Team:DTU-Denmark/Notebook/20 August 2013
From 2013.igem.org
(→Lab 208) |
|||
| (20 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:DTU-Denmark/Templates/StartPage|20 August 2013}} | {{:Team:DTU-Denmark/Templates/StartPage|20 August 2013}} | ||
| - | |||
Navigate to the [[Team:DTU-Denmark/Notebook/19_August_2013|Previous]] or the [[Team:DTU-Denmark/Notebook/21_August_2013|Next]] Entry | Navigate to the [[Team:DTU-Denmark/Notebook/19_August_2013|Previous]] or the [[Team:DTU-Denmark/Notebook/21_August_2013|Next]] Entry | ||
| - | + | =Lab 208= | |
| - | = | + | |
<hr/> | <hr/> | ||
==Main purpose== | ==Main purpose== | ||
<hr/> | <hr/> | ||
| + | *PCR to linearize pZA21::RFP::araBAD | ||
==Who was in the lab== | ==Who was in the lab== | ||
<hr/> | <hr/> | ||
| + | |||
| + | Kristian, Helen, Henrike, Julia | ||
==Procedure== | ==Procedure== | ||
<hr/> | <hr/> | ||
| + | ===PCR to linearize pZA21::RFP::araBAD=== | ||
| + | |||
| + | *Template: pZA21::RFP::araBAD | ||
| + | *Primers: 55a and 55b | ||
| + | *Program: | ||
| + | {| class="wikitable" style="text-align: right" | ||
| + | ! temperature !! time !! cycles | ||
| + | |- | ||
| + | | 98C || 2:00 || - | ||
| + | |- | ||
| + | | 98C || 0:10 || 36 | ||
| + | |- | ||
| + | | 65C || 1:00 || 36 | ||
| + | |- | ||
| + | | 72C || 3:00 || 36 | ||
| + | |- | ||
| + | | 72C || 5:00 || - | ||
| + | |- | ||
| + | | 10C || hold || - | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | The table shows the composition of each reaction: | ||
| + | {| class="wikitable" style="text-align: right" | ||
| + | ! component (per reaction) !! 1 !! 2 !! 3 !! 4 !! 5 !! 6 !! Neg | ||
| + | |- | ||
| + | | dNTPs || 1uL || 1uL || 1uL || 1uL || 1uL || 1uL || - | ||
| + | |- | ||
| + | | buffer (GC/HF + volume) || HF + 10uL || HF + 10uL || HF + 10uL || GC + 10uL || HF + 10uL || HF + 10uL || HF + 10uL | ||
| + | |- | ||
| + | | X7 polymerase || 0.5uL || 0.5uL || 0.5uL || 0.5uL || 0.5uL || 0.5uL || 0.5uL | ||
| + | |- | ||
| + | | MilliQ water || 29uL || 21.5uL || 31.5uL || 29uL || 31.5uL || 31.5uL || 32.5uL | ||
| + | |- | ||
| + | | template || 1uL || 1uL || 1uL || 1uL || 1uL || 1uL || - | ||
| + | |- | ||
| + | | FW primer 55a || 3uL || 3uL || 3uL || 3uL || 3uL || 3uL || 3uL | ||
| + | |- | ||
| + | | RV primer 55b || 3uL || 3uL || 3uL || 3uL || 3uL || 3uL || 3uL | ||
| + | |- | ||
| + | | DMSO (100%) || 2.5uL (5%) || - || - || 2.5uL (5%) || - || - || - | ||
| + | |- | ||
| + | | Betