20 µL | 22 µL | LB (+Amp) |
| 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT | 2 µL | 20 µL | 22 µL | LB (+Amp) |
| 9/23 Pcon-pT181antisense-DT | 2 µL | 20 µL | 22 µL | LB (+CP) |
| 9/23 Pcon-pT181attenuator-RBS-GFP-DT | 2 µL | 20 µL | 22 µL | LB (+CP) |
| 9/23 Pcon-pT181attenuator-aptamer12-1R-DT | 2 µL | 20 µL | 22 µL | LB (+CP) |
| PUC19 | 2 µL | 20 µL | 22 µL | LB (+Amp) |
| Ptet-RBS-lacZα-DT | 2 µL | 20 µL | 22 µL | LB (+CP) |
| pSB1C3 | 2 µL | 20 µL | 22 µL | LB (+CP) |
Liquid Culture
| Sample | medium |
|---|---|
| aptamer12-1R-DT(Master plate) | LB(+CP) |
Restriction Enzyme Digestion(PCR product)
Concentrate PCR product by evaporator until volume become less than 1µL, add all sulution and pipetting.
| 10*cut smart buffer | BsaI-HF | MilliQ | total | |
|---|---|---|---|---|
| 9/22 Pcon-pT181 attenuator (E-0) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-pT181 attenuator (2-3A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 spinach-DT(3-2) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Ptet-pT181 antisense (1-3) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-tet aptamer-DT (E-1A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-spinach-DT (E-1A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-pT181 attenuator (E-1A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 DT (1-3) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 tet aptamer-DT (1-3) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-pT181 attenuator (1-SA) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-RBS-tetR-DT (1-2A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-RBS-tetR-DT (3-2A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-RBS-tetR-DT (E-2A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-RBS-GFP-DT (1-2A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/22 Pcon-RBS-GFP-DT (2-SA) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/23 Pcon-pT181 attenuator-DT (E-1A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/23 Pcon-pT181 attenuator-DT (0-1A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/23 Pcon-pT181 antisense-spinach-DT (1-2A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/23 Pcon-pT181 antisense (1-2A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/23 Ptet-1 (2-S) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
| 9/23 Plac-2 (E-1A) | 2.0 µL | 0.3 µL | 17 µL | 19.3µL |
Column Refining
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 8/22 Pcon-pT181 attenuator(E-0) | 7.8 | 1.70 | 0.03 |
| 8/22 Pcon-pT181 attenuator(2-SA) | 17.3 | 1.93 | 0.03 |
| 8/22 spinach-DT (3-2) | 22.4 | 1.93 | 0.09 |
| 8/22 Ptet-PT181 anisense (1-3) | 18.3 | 1.58 | 0.14 |
| 8/22 Pcon-aptamer12_1R-DT (E-1A) | 8.2 | 2.10 | 0.05 |
| 8/22 Pcon-spinach-DT(E-1A) | 5.6 | 0.91 | 0.05 |
| 8/22 Pcon-pT181 attenuator(E-1A) | 18.5 | 1.80 | 0.12 |
| 8/22 DT(1-3) | 7.3 | 1.13 | 0.04 |
| 8/22 tetR aptamer 12_1R-DT(1-3) | 14.6 | 1.74 | 0.06 |
| 8/22 Pcon-pT181 attenuator(1-SA) | 7.9 | 1.57 | 0.04 |
| 8/22 Pcon-RBS-tetR-DT(1-2Ⓐ) | 41.6 | 1.81 | 0.25 |
| 8/22 Pcon-RBS-tetR-DT(3-2Ⓐ) | 30.2 | 1.75 | 0.15 |
| 8/22 Pcon-RBS-tetR-DT(E-2Ⓐ) | 36.9 | 1.75 | 0.56 |
| 8/22 Pcon-RBS-GFP-DT(1-SⒶ) | 43.5 | 1.83 | 0.33 |
| 8/22 Pcon-RBS-GFP-DT(2-SⒶ) | 64.6 | 1.82 | 0.30 |
| 8/23 Pcon-pT181 attenuator-DT (E-1A) | 31.2 | 1.83 | 0.09 |
| 8/23 Pcon-pT181 attenuator-DT (0-1A) | 64.5 | 1.80 | 0.43 |
| 8/23 Pcon-pT181 antisense-spinach-DT (1-2A) | 36.4 | 1.88 | 0.21 |
| 8/23 Pcon-pT181 antisense-spinach-DT (E-1A) | 27.9 | 1.92 | 0.16 |
| 8/23 Ptet(1)(2-S) | 37.0 | 1.91 | 0.16 |
| 9/23 Plac-2(E-1A) | 21.2 | 1.82 | 0.13 |
Transformation
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| 9/23 Pcon-attenuator-aptamer12-1R-DT | 2 µL | 20 µL | 22 µL | LB (+CP) |
| 9/23 pT181attenuator(1C3) | 2 µL | 20 µL | 22 µL | LB (+CP) |
| 9/23 Pcon-pT181antisense-DT | 2 µL | 20 µL | 22 µL | LB (+CP) |
| 9/23 Pcon-pT181attenuator-RBS--GFP-DT | 2 µL | 20 µL | 22 µL | LB (+CP) |
Liquid Culture
| Sample | medium |
|---|---|
| pSB1C3 (Master plate-1) | LB(+CP) |
Restriction Enzyme Digestion
| 8/21 pT181attenuator (330 µg/mL) | XbaI | PstI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 6.1 | 1 | 1 | 3 | 3 | 15.9 | 30 |
| NC | 0.3 | 0 | 0 | 1 | 1 | 7.7 | 10 |
| 9/22 Pcon-pT181attenuator (368.3 µg/mL) | EcoRI | SpeI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 5.4 | 1 | 1 | 3 | 0 | 19.6 | 30 |
| NC | 0.3 | 0 | 0 | 1 | 0 | 8.7 | 10 |
| 9/16 pSB1C3 | EcoRI | SpeI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 18 | 1 | 1 | 3 | 0 | 7.0 | 30 |
| NC | 0.9 | 0 | 0 | 1 | 0 | 8.1 | 10 |
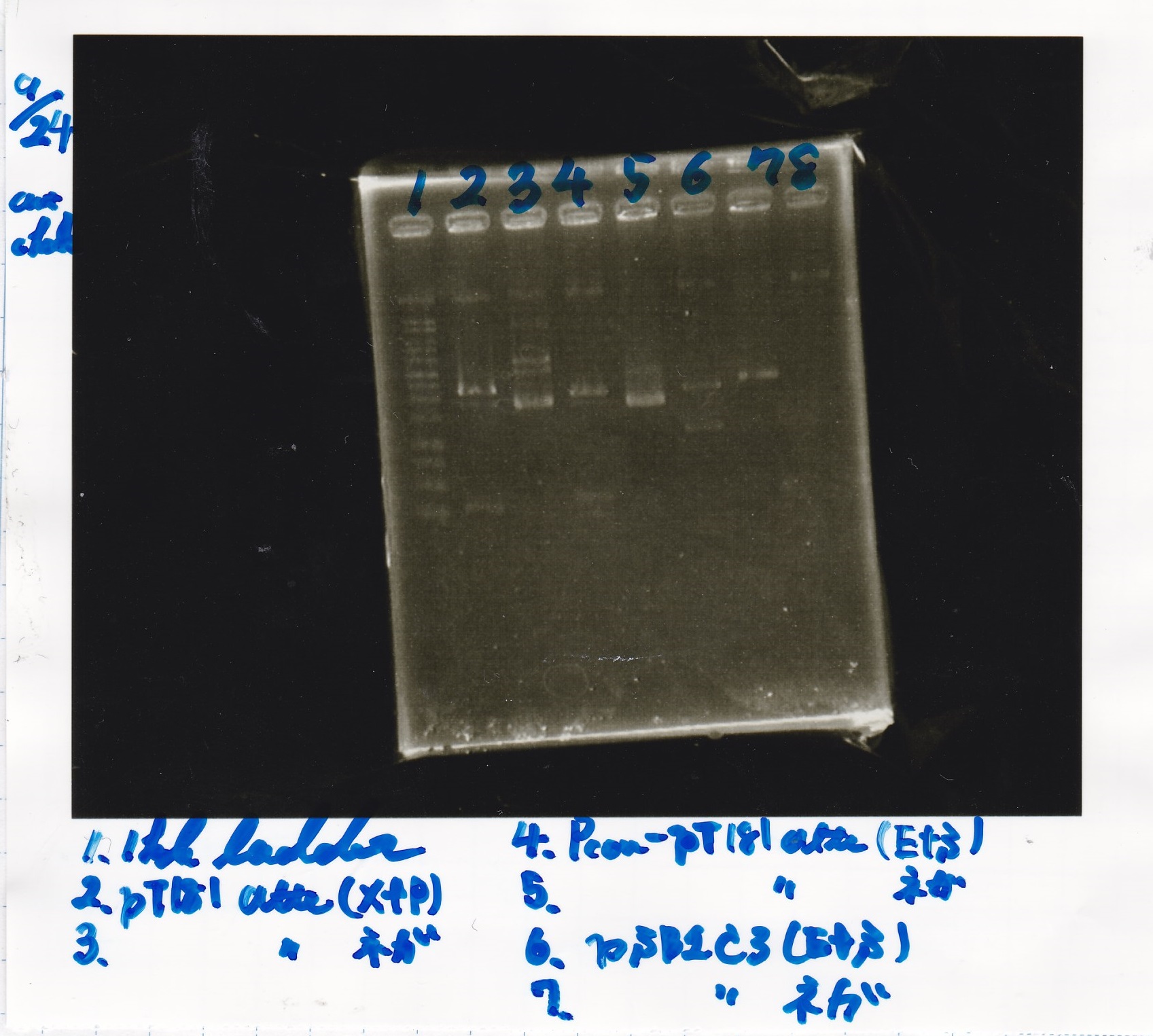
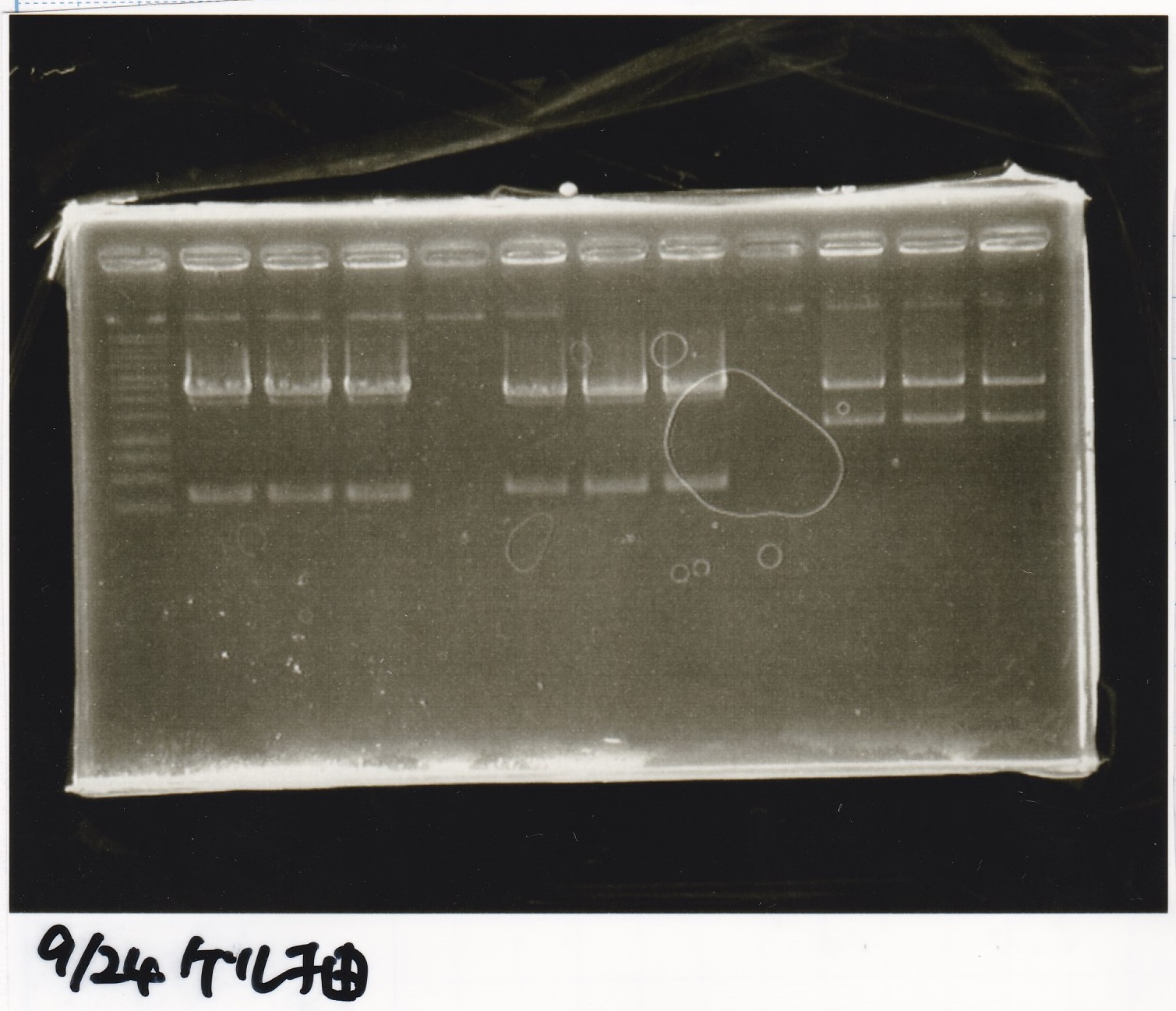
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kb ladder | - | - |
| 2 | pT181attenuator | XbaI | PstI |
| 3 | pT181attenuator | -- | -- |
| 4 | Pcon-pT181attenuator | EcoRI | SpeI |
| 5 | Pcon-pT181attenuator | -- | -- |
| 6 | pSB1C3 | EcoRI | SpeI |
| 7 | pSB1C3 | -- | -- |
Miniprep
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 9/24 aptamer12-1R-DT | 144.7 | 1.03 | 1.21 |
Gel Extraction
| Lane | DNA | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 100kb ladder | -- | -- |
| 2 | pT181attenuator | XbaI | PstI |
| 3 | pT181attenuator | XbaI | PstI |
| 4 | pT181attenuator | XbaI | PstI |
| 5 | -- | -- | -- |
| 6 | Pcon-pT181attenuator | EcoRI | SpeI |
| 7 | Pcon-pT181attenuator | EcoRI | SpeI |
