From 2013.igem.org
Sep 6
File:Igku Notebook Sep6 1.jpg
File:Igku Notebook Sep6 2.jpg
File:Igku Notebook Sep6 3.jpg
File:Igku Notebook Sep6 4.jpg
File:Igku Notebook Sep6 5.jpg
File:Igku Notebook Sep6 6.jpg
File:Igku Notebook Sep6 7.jpg
File:Igku Notebook Sep6 8.jpg
File:Igku Notebook Sep6 9.jpg
Electrophoresis
Hirano
| Lane | Sample
|
| 1 | 100bp ladder
|
| 2 | 9/4 RBS-lysis3-DT -3
|
| 3 | 9/4 RBS-lysis3-DT -4
|
| 4 | 9/4 RBS-lysis3-DT -5
|
| 5 | 9/4 RBS-lysis1-DT -3
|
| 6 | 9/4 Ptrc-KaiC -1
|
| 7 | 9/4 pSB4K5 -1
|
| 8 | 100bp ladder
|

Colony PCR
No name
| Sample | base pair
|
| Pconst-spinach-DT-(1~4) | 563
|
| Plac-spinach-DT-(1~4) | 659
|
| Pcon-pT181 antisense-(1~4) | 468
|
| Ptet-pT181 antisense-(1~4) | 604
|
| Pconst-pT181 attenuator-(1~4) | 568
|
| Pconst+RBS-tetR-DT-(1~4) | 1121
|
| Ptet+RBS-lacZα-DT-(1~4) | 1162
|
| PreDenature | Denature | Annealing | Extension | cycle
|
| 94°C | 94°C | 55°C | 68°C | --
|
| 5min | 30s | 30s | 40s | 30cycles
|
Liquid Culture
No name
| Sample | medium
|
| 9/4 RBS-Lysis3+DT3 | Plusgrow medium(+CP)
|
| 9/4 RBS-Lysis1+DT3 | Plusgrow medium(+CP)
|
| 9/4 pSB4K5 | Plusgrow medium(+Kan)
|
Electrophoresis
No name
| Lane | Sample
|
| 1 | 100bp ladder
|
| 2 | Pconst-spinach-DT -(1)
|
| 3 | Pconst-spinach-DT -(2)
|
| 4 | Pconst-spinach-DT -(3)
|
| 5 | Pconst-spinach-DT -(4)
|
| 6 | Plac-spinach-DT -(1)
|
| 7 | Plac-spinach-DT -(2)
|
| 8 | Plac-spinach-DT -(3)
|
| 9 | Plac-spinach-DT -(4)
|
| 10 | Pcon-pT181 antisense -(1)
|
| 11 | Pcon-pT181 antisense -(2)
|
| 12 | Pcon-pT181 antisense -(3)
|
| 13 | Pcon-pT181 antisense -(4)
|
