Team:Paris Saclay/Notebook/August/12
From 2013.igem.org
Contents |
Notebook : August 9
Lab work
A - Aerobic/Anaerobic regulation system
1 - Obtaining Pfnr_RBS-LacZ-Term in PSB1C3
Anaïs, Nadia, XiaoJing
Protocol : digestion
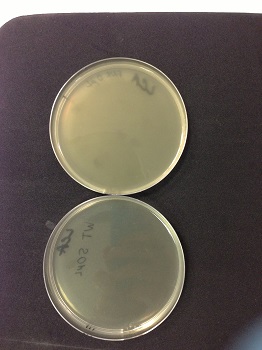
10µl,50µl and 100µl petri dishes are clear so phages are multiplied.
We let the antibiotic over night to select the right strain.
2 - Obtaining RBS_lacZ_ter.
Abdou
Clone 10,14 and 15 plamid extraction using nucleospin kit.
Protocol : hight copy plamid extraction
Results: concentration measured by nanodrop
- clone 10: C=38ng/µl 260/280= 1.78
- clone 14: C=48.5ng/µl 260/280=1.90
- clone 15: C=52 ng/µl 260/280=1.78
We still have to sequence plasmids in order to verify our results.
A - Aerobic/Anaerobic regulation system / B - PCB sensing system
1 - Gibson assembly.
August 1st PCR purification to be sure about the experiment. BphR2 part1, BphR2 part2 and RBS_BphR2_part1, FNR part1, FNR part2 and RBS_FNR_part1.
Protocol : PCR_clean_up
Results:
| IMAGE |
|
we have no fragment so we must do again these PCRs
 "
"