| 8 | Pcon-pT181attenuator | EcoRI | SpeI |
| 9 | -- | -- | -- |
| 10 | pSB1C3 | EcoRI | SpeI |
| 11 | pSB1C3 | EcoRI | SpeI |
| 12 | pSB1C3 | EcoRI | SpeI |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| pT181attenuator(XbaI&PstI) | 10.1 | 2.02 | 0.43 |
| Pcon-pT181attenuator(EcoRI&SpeI) | 10.3 | 1.81 | 0.32 |
| pSB1C3(EcoRI&SpeI) | 10.0 | 2.81 | 0.02 |
Colony PCR
| Sample | base pair |
|---|---|
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 1 | 1414 |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 2 | 1414 |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 3 | 1414 |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 4 | 1414 |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 5 | 1414 |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 6 | 1414 |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 7 | 1414 |
| 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 8 | 1414 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 1 | 1631 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 2 | 1631 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 3 | 1631 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 4 | 1631 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 5 | 1631 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 6 | 1631 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 7 | 1631 |
| 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 8 | 1631 |
| 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 1 | 2042 |
| 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 2 | 2042 |
| 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 3 | 2042 |
| 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 4 | 2042 |
| 9/23 Pcon-pT181antisense-DT 1 | 508 |
| 9/23 Pcon-pT181antisense-DT 2 | 508 |
| 9/23 Pcon-pT181antisense-DT 3 | 508 |
| 9/23 Pcon-pT181antisense-DT 4 | 508 |
| 9/23 Pcon-pT181antisense-DT 5 | 508 |
| 9/23 Pcon-pT181antisense-DT 6 | 508 |
| 9/23 Pcon-pT181antisense-DT 7 | 508 |
| 9/23 Pcon-pT181antisense-DT 8 | 508 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94 °C | 94 °C | 55 °C | 68 °C | -- |
| 5 min | 30 sec | 30 sec | 2min | 30 cycles |
| Lane | Sample |
|---|---|
| 1 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 1 |
| 2 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 2 |
| 3 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 3 |
| 4 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 4 |
| 5 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 5 |
| 6 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 6 |
| 7 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 7 |
| 8 | 9/23 Pcon-aptamer12-1R-DT+Pcon-RBS-GFP-DT 8 |
| 9 | 1kb ladder |
| 10 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 1 |
| 11 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 2 |
| 12 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 3 |
| 13 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 4 |
| 14 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 5 |
| 15 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 6 |
| 16 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 7 |
| 17 | 9/23 Pcon-pT181attenuator-DT+Pcon-RBS-GFP-DT 8 |
| Lane | Sample |
|---|---|
| 1 | 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 1 |
| 2 | 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 2 |
| 3 | 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 3 |
| 4 | 9/23 Pcon-RBS-tetR-DT+Pcon-RBS-GFP-DT 4 |
| 5 | 9/23 Pcon-pT181antisense-DT 1 |
| 6 | 9/23 Pcon-pT181antisense-DT 2 |
| 7 | 1kb ladder |
| 8 | 9/23 Pcon-pT181antisense-DT 3 |
| 9 | 9/23 Pcon-pT181antisense-DT 4 |
| 10 | 9/23 Pcon-pT181antisense-DT 5 |
| 11 | 9/23 Pcon-pT181antisense-DT 6 |
| 12 | 9/23 Pcon-pT181antisense-DT 7 |
| 13 | 9/23 Pcon-pT181antisense-DT 8 |
| 14 | -- |
Restriction Enzyme Digestion
| 9/24 aptamer12-1R-DT | EcoRI | XbaI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 17.5µL | 1.0µL | 1.0µL | 3.0µL | 3.0µL | 4.5µL | 30µL |
| NC | 0.9µL | 0µL | 0µL | 1.0µL | 1.0µL | 7.1µL | 10µL |
| 9/20 Ptet | SpeI | PstI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 9.5µL | 1.0µL | 1.0µL | 3.0µL | 0µL | 15.5µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 1.0µL | 0µL | 8.5µL | 10µL |
| 9/22 RBS-GFP-DT | EcoRI | SpeI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 3.8µL | 1.0µL | 1.0µL | 3.0µL | 0µL | 21.2µL | 30µL |
| NC | 0.2µL | 0µL | 0µL | 1.0µL | 0µL | 8.8µL | 10µL |
| 9/22 Pcon-RBS-GFP-DT | EcoRI | XbaI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 2.4µL | 1.0µL | 1.0µL | 3.0µL | 3.0µL | 19.6µL | 30µL |
| NC | 0.1µL | 0µL | 0µL | 1.0µL | 1.0µL | 7.9µL | 10µL |
| 9/22 Pcon-pT181attenuator 1 | EcoRI | XbaI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 5.4µL | 1.0µL | 1.0µL | 3.0µL | 3.0µL | 16.6µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1.0µL | 1.0µL | 7.7µL | 10µL |
| 9/22 RBS-GFP-DT | XbaI | PstI | buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 3.8µL | 1.0µL | 1.0µL | 3.0µL | 3.0µL | 18.2µL | 30µL |
| NC | 0.2µL | 0µL | 0µL | 1.0µL | 1.0µL | 7.8µL | 10µL |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/22 aptamer12-1R-DT(EcoRI&XbaI) | 1.4 µL | 9/24 Pcon-pT181attenuator(EcoRI&SpeI) | 26 µL | 3.5 µL |
| experiment | 9/4 Ptet(SpeI&PstI) | 2.0 µL | 9/21 RBS-GFP-DT(XbaI&PstI) | 9.0 µL | 3.5 µL |
- incubated at 37 °C for 1 hour.