| Lane | Sample
|
| 1 | 100bp ladder
|
| 2 | Ptet-pT181 antisense -(1)
|
| 3 | Ptet-pT181 antisense -(2)
|
| 4 | Ptet-pT181 antisense -(3)
|
| 5 | Ptet-pT181 antisense -(4)
|
| 6 | Plac-pT181 attenuator -(1)
|
| 7 | Plac-pT181 attenuator -(2)
|
| 8 | Plac-pT181 attenuator -(3)
|
| 9 | Plac-pT181 attenuator -(4)
|
Miniprep
Honda and Kojima
| DNA | concentration [µg/mL] | 260/280 | 260/230
|
| pT181 antisense-(1) | 238.7 | 1.68 | 1.44
|
| pT181 antisense-(2) | 248.8 | 1.83 | 1.77
|
| pT181 antisense-(3) | 223.2 | 1.78 | 1.61
|
| pT181 antisense-(4) | 220.8 | 1.89 | 1.92
|
| tetR aptamer-(1) | 252.5 | 1.89 | 1.90
|
| tetR aptamer-(2) | 211.2 | 1.90 | 1.99
|
| DT181 attenuator | 102.2 | 1.98 | 2.17
|
| 121R aptamer-DT | 251.7 | 1.92 | 2.01
|
| pSB4K5 | 212.9 | 1.80 | 1.90
|
| RBS-lacZα-DT | 185.7 | 1.78 | 1.79
|
| RBS-lysis2-DT-(1) | 84.8 | 1.85 | 2.24
|
Restriction Enzyme Digestion
Tatsui
| | 9/6 RBS-lysis2-DT (78.3 µg/mL) | XbaI | PstI | BSA | Buffer | MilliQ | total
|
| 2 cuts | 22 µL | 1 µL | 1 µL | 3 µL | 3 µL | 0 µL | 30 µL
|
| NC | 1.3 µL | 0 µL | 0 µL | 1 µL | 1 µL | 6.7 µL | 10 µL
|
| | aptamer-DT (251.7 µg/mL) | XbaI | PstI | BSA | Buffer | MilliQ | total
|
| 2 cuts | 0.4 µL | 1 µL | 1 µL | 3 µL | 3 µL | 14.1 µL | 30 µL
|
| NC | 0.4 µL | 0 µL | 0 µL | 1 µL | 1 µL | 7.6 µL | 10 µL
|
| | Pλ-luxI (86.4 µg/mL) | EcoRI | SpeI | BSA | Buffer | MilliQ | total
|
| 2 cuts | 23.1 µL | 1 µL | 1 µL | 3 µL | 3 µL | 1.9 µL | 30 µL
|
| NC | 1.2 µL | 0 µL | 0 µL | 1 µL | 1 µL | 6.8 µL | 10 µL
|
Electrophoresis
Hirano
| Lane | Sample
|
| 1 | 100bp ladder
|
| 2 | Pconst+RBS-tetR-DT -(1)