colony PCR
| Sample | base pair |
|---|---|
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 5 | 2042bp |
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 6 | 2042bp |
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 7 | 2042bp |
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 8 | 2042bp |
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 9 | 2042bp |
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 10 | 2042bp |
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 11 | 2042bp |
| 9/23 Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 12 | 2042bp |
| 9/23 Pcon-pT181antisense-DT 9 | 508bp |
| 9/23 Pcon-pT181antisense-DT 10 | 508bp |
| 9/23 Pcon-pT181antisense-DT 11 | 508bp |
| 9/23 Pcon-pT181antisense-DT 12 | 508bp |
| 9/23 Pcon-pT181antisense-DT 13 | 508bp |
| 9/24 Pcon-pT181attenuator-RBS-GFP-DT 1 | 1527bp |
| 9/24 Pcon-pT181attenuator-RBS-GFP-DT 2 | 1527bp |
| 9/24 Pcon-pT181attenuator-RBS-GFP-DT 3 | 1527bp |
| 9/24 Pcon-pT181attenuator-RBS-GFP-DT 4 | 1527bp |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT 1 | 859bp |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT 2 | 859bp |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT 3 | 859bp |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT 4 | 859bp |
| 9/24 pT181attenuator(pSB1C3) 1 | 390bp |
| 9/24 pT181attenuator(pSB1C3) 2 | 390bp |
| 9/24 pT181attenuator(pSB1C3) 3 | 390bp |
| 9/24 pT181attenuator(pSB1C3) 4 | 390bp |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 2min | 30cycle |
Miniprep
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| pSB1C3 | 183.9 | 1.91 | 1.59 |
| Pcon-pT181attenuator-aptamer12-1R-DT | 231.7 | 1.82 | 2.35 |
| Plac-aptamer12-1R-DT | 208.9 | 1.98 | 1.79 |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kb ladder | -- | -- |
| 2 | 9/24 aptamer12-1R-DT | EcoRI | XbaI |
| 3 | 9/24 aptamer12-1R-DT | -- | -- |
| 4 | 9/24 Ptet | SpeI | PstI |
| 5 | 9/24 Ptet | -- | -- |
| 6 | 9/24 RBS-GFP-DT | EcoRI | SpeI |
| 7 | 9/24 RBS-GFP-DT | -- | -- |
| 8 | 9/24 Pcon-RBS-GFP-DT | EcoRI | XbaI |
| 8 | 9/24 Pcon-RBS-GFP-DT | -- | -- |
| 9 | 9/24 Pcon-RBS-GFP-DT | -- | -- |
| 10 | 9/24 Pcon-pT181attenuator 1 | EcoRI | XbaI |
| 11 | 9/24 Pcon-pT181attenuator 1 | -- | -- |
| 12 | 9/24 RBS-GFP-DT | XbaI | PstI |
| 13 | 9/24 RBS-GFP-DT | -- | -- |
| 14 | 1kb ladder | -- | -- |
Gel Extraction
1
| Lane | DNA | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kb ladder | -- | -- |
| 2 | aptamer12-1R-DT | EcoRI | XbaI |
| 3 | aptamer12-1R-DT | EcoRI | XbaI |
| 4 | aptamer12-1R-DT | EcoRI | XbaI |
| 5 | -- | -- | -- |
| 6 | Ptet | SpeI | PstI |
| 7 | Ptet | SpeI | PstI |
| 8 | Ptet | SpeI | PstI |
| 9 | -- | -- | -- |
| 10 | RBS-GFP-DT | EcoRI | SpeI |
| 11 | RBS-GFP-DT | EcoRI | SpeI |
| 12 | RBS-GFP-DT | EcoRI | SpeI |
| Lane | DNA | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kb ladder | -- | -- |
| 2 | Pcon-RBS-GFP-DT | EcoRI | XbaI |
| 3 | Pcon-RBS-GFP-DT | EcoRI | XbaI |
| 4 | Pcon-RBS-GFP-DT | EcoRI | XbaI |
| 5 | -- | -- | -- |
| 6 | Pcon-pT181attenuator | EcoRI | XbaI |
| 7 | Pcon-pT181attenuator | EcoRI | XbaI |
| 8 | Pcon-pT181attenuator | EcoRI | XbaI |
| 9 | -- | -- | -- |
| 10 | RBS-GFP-DT | XbaI | PstI |
| 11 | RBS-GFP-DT | XbaI | PstI |
| 12 | RBS-GFP-DT | XbaI | PstI |
Electrophoresis
1
| Lane | Sample |
|---|---|
| 1 | 1kb ladder |
| 2 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 5 |
| 3 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 6 |
| 4 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 7 |
| 5 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 8 |
| 6 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 9 |
| 7 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 10 |
| 8 | 1kb ladder |
2
| Lane | Sample |
|---|---|
| 1 | 1kb ladder |
| 2 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 11 |
| 3 | Pcon-RBS-tetR-DT-Pcon-RBS-GFP-DT 12 |
| 4 | Pcon-pT181attenuator-RBS-GFP-DT 1 |
| 5 | Pcon-pT181attenuator-RBS-GFP-DT 2 |
| 6 | Pcon-pT181attenuator-RBS-GFP-DT 3 |
| 7 | Pcon-pT181attenuator-RBS-GFP-DT 4 |
| 8 | 1kb ladder |
3
| Lane | Sample |
|---|---|
| 1 | Pcon-pT181antisense-DT 9 |
| 2 | Pcon-pT181antisense-DT 10 |
| 3 | Pcon-pT181antisense-DT 11 |
| 4 | Pcon-pT181antisense-DT 12 |
| 5 | Pcon-pT181antisense-DT 13 |
| 6 | 100bp ladder |
| 7 | Pcon-pT181attenuator-aptamer12-1R-DT 1 |
| 8 | Pcon-pT181attenuator-aptamer12-1R-DT 2 |
4
| Lane | Sample |
|---|---|
| 1 | Pcon-pT181attenuator-aptamer12-1R-DT 3 |
| 2 | Pcon-pT181attenuator-aptamer12-1R-DT 4 |
| 3 | 100bp ladder |
| 4 | pT181attenuator(pSB1C3) 1 |
| 5 | pT181attenuator(pSB1C3) 2 |
| 6 | pT181attenuator(pSB1C3) 3 |
| 7 | pT181attenuator(pSB1C3) 4 |
Ligation
| state | Vector | Inserter1 | Inserter2 | Inserter3 | Inserter4 | (NEB)T4ligase | (NEB)Buffer | milliQ | total | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | tetR-aptamer12-1R-DT(1-3) | 1 µL | Ptet(2-S) | 0.3µL | Pcon-tetR-aptamer12-1R-DT(3-2A) | 1.1µL | Plac(E-1A) | 0.5µL | 0.5µL | 0.5µL | 0.2 µL | 5µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-pT181attenuator(E-1A) | 1 µL | Pcon-tetR-aptamer12-1R-DT(1-2A) | 1µL | Ptet(2-S) | 0.3µL | -- | -- | 0.5µL | 0.5µL | 0.7 µL | 5µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-tetRaptamer12-1R-DT(E-1A) | 2µL | Pcon-tetRaptamer12-1R-DT(1-2A) | 1µL | Ptet(2-S) | 0.3µL | -- | -- | 0.5µL | 0.5µL | 0 µL | 5.3µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-spinach-DT(E-1A) | 3µL | Pcon-tetRaptamer12-1R-DT(1-2A) | 1µL | Ptet(2-S) | 0.3µL | -- | -- | 0.5µL | 0.5µL | 0 µL | 6.3µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-pT181antisense-DT(E-1A) | 0.5µL | DT(1-3) | 2µL | Ptet(2-S) | 0.3µL | Pcon-tetRaptamer12-1R-DT(3-2A) | 1.1µL | 0.5µL | 0.5µL | 0.1 µL | 5µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-pT181attenuator(E-1A) | 1µL | tetRaptamer12-1R-DT(1-3) | 1µL | Ptet(2-S) | 0.3µL | Pcon-tetRaptamer12-1R-DT(3-2A) | 1.1µL | 0.5µL | 0.5µL | 0µL | 5.4µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Ptet(2-S) | 0.3µL | Pcon-tetRaptamer12-1R-DT(E-2A) | 1µL | -- | -- | -- | -- | 0.5µL | 0.5µL | 1.7µL | 5µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-GFP-DT(2-SA) | 0.5µL | 1µL | -- | -- | -- | -- | -- | 0.5µL | 0.5µL | 1.5µL | 5µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-tetRaptamer12-1R-DT(E-1A) | 2µL | Pcon-pT181attenuator(1-SA) | 2.1µL | -- | -- | -- | -- | 0.5µL | 0.5µL | 0µL | 6.1µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-pT181attenuator-DT(E-1A) | 1µL | Pcon-pT181attenuator(1-SA) | 2.1µL | -- | -- | -- | -- | 0.5µL | 0.5µL | 0µL | 5.1µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-tetR12-1R-DT(E-1A) | 2µL | Pcon-GFP-DT(2-SA) | 0.5µL | spinach-DT(3-2) | 1µL | Ptet-pT181antisense(1-3) | 1µL | 0.5µL | 0.5µL | 0µL | 6.5µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-tetRaptamer12-1R-DT(E-1A) | 2µL | Pcon-pT181antisense(1-2A) | 0.7µL | Pcon-pT181attenuator(2-SA) | 2.1µL | -- | -- | 0.5µL | 0.5µL | 0µL | 6.8µL |
| experiment | RBS-GFP-DT(pSB1C3) | 1µL | Pcon-tetRaptamer12-1R-DT(E-1A) | 2µL | Pcon-pT181attenuator(2-SA) | 2.1µL | Spinach-DT(3-2) | 1µL | Ptet-pT181antisense(1-3) | 1µL | 0.5µL | 0.5µL | 0µL | 8.1µL |
Concentrate total (more than 5µL) by evaporater until volume become less than 5 µL
PCR
| genome DNA | KOD plus | 10x buffer | dNTP | MgSO4 | F primer | R primer | KOD-plus | MilliQ | total |
|---|---|---|---|---|---|---|---|---|---|
| Pcon antisense(1C3-GGE-fwd,1A2-GG2-rev) | 1µL | 2.5µL | 2.5µL | 1.5µL | 0.75µL | 0.75µL | 0.5µL | 15.5 | 25µL |
| DT-(1C3-GG1-fwd,1C3-GG2-rev) | 1µL | 2.5µL | 2.5µL | 1.5µL | 0.75µL | 0.75µL | 0.5µL | 15.5 | 25µL |
| Spinach-DT-(1C3-GG1-fwd,1C3-GG2-rev) | 1µL | 2.5µL | 2.5µL | 1.5µL | 0.75µL | 0.75µL | 0.5µL | 15.5 | 25µL |
| Pcon-antisense-spinach-DT(1C3-GGE-fwd,1A2-GG1-rev) | 1µL | 2.5µL | 2.5µL | 1.5µL | 0.75µL | 0.75µL | 0.5µL | 15.5 | 25µL |
| Ptet(1C3-GGE-fwd,1C3-GG0-rev) | 1µL | 2.5µL | 2.5µL | 1.5µL | 0.75µL | 0.75µL | 0.5µL | 15.5 | 25µL |
| Pcon attenuator(1C3-GGE-fwd,1C3-GG0-rev) | 1µL | 2.5µL | 2.5µL | 1.5µL | 0.75µL | 0.75µL | 0.5µL | 15.5 | 25µL |
| DT(1C3-GG2-fwd,1C3-GG1-rev) | 1µL | 2.5µL | 2.5µL | 1.5µL | 0.75µL | 0.75µL | 0.5µL | 15.5 | 25µL |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 98°C | 57°C | 68°C | -- |
| 2min | 10sec | 30sec | 36sec | 30 |
Sep 25
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 100bp ladder | -- | -- |
| 2 | Pcon-antisense (1C3-66E-fwd A2-662 rev) | -- | -- |
| 3 | DT (1C3-661-fwd 1C3 662 rev) | -- | -- |
| 4 | spinach-DT(1C3-661-fwd 1C3 662 rev) | -- | -- |
| 5 | Pcon-antisense-spinach-DT(1C3-66E fwd A2-661 rev) | -- | -- |
| 6 | 100bp ladder | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 2 | 100bp ladder | -- | -- |
| 3 | Ptet (1C3 66E fwd-1C3 660 rev) | -- | -- |
| 4 | Pcon-attenuator(1C3-660 fwd A2-661 rev) | -- | -- |
| 5 | DT(1C3 662 fwd-1C3 661 rev) | -- | -- |
| 6 | Pcon-attenuator-aptamer-DT(1C3 66E fwd-A2-661-rev) | -- | -- |
| 7 | 100bp ladder | -- | -- |
Colony PCR
| Sample | base pair |
|---|---|
| 9/23 Pcon-PT181 attenuator-aptamer12-1R-DT-(5~12) | 859 |
| 9/23 Pcon-PT181 attenuator-RBS-GFP-DT-(5~8) | 1527 |
| 9/23 Pcon-PT181 attenuator-pSB1C3-(5~8) | 601 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94 °C | 94 °C | 55 °C | 68 °C | -- |
| 2 min | 30 s | 30 s | 1min5s | 30 cycles |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | Pcon-PT181 attenuator-aptamer12-1R-DT-5 | -- | -- |
| 2 | Pcon-PT181 attenuator-aptamer12-1R-DT-6 | -- | -- |
| 3 | Pcon-PT181 attenuator-aptamer12-1R-DT-7 | -- | -- |
| 4 | Pcon-PT181 attenuator-aptamer12-1R-DT-8 | -- | -- |
| 5 | Pcon-PT181 attenuator-aptamer12-1R-DT-9 | -- | -- |
| 6 | Pcon-PT181 attenuator-aptamer12-1R-DT-10 | -- | -- |
| 7 | Pcon-PT181 attenuator-aptamer12-1R-DT-11 | -- | -- |
| 8 | Pcon-PT181 attenuator-aptamer12-1R-DT-12 | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | Pcon-PT181 attenuator-RBS-GFP-DT-5 | -- | -- |
| 11 | Pcon-PT181 attenuator-RBS-GFP-DT-6 | -- | -- |
| 12 | Pcon-PT181 attenuator-RBS-GFP-DT-7 | -- | -- |
| 13 | Pcon-PT181 attenuator-RBS-GFP-DT-8 | -- | -- |
| 14 | Pcon-PT181 attenuator-pSB1C3-5 | -- | -- |
| 15 | Pcon-PT181 attenuator-pSB1C3-6 | -- | -- |
| 16 | Pcon-PT181 attenuator-pSB1C3-7 | -- | -- |
| 17 | Pcon-PT181 attenuator-pSB1C3-8 | -- | -- |
Transformation
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| Plac+tet aptamer12+Pcon tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181attenuator-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-Spinach-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181antisense+DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181attenuator+tet aptamer12-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon tetR-DT+Pcon-GFP-DT+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181attenuator-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Ptet-pT181antisense+Spinach-DT+Pcon-RBS-GFP-DT+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Pcon-pT181antisense+Spinach-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Ptet-pT181antisense+Spinach-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| aptamer12-DT(EcoRI+XbaI)+Pcon-pT181attenuator(EcoRI+SpeI) | 2µL | 20µL | 22µL | -- |
| Ptet(SpeI+PstI)+RBS-GFP-DT(XbaI+PstI)) | 2µL | 20µL | 22µL | -- |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/25 aptamer12_1R-DT(EcoRI&XbaI) | 4.5 µL | Pcon-pT181 attenuator(EcoRI&SpeI) | 7.7 µL | 6.1 µL |
| experiment | 9/17 pSB1C3(EcoRI&SpeI) | 2.0 µL | 9/3 pT181 attenuator(EcoRI&SpeI) | 14 µL | 8 µL |
| experiment | 9/25 Pcon-pT181 attenuator(EcoRI&XbaI) | 4.5 µL | 9/25 RBS-GFP-DT(EcoRI&SpeI) | 19.3 µL | 11.9 µL |
| experiment | 9/25 Pcon-RBS-GFP-DT(EcoRI&XbaI) | 4.2 µL | Pcon-RBS-tetR-DT(EcoRI&SpeI) | 20 µL | 12.1 µL |
Restriction Enzyme Digestion