|
| 3 | Pconst+RBS-tetR-DT -(2)
|
| 4 | Pconst+RBS-tetR-DT -(3)
|
| 5 | Pconst+RBS-tetR-DT -(4)
|
| 6 | Ptet+RBS-lacZα-DT -(1)
|
| 7 | Ptet+RBS-lacZα-DT -(2)
|
| 8 | 100bp ladder
|
| 9 | Ptet+RBS-lacZα-DT -(3)
|
| 10 | Ptet+RBS-lacZα-DT -(4)
|
| 11 | 100bp ladder
|
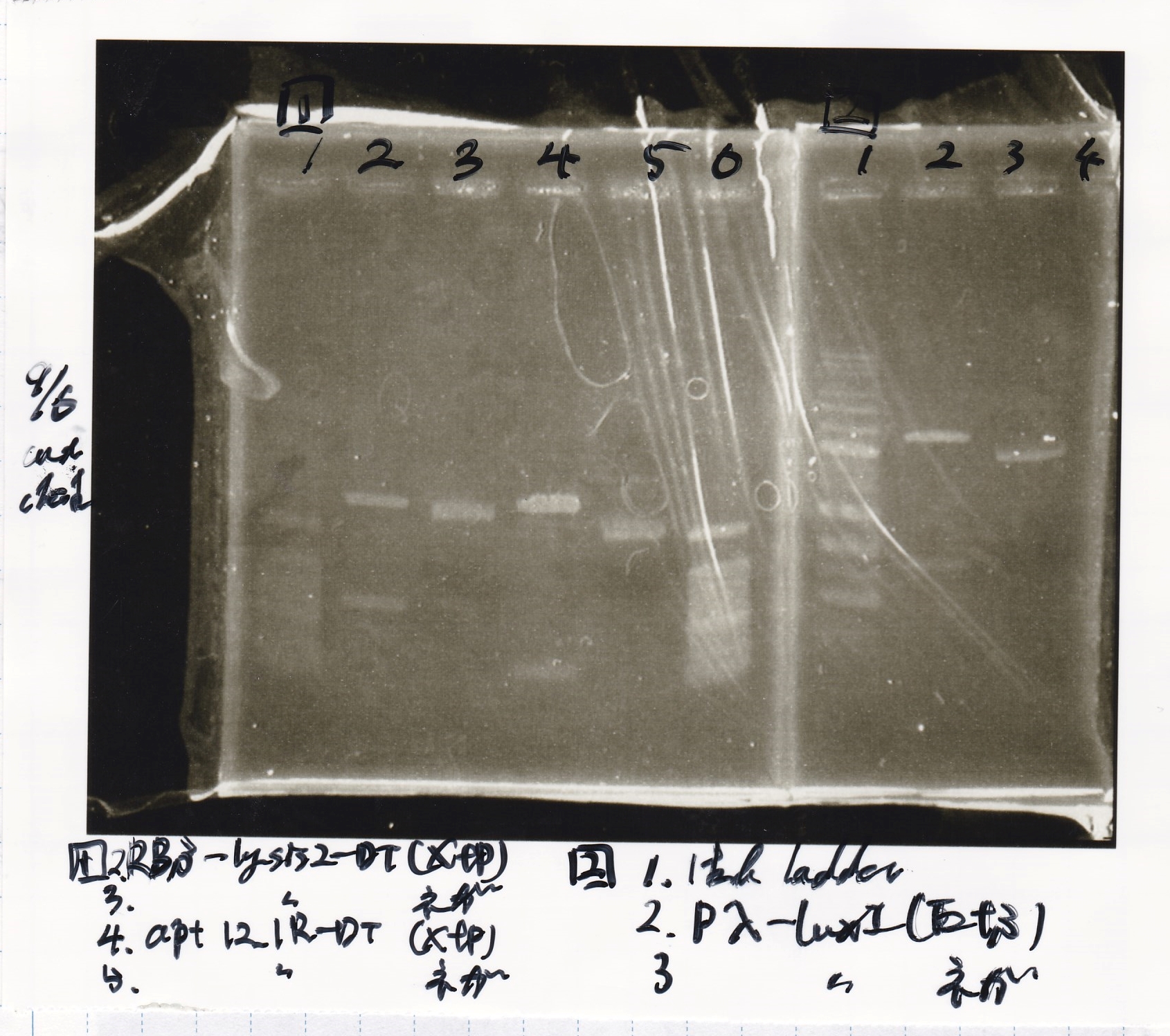
File:Igku xxxxxxx.xxx
No name
| Lane | DNA | Enzyme
|
| 1 | 100bp ladder | --
|
| 2 | RBS-lysis2-DT | XbaI & PstI
|
| 3
|
| 4 | -- | --
|
| 5 | aptamer 12_1R-DT | XbaI & PstI
|
| 6
|
| Lane | DNA | Enzyme
|
| 1 | 100bp ladder | --
|
| 2 | Pλ-luxI | EcoRI & SpeI
|
| 3
|
File:Igku xxxbeforexxxx.xxx
File:Igku xxxafterxxx.xxx
| Name | concentration[µg/mL] | 260/280 | 260/230
|
| RBS-lysis2-DT (XbaI & PstI) | 7.1 | 1.88 | 0.07
|
| aptamer 12_1R-DT (XbaI & PstI) | 15.3 | 1.93 | 0.70
|
| Pλ-luxI (EcoRI & SpeI) | 4.5 | 1.79 | 0.31
|
Liquid Culture
No name
| Sample | medium
|
| 9/5 Pconst-spinach-DT-(1) | Plusgrow medium(+Amp&CP)
|
| 9/5 Ptet-pT181 antisense-(1) | Plusgrow medium(+CP)
|
| 9/5 Pconst+RBS-tetR-DT-(1) | Plusgrow medium(+Amp&CP)
|
| 9/5 Ptet+RBS-lacZα-DT-(1) | Plusgrow medium(+CP)
|
| 9/5 Pcon-pT181 attenuator-(1) | Plusgrow medium(+CP)
|
| 9/4 pT181 antisense-(1) | Plusgrow medium(+CP)
|
| 9/4 aptamer 12_1R-(1) | Plusgrow medium(+CP)
|
Colony PCR
No name
| Sample | base pair
|
| 9/5 Plac-spinach-DT-(5~12) | 563
|
| 9/5 Pcon-antisense-(5~12) | 468
|
| 9/5 Plac-attenuator-(5~12) | 664
|
| 9/5 pT181 antisense+Plac-(1) | -
|
| PreDenature | Denature | Annealing | Extension | cycle
|
| 94°C | 94°C | 55°C | 68°C | --
|
| 5min | 30s | 30s | 42s | 30cycles
|
 "
"