| 9/24 Pcon-attenuator-aptamer-DT(µL) | EcoRI(µL) | SpeI(µL) | Buffer(µL) | MilliQ(µL) | total(µL) | |
|---|---|---|---|---|---|---|
| 2 cuts | 8.6 | 1 | 1 | 3 | 16.4 | 30 |
| NC | 0.4 | 0 | 0 | 1 | 8.6 | 10 |
| 9/11 DT(µL) | EcoRI(µL) | XbaI(µL) | Buffer(µL) | BSA(µL) | MilliQ(µL) | total(µL) | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 10.4 | 1 | 1 | 3 | 3 | 11.6 | 30 |
| NC | 0.5 | 0 | 0 | 1 | 1 | 7.5 | 10 |
Liquid Culture
| Sample | medium |
|---|---|
| DT | -- |
| pT181attenuator | -- |
| Pcon-RBS-tetR-DT | -- |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2 | Pcon-pT181antisense | -- |
| 3 | ||
| 4 | ||
| 5 | -- | -- |
| 6 | DT | -- |
| 7 | ||
| 8 | ||
| 9 | -- | -- |
| 10 | Spinach-DT | -- |
| 11 | ||
| 12 |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 3 | Pcon-antisense-spinach-DT(E-1A) | -- |
| 4 | Pcon-antisense-spinach-DT(E-1A) | -- |
| 5 | Pcon-antisense-spinach-DT(E-1A) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 7 | 100bp ladder | -- |
| 9 | Ptet(E-0) | -- |
| 10 | Ptet(E-0) | -- |
| 11 | Ptet(E-0) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 13 | 100bp ladder | -- |
| 15 | Pcon-attenuator(0-1A) | -- |
| 16 | Pcon-attenuator(0-1A) | -- |
| 17 | Pcon-attenuator(0-1A) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 19 | 100bp ladder | -- |
| 21 | Pcon-attenuator-aptamer-DT(E-1A) | -- |
| 22 | Pcon-attenuator-aptamer-DT(E-1A) | -- |
| 23 | Pcon-attenuator-aptamer-DT(E-1A) | -- |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| DT(2-1) | 24.9 | 1.73 | 0.37 |
| Ptet(E-0) | 12.8 | 1.59 | 0.90 |
| Pcon-antisense(E-2A) | 16.5 | 1.72 | 1.07 |
| Pcon-attenuator-aptamer-DT(E-1A) | 22.1 | 1.75 | 0.94 |
| Pcon-antisense-spinach-DT(E-1A) | 31.0 | 1.78 | 1.15 |
| spinach-DT(1-2) | 23.4 | 1.85 | 1.11 |
| Pcon-attenuator(0-1A) | 22.5 | 1.76 | 0.46 |
| DT(1-2) | 25.1 | 1.78 | 1.21 |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kbp ladder | -- | -- |
| 2 | Pcon-pT181 attenuator-aptamer 12_1R-DT | EcoRI | SpeI |
| 3 | Pcon-pT181 attenuator-aptamer 12_1R-DT | -- | -- |
| 4 | DT | EcoRI | XbaI |
| 5 | DT | -- | -- |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | DT | EcoRI+XbaI |
| 2 | DT | EcoRI+XbaI |
| 3 | Pcon-pT181 attenuator-aptamer12_1R-DT | EcoRI+SpeI |
| 4 | Pcon-pT181 attenuator-aptamer12_1R-DT | EcoRI+SpeI |
| 5 | Pcon-pT181 attenuator-aptamer12_1R-DT | EcoRI+SpeI |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-pT181 attenuator-aptamer12_1R-DT(EcoRI&SpeI) | 11.8 | 1.80 | 0.46 |
| DT(EcoRI+XbaI) | 42.6 | 1.84 | 0.98 |
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-pT181anntisense-spinach-DT(4k5) | - |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/24 pSB1C3(EcoRI+Spel) | 10µL | 9/25 PconpT181attenuator-aptamer12-1R-DT(EcoRI+Spel) | 5.9µL | 8µL |
PCR
| 800pg/μL DT | KOD plus | 10x buffer | dNTP | MgSO4 | 1C3_GG2_fwd | iC3_GG1_rev | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| 1 | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 15.5 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 98°C | 57°C | 68°C | -- |
| 2min | 10s | 30s | 24s | 30cycles |
Transformation
| Name | Plate |
|---|---|
| aptamer12-1R-DT | CP |
| pSB1C3(EcoRI+SpeI)+pT181attenuator(EcoRI+Spal) | CP |
| Pcon-pT181attenuator(EcoRI+XbaI)+RBS-GFP-DT(EcoRI+SpeI) | Amp |
| Pcon-RBS-GFP-DT(EcoRI+XbaI)+Pcon-RBS-tetR-DT(EcoRI+SpeI) | Amp |
| pSB1C3(EcoRI+SpeI)+Pcon-pT181attenuator-aptamer12-1R-DT(EcoRI+SpeI) | CP |
Electrophoresis
Column Refining
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 8/25 DT(2-1) | -- | -- | -- |
Restriction Enzyme Digestion
Concentrate PCR product by evaporator until volume become less than 3µL, add all sulution and pipetting.
| 9/25 DT(2-1) | 10*cut smart Buffer | BsaI-HF | MilliQ | total |
|---|---|---|---|---|
| 3.0 µL | 2.0 µL | 0.3 µL | 14.7 µL | 20.0 µL |
Incubate 37°C 12hour
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-GFP-DT | LB(Amp) |
| RBS-GFP-DT | LB(CP) |
| Pcon-spinach-DT | LB(Amp) |
| Pcon-tetRaptamer-DT | LB(Amp) |
| Pcon-attenuator-DT | LB(Amp) |
| spinach-DT | LB(CP) |
| Pcon-antisense-spinach-DT | LB(Amp) |
Plating
Mix Liquid Culture 1mL and LB(+Amp), incubate 37°C 30min
| Sample | Use plate | The number of plate |
|---|---|---|
| Pcon-GFP-DT | M9(+Amp) | 3 |
| Pcon-GFP-DT | LB(+Amp) | 3 |
</div> incubate 37°C
Colony PCR
| Sample | base pair |
|---|---|
| 9/24 GGA-1(1~8) | 3177 |
| 9/24 GGA-2(1~8) | 3189 |
| 9/24 GGA-3(1~8) | 2989 |
| 9/24 GGA-4(1~8) | 2939 |
| 9/24 GGA-7(1) | 2489 |
| GGA-16(1) | 2717 |
| RBS-GFP-DT | -- |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 3min15s | 30cycles |
RNA Extraction
sumple 0.25mL
Add 1mLISOGEN-LS
Lysis or homogenization
Store for 5 min, at room temperature
Add 0.2mL Chloroform
Shake vigorously for 15sec.
Store for 2~3min.at room temperature
Centrifuge 12K*g for 15min. at 4°C
Extract aqueous phase
Add 0.5mL isopropanol
Store for 5~10min. at room temperature
Centrifuge 12K*g for 10min. at 4°C
Remove aqueous phase
Add at least 1mL 70% ethanol
Vortex
Centrifuge 7.5K*g for 5min. at 4°C
Remove aqueous phase
Dry briefly
Add ddH2O, TE(pH8.0) or 0.5% SDS
Dissolve
Total RNA solution
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-RBS-GFP-DT | 2783 | 1.95 | 1.48 |
Add 91.2µL, Store at freezer
Total 250µL</div>
cDNA Synthesis
| volume | 7 µL |
|---|---|
| 7*cDNA Wipcout Buffer | 1 µL |
| Template RNA(9/25 Pcon-RBS-GFP-DT) | 3 µL |
| RNase-frec water | 4 µL |
Incubate 42°C 2min, then on ice immediately
Add
| 5*Quantiscript Reverse Transcriptase | 0.5 µL |
|---|---|
| 5*Quantiscript RT Buffer | 2 µL |
| RT Primer Mix | 0.5µL |
Incubate 42°C 15min, then incubate 95°C 3min
Electrophoresis
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/13 Pcon-pT181-attenuator(SpeI&PstI) | 3.5µL | 9/25 RBS-GFP-DT(XbaI&PstI) | 13.5µL | 8.5 µL |
Sep 26
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA1-1 | -- | -- |
| 2 | GGA1-2 | -- | -- |
| 3 | GGA1-3 | -- | -- |
| 4 | GGA1-4 | -- | -- |
| 5 | GGA1-5 | -- | -- |
| 6 | GGA1-6 | -- | -- |
| 7 | GGA1-7 | -- | -- |
| 8 | GGA1-8 | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | GGA2-1 | -- | -- |
| 11 | GGA2-2 | -- | -- |
| 12 | GGA2-3 | -- | -- |
| 13 | GGA2-4 | -- | -- |
| 14 | GGA2-5 | -- | -- |
| 15 | GGA2-6 | -- | -- |
| 16 | GGA2-7 | -- | -- |
| 17 | GGA2-8 | -- | -- |
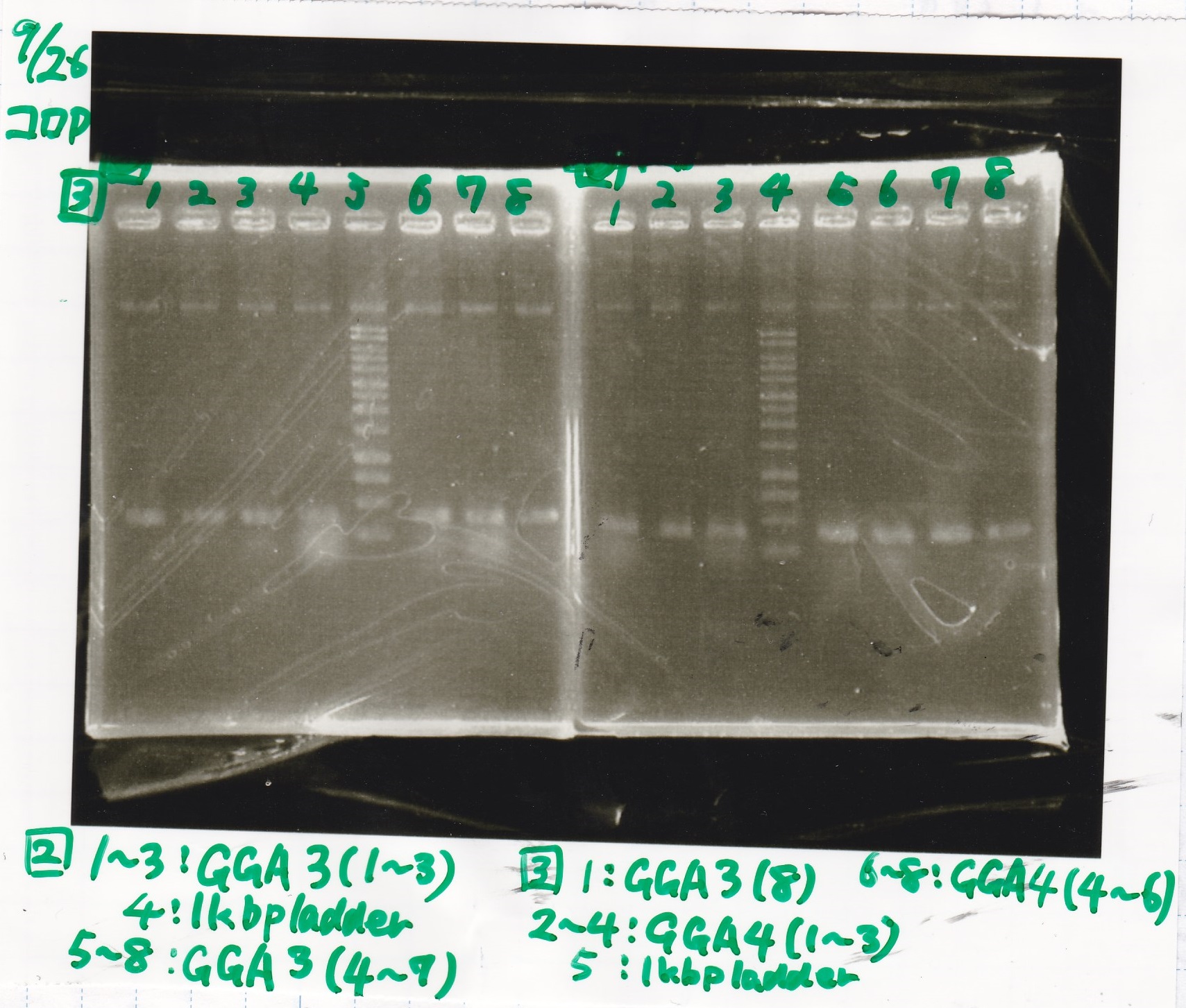
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA3-1 | -- | -- |
| 2 | GGA3-2 | -- | -- |
| 3 | GGA3-3 | -- | -- |
| 4 | 1kbp ladder | -- | -- |
| 5 | GGA3-4 | -- | -- |
| 6 | GGA3-5 | -- | -- |
| 7 | GGA3-6 | -- | -- |
| 8 | GGA3-7 | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA3-8 | -- | -- |
| 2 | GGA4-1 | -- | -- |
| 3 | GGA4-2 | -- | -- |
| 4 | GGA4-3 | -- | -- |
| 5 | 1kbp ladder | -- | -- |
| 6 | GGA4-4 | -- | -- |
| 7 | GGA4-5 | -- | -- |
| 8 | GGA4-6 | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA4-7 | -- | -- |
| 2 | GGA4-8 | -- | -- |
| 3 | 1kbp ladder | -- | -- |
| 4 | GGA7-1 | -- | -- |
| 5 | GGA16-1 | -- | -- |
| 6 | NC(RBS-GFP-DT) | -- | -- |
Ligasion(Golden Gate Assenbly)
| state | Vector | Inserter1 | Inserter2 | Inserter3 | Inserter4 | Inserter5 | (NEB)T4 ligase | (NEB)10*T4 ligase | MilliQ | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| experiment | RBS-GFP-DT | 1.0 µL | Pcon-pT181 antisense(E-2A) | 0.6µL | DT(2-1) | 0.5µL | Pcon-pT181 attenuator(1-SA) | 2.1 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | pSB1C3(EcoRI&SpeI) | 1.8 µL | Pcon-pT181 antisense(E-2A) | 0.9 µL | DT(2-1) | 0.5 µL | Pcon-GFP-DT(1-SA) | 1.5 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Ptet(E-0) | 0.7µL | Pcon pT181 attenuator(0-1A) | 0.3µL | tet aptamer(1-3) | 0.3 µL | Pcon-tetR-DT(3-2A) | 1.1 µL | Ptet(2-5) | 0.3 µL | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Ptet(E-0) | 0.7 µL | Pcon-pT181 attenuator-DT(0-1A) | 1.0 µL | Pcon-tetR-DT(1-2A) | 1.0 µL | Ptet(2-5) | 0.3µL | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Pcon-pT181 attenuator-tetR-DT(E-1A) | 1.0 µL | Pcon antisense-spinach-DT(1-2A) | 0.7µL | Pcon-pT 181 attenuator(2-SA) | 2.1 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | RBS-GFP-DT | 1.0 µL | Pcon antisense(E-1A) | 0.5 µL | spinach-DT(1-2) | 1.0 µL | Pcon-GFP-DT(1-2) | 0.8 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | pSB1C3(EcoRI&SpeI) | 1.8 µL | Pcon-pT181 antisense(E-1A) | 0.8µL | spinach-DT(1-2) | 1.0µL | Pcon-GFP-DT(2-SA) | 0.8 µL | -- | -- | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
| experiment | pSB1C3(EcoRI&SpeI) | 1.8 µL | Pcon-tet aptamer-DT(E-1A) | 3.0 µL | Ptet-pT181 antisense(1-3) | 1.5 µL | spinach-DT(3-2) | 1.5 µL | Pcon-GFP-DT(2-5A) | 0.8 µL | -- | -- | 0.5 µL | 0.5 µL | up to 5 µL |
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-GFP-DT | Amp |
| RBS-GFP-DT | CP |
| Pcon-pT181 attenuator-DT | Amp |
| Pcon-tet aptamer-DT | Amp |
| Pcon-spinach-DT | Amp |
| Pcon-pT181 antisense-spinach-DT | Amp |
| spinach-DT | CP |
BSAI Digestion
| DNA | MilliQ | BSAI-HF | 10*cut smart | total | |
|---|---|---|---|---|---|
| Pcon | 7 µL | 10.7 µL | 0.3 µL | 2 µL | 20 µL |
| RBS-luxR-DT | 10 µL | 7 µL | 0.3 µL | 2 µL | 20 µL |
Colony PCR
| Sample | base pair |
|---|---|
| GGA-1 (9~16) | 3177 |
| GGA-2 (9~16) | 3189 |
| GGA-3 (9~21) | 2989 |
| GGA-4 (9~16) | 2939 |
| GGA-6 (1) | 3359 |
| GGA-16(2~4) | 2939 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 3min30s | 30cycles |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-1 (9) | -- | -- |
| 2 | GGA-1 (10) | -- | -- |
| 3 | GGA-1 (11) | -- | -- |
| 4 | GGA-1 (12) | -- | -- |
| 5 | GGA-1 (13) | -- | -- |
| 6 | GGA-1 (14) | -- | -- |
| 7 | GGA-1 (15) | -- | -- |
| 8 | GGA-1 (16) | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | GGA-2 (9) | -- | -- |
| 11 | GGA-2 (10) | -- | -- |
| 12 | GGA-2 (11) | -- | -- |
| 13 | GGA-2 (12) | -- | -- |
| 14 | GGA-2 (13) | -- | -- |
| 15 | GGA-2 (14) | -- | -- |
| 16 | GGA-2 (15) | -- | -- |
| 17 | GGA-2 (16) | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-4 (9) | -- | -- |
| 2 | GGA-4 (10) | -- | -- |
| 3 | GGA-4 (11) | -- | -- |
| 4 | GGA-4 (12) | -- | -- |
| 5 | GGA-4 (13) | -- | -- |
| 6 | GGA-4 (14) | -- | -- |
| 7 | GGA-4 (15) | -- | -- |
| 8 | GGA-4 (16) | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | GGA-6 (1) | -- | -- |
| 11 | GGA-16 (2) | -- | -- |
| 12 | GGA-16 (3) | -- | -- |
| 13 | GGA-16 (4) | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-3 (9) | -- | -- |
| 2 | GGA-3 (10) | -- | -- |
| 3 | GGA-3 (11) | -- | -- |
| 4 | GGA-3 (12) | -- | -- |
| 5 | 1kbp ladder | -- | -- |
| 6 | GGA-3 (13) | -- | -- |
| 7 | GGA-3 (15) | -- | -- |
| 8 | GGA-3 (14) | -- | -- |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | GGA-3 (17) | -- | -- |
| 2 | GGA-3 (18) | -- | -- |
| 3 | GGA-3 (19) | -- | -- |
| 4 | 1kbp ladder | -- | -- |
| 5 | GGA-3 (20) | -- | -- |
| 6 | GGA-3 (21) | -- | -- |
| 7 | GGA-3 (16) | -- | -- |
Electrophoresis
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-GFP-DT | -- |
| RBS-GFP-DT | -- |
| Pcon-tet aptamer-DT | -- |
| Pcon-spinach-DT | -- |
| Pcon-pT181 antisense-spinach-DT | -- |
| spinach-DT | -- |
Colony PCR
| Sample | base pair |
|---|---|
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-1 | -- |
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-2 | -- |
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-3 | -- |
| 9/25 Pcon-pH181 attenuator+RBS-GFP-DT-4 | -- |
| 9/24 Ptet-RBS-GFP-DT-1 | -- |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 1min40s | 30cycles |
Ligasion(Golden Gate Assenbly)
| state | Vector | Inserter1 | Inserter2 | Inserter3 | Inserter4 | Inserter5 |
|---|---|---|---|---|---|---|
| experiment | GGA2 | Pcon-attenuator(E-1A) | Pcon-tetR-DT(1-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) | |
| experiment | GGA3 | Pcon-tetap-DT(E-1A) | Pcon-tetR-DT(1-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) | |
| experiment | GGA4 | Pcon-spi-DT(E-1A) | Pcon-tetR-DT(1-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) | |
| experiment | GGA5 | Pcon-antisense(E-1A) | DT(1-3) | Pcon-tetR-DT(3-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) |
| experiment | GGA6 | Pcon-attenuator(E-1A) | tetap-DT(1-3) | Pcon-tetR-DT(3-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) |
| experiment | GGA7 | Pcon-tetR-DT(E-2A) | Ptet(2-S) | RBS-GFP-DT(EcoRI&XbaI) |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 37°C | 16°C | 50°C | 80°C | -- |
| 3min | 4min | 5min | 5min | 25cycles |
RNA Extraction
| Sample |
|---|
| Pcon-Spinach-DT |
| Pcon-tetRaptamer-DT |
| Pcon-attenuator-DT |
| Spinach-DT |
| Pcon-antisense-Spinach-DT |
Liquid Culture
| Sample |
|---|
| Pcon-attemuater-DT |
| Pcon-aptamer-DT |
| Pcon-spinach-DT |
| Pcon-RBS-tetR-DT |
| Pcon-antisense-spinach-DT(Master 1) |
| Pcon-antisense |
Colony PCR
| Sample | base pair |
|---|---|
| 9/25 9/19pSB1C3&9/3PT181 attenuator(1~3) | 601bp |
| 9/25 aptamer12_1R-DT&9/24Pcon-pT181 attenuator(1~4) | 859bp |
| Pcon-attenuator-aptamer-DT(1) | 859bp |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 65°C | -- |
| 5min | 30sec | 30sec | 54sec | 30 |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kbp ladder | -- | -- |
| 2 | 9/25 9/23 1C3&9/3attenuator | -- | -- |
| 3 | 9/25 9/23 1C3&9/3attenuator | -- | -- |
| 4 | 9/25 9/23 1C3&9/3attenuator | -- | -- |
| 5 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
| 6 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
| 7 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
| 8 | 9/24 Pcon attenuator-aptamer-DT(1) | -- | -- |
| 9 | 9/25aptamer-DT&9/24Pcon-attenuator | -- | -- |
Plating
| Sample | Use plate |
|---|---|
| 9/26 GGA2 | LB(CP) |
| 9/26 GGA3 | LB(CP) |
| 9/26 GGA4 | LB(CP) |
| 9/26 GGA5 | LB(CP) |
| 9/26 GGA6 | LB(CP) |
| 9/26 GGA7 | LB(CP) |
| 9/26 GGA11 | LB(CP) |
| 9/26 GGA312 | LB(CP) |
| 9/26 GGA13 | LB(CP) |
| 9/26 GGA14 | LB(CP) |
| 9/26 GGA15 | LB(CP) |
| 9/26 GGA20 | LB(CP) |
| 9/26 GGA23 | LB(CP) |
| 9/26 GGA24 | LB(CP) |
| 9/26 Ptet+RBS-GFP-DT | LB(CP) |
incubate 37°C
Restriction Enzyme Digestion
| 9/24 pSB1C3 | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 11µL | 1µL | 1µL | 3µL | 14µL | 30µL |
| NC | 0.5µL | 0µL | 0µL | 1µL | 8.5µL | 10µL |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 8.7µL | 1µL | 1µL | 3µL | 16.3µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 8.6µL | 10µL |
| 9/24 pT181attenuator | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 3.1µL | 1µL | 1µL | 3µL | 21.9µL | 30µL |
| NC | 0.2µL | 0µL | 0µL | 1µL | 8.8µL | 10µL |
| 8/17 RBS-GFP-DT | EcoRI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 1cut | 16.4µL | 1µL | 3µL | 3µL | 6.6µL | 30µL |
| NC | 0.4µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/17 pSB4K5 | EcoRI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 8.5µL | 1µL | 1µL | 3µL | 3µL | 13.5µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/14 pSB4K5 | EcoRI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 7.3µL | 1µL | 1µL | 3µL | 3µL | 14.7µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/22 Pcon-pT181attenuator-DT | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 5.4µL | 1µL | 1µL | 3µL | 19.6µL | 30µL |
| NC | 0.3µL | 0µL | 0µL | 1µL | 8.7µL | 10µL |
| 8/7 RBS-GFP-DT 1 | XbaI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 8.1µL | 1µL | 1µL | 3µL | 3µL | 13.9µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
| 9/26 RBS-GFP-DT 1 | XbaI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 1cut | 8.1µL | 1µL | 3µL | 3µL | 14.9µL | 30µL |
| 9/16 Pλ-luxI | EcoRI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 14.7µL | 1µL | 1µL | 3µL | 3µL | 7.3µL | 30µL |
| NC | 0.7µL | 0µL | 0µL | 1µL | 1µL | 7.3µL | 10µL |
| 9/21 Ptet | EcoRI | SpeI | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2cuts | 14.2µL | 1µL | 1µL | 3µL | 10.8µL | 30µL |
| NC | 0.7µL | 0µL | 0µL | 1µL | 8.3µL | 10µL |
| 9/24 Pcon-pT181attenuator-aptamer12-1R-DT | XbaI | PstI | BSA | buffer | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2cuts | 8.7µL | 1µL | 1µL | 3µL | 3µL | 13.3µL | 30µL |
| NC | 0.4µL | 0µL | 0µL | 1µL | 1µL | 7.6µL | 10µL |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | pSB1C3 | EcoRI | SpeI |
| 2 | pSB1C3 | -- | -- |
| 3 | Pcon-pT181attenuator-aptamer12-1R-DT | EcoRI | SpeI |
| 4 | Pcon-pT181attenuator-aptamer12-1R-DT | -- | -- |
| 5 | 1kb ladder | -- | -- |
| 6 | pT181attenuator | EcoRI | SpeI |
| 7 | pT181attenuator | -- | -- |
| 8 | RBS-GFP-DT | EcoRI | -- |
| 9 | RBS-GFP-DT | -- | -- |
Sep 27
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kbp ladder | -- |
| 2 | pSB1C3 | EcoRI+SpeI |
| 3 | ||
| 4 | ||
| 6 | Pcon-attenuator-aptmer-DT | EcoRI+SpeI |
| 7 | ||
| 8 | ||
| 10 | Pcon-attenuator-aptmer-DT | EcoRI+SpeI |
| 11 | ||
| 12 |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-attenuator-DT(EcoRI+SpeI) | 6.8 | 1.98 | 0.43 |
| pT181-attenuator(EcoRI+SpeI) | 7.5 | 1.84 | 0.06 |
| pSB1C3(EcoRI+SpeI) | 17.7 | 1.97 | 0.71 |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kbp ladder | -- |
| 3 | pSB4K5 | EcoRI+PstI |
| 4 | pSB4K5 | EcoRI+PstI |
| 5 | pSB4K5 | EcoRI+PstI |
| 7 | pSB6A1 | EcoRI+PstI |
| 8 | pSB6A1 | EcoRI+PstI |
| 9 | pSB6A1 | EcoRI+PstI |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | pSB4K5 | EcoRI | PstI |
| 2 | pSB4K5 | - | - |
| 3 | 1kbp ladder | - | - |
| 4 | Pcon-attenuator | EcoRI | SpeI |
| 5 | Pcon-attenuator | - | - |
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | RBS-GFP-DT | XbaI | PstI |
| 2 | RBS-GFP-DT | - | - |
| 3 | RBS-GFP-DT | EcoRI | XbaI |
| 4 | 1kbp ladder | - | - |
| 5 | Ptet | EcoRI | SpeI |
| 6 | Ptet | - | - |
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-GFP | Amp |
| RBS-GFP-DT | CP |
| Pcon-attenuator | Amp |
| Pcon-attenuator-aptamer-DT | Amp |
| Pcon-tetR-DT | Amp |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/17 pSB1C3 (EcoRI+SpeI) 10.0ng/µL | 10µL | 9/26 pT181 attenuator(EcoRI+SpeI) 7.5ng/µL | 1.3µL | 3.5 µL |
| experiment | 9/25 DT (EcoRI+XbaI) 42.8ng/µL | 2.3µL | 9/22 Pcon-pT181 antisense(EcoRI+SpeI) 46.7ng/µL | 1.5µL | 1.9 µL |
| experiment | 9/27 pSB4K5(EcoRI+PstI) 12.9ng/µL | 7.8µL | 9/10 Pcon-attenuator(EcoRI+SpeI) 13.8ng/µL | 8.0µL | 3.5 µL |
| experiment | 9/27 pSB4K5(EcoRI+PstI) 12.9ng/µL | 7.8µL | 9/25 RBS-GFP-DT(XbaI+PstI) 13.9ng/µL | 12µL | 3.5 µL |
| experiment | 9/8 pSB4K5 (EcoRI+SpeI) 17.6ng/µL | 5.7 µL | 8/21 Pcon-GFP-DT(EcoRI+SpeI) 11ng/µL | 12.6µL | 3.5 µL |
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-spinach-DT | plusgrow Amp |
| Pcon-tetRaptamer-DT | plusgrow Amp |
| Pcon-antisense-spinach-DT | plusgrow Amp |
| spinach-DT | plusgrow CP |
RNA Extraction
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 9/26 Pcon-GFP-DT | 423.0 | 1.83 | 1.46 |
| 9/26 Pcon-GFP-DT K | 298.7 | 1.82 | 1.50 |
Electrophoresis
Miniprep
| DNA |
|---|
| Pcon-aptamer-DT |
| Pcon-spinach-DT |
| Pcon-antisense |
| Pcon-tetR-DT |
| Pcon-attenuator-DT |
| Pcon-antisense-spinach-DT |
Transformation
| Name1 | Name2 | Sample1(µL) | Sample2(µL) | Competent Cells(µL) | Total(µL) |
|---|---|---|---|---|---|
| Pcon-attenuator+RBS-GFP-DT(4K5) | Pcon-attenuator-DT(1A2) | 2 | 1 | 30 | 33 |
| Pcon-attenuator+RBS-GFP-DT(4K5) | Pcon-antisense-spinach-DT(1A2) | 2 | 1 | 30 | 33 |
| Pcon-attenuator+RBS-GFP-DT(4K5) | Pcon-tetR aptamer-DT(1A2) | 2 | 1 | 30 | 33 |
| Pcon-RBS-GFP-DT(4K5) | Pcon-attenuator-DT(1A2) | 2 | 1 | 30 | 33 |
| Pcon-RBS-GFP-DT(4K5) | Pcon-antisense-spinach-DT(1A2) | 2 | 1 | 30 | 33 |
| Pcon-RBS-GFP-DT(4K5) | Pcon-tetR aptamer-DT(1A2) | 2 | 1 | 30 | 33 |
| Pcon-attenuator+RBS-GFP-DT(4K5) | -- | 2 | -- | 20 | 22 |
| Pcon-RBS-GFP-DT(4K5) | -- | 2 | -- | 20 | 22 |
| attenuator(1C3) | -- | 2 | -- | 20 | 22 |
Miniprep
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-RBS-GFP-DT | 137.6 | 1.95 | 1.78 |
| RBS-GFP-DT | 77.0 | 2.06 | 1.96 |
| Pcon-PT181 attenuator | 231.0 | 1.89 | 1.89 |
| Pcon-antisense-Spinach-DT(2) | 130.0 | 1.56 | 1.62 |
| Pcon-aptamer-DT | 218.8 | 1.92 | 1.72 |
| Pcon-attenuator-DT | 180.7 | 1.95 | 1.78 |
| Pcon-Spinach-DT | 177.6 | 1.96 | 1.06 |
| Pcon-antisense-Spinach-DT(1) | 181.1 | 1.96 | 1.92 |
| Pcon-antisense | 152.9 | 2.02 | 1.96 |
| Pcon-RBS+ezR-DT | 164.1 | 1.93 | 1.91 |
Restriction Enzyme Digestion
| 9/27 RBS-GFP-DT | XbaI | PstI | Buffer | BSA | MilliQ | total | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 12µL | 1µL | 1µL | 3µL | 3µL | 20µL | 40µL |
| NC | 1.3µL | 0µL | 0µL | 1µL | 1µL | 6.7µL | 10µL |
| 9/27 Pcon-tetR-DT | EcoRI | PstI | Buffer | MilliQ | total | |
|---|---|---|---|---|---|---|
| 2 cuts | 12.2µL | 1µL | 1µL | 3µL | 12.8µL | 30µL |
| NC | 0.6µL | 0µL | 0µL | 1µL | 8.4µL | 10µL |
RNA Extraction
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 9/27 Pcon-RBS-GFP-DT | 686.6 | 1.99 | 1.34 |
| 9/27 Pcon-RBS-GFP-DT (+High salt) | 127.6 | 1.76 | 0.27 |
DNA sythesis
| genome DNA | DNA volume | genome DNA wipeout buffer | RNase freewater | RT | RT buffer | Primer Mix | total |
|---|---|---|---|---|---|---|---|
| 9/27 Pcon-RBS-GFP-DT | 0.8µL | 1µL | 5.2µL | 0.5µL | 2µL | 0.5µL | 10µL |
| 9/27 pcon-RBS-GFP-DT(+high salt) | 2.5µL | 1µL | 3.5µL | 0.5µL | 2µL | 0.5µL | 10µL |
| 9/27 Pcon-RBS-GFP-DT(revised concentration) | 0.5µL | 1µL | 5.5µL | 0.5µL | 2µL | 0.5µL | 10µL |
Electrophoresis
Liquid Culture
| Sample | medium |
|---|---|
| Pcon spinach DT | plusgrow 2µL(Amp) |
| Pcon aptamer DT | plusgrow 2µL(Amp) |
| Pcon antisense Spinach DT | plusgrow 2µL(Amp) |
| Spinach DT | plusgrow 4µL(CP) |
RT PCR
| genome DNA | KOD plus | 10x buffer for KOD-plus-ver.2 | 2nM dNTPs | 25mM MgSO4 | F primer | R primer | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| 0.5 | 1 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 15.5 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 98°C | 49.8°C | 68°C | -- |
| 2min | 10sec | 30sec | 10sec | 30 |
Electrophoresis
| Lane | Sample |
|---|---|
| 1 | Pcon-GFP-DT(GFP) |
| 2 | Pcon-GFP-DT(internal) |
| 3 | Pcon-GFP-DT(HS)(GFP) |
| 4 | Pcon-GFP-DT(internal) |
Digesting cut check
| Lane | Sample |
|---|---|
| 1 | RBS-GFP-DT(XbaI&PstI) |
| 2 | RBS-GFP-DT NC(XbaI&PstI) |
| 3 | 1kbp ladder |
| 4 | Pcon-RBS-tetR-DT(EcoRI&SpeI) |
| 5 | Pcon-RBS-tetR-DT NC(EcoRI&SpeI) |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 1kbp ladder | -- |
| 3 | RBS-GFP-DT | Xbal+PstI |
| 4 | RBS-GFP-DT | Xbal+PstI |
| 5 | RBS-GFP-DT | Xbal+PstI |
| 7 | 1kbp ladder | -- |
| 9 | Pcon-tetR-DT | EcoRI+SpeI |
| 10 | Pcon-tetR-DT | EcoRI+SpeI |
| 11 | Pcon-tetR-DT | EcoRI+SpeI |
RNA Extraction
| 0 | 5min | 15min | 20min | 23min | 28min | 33min | 36min | |
|---|---|---|---|---|---|---|---|---|
| aptamer generator | 0.348 | 0.408 | 0.550 | 0.865 | ||||
| Pcon-spinach-DT | 0.250 | 0.240 | 0.305 | 0.426 | 0.471 | 0.554 | 0.668 | 0 |
| GFP generator | 0.389 | 0.420 | 0.516 | 0.884 | ||||
| antisense-spinach | 0.371 | 0.438 | 0.537 | 0.733 | ||||
| Spinach-DT | 0.084 | 0.110 | 0.134 | 0.433 | 0.269 | 0.326 | 0.409 | 0.742 |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Spinach-DT | 83.9 | 1.63 | 0.17 |
| Pcon-tetRaptamer-DT | 425.1 | 2.10 | 0.64 |
| Pcon-RBS-GFP-DT | 262.4 | 2.01 | 0.84 |
| Pcon-antisense-Spinach-DT | 489.1 | 2.05 | 0.68 |
| Pcon-Spinach-DT | 266.6 | 2.01 | 0.66 |
cDNA Synthesis
| 9/27 Spinach-DT | 9/23 Pcon-tetRaptamer-DT | 9/23 Pcon-RBS-GFP-DT | 9/23 Pcon-antisense-spinach-DT | 9/23 Pcon-Spinach-DT | |
|---|---|---|---|---|---|
| RNA | 1 | 1 | 1 | 1 | 1 |
| gDNA wipeout | 6 | 1.2 | 1.9 | 1.0 | 1.9 |
| RNase-free water | 0 | 4.8 | 4.1 | 5.0 | 4.1 |
| RT | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 |
| RT Buffer | 2 | 2 | 2 | 2 | 2 |
| Primer mix | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 |
| total | 10µL | 10µL | 10µL | 10µL | 10µL |
 "
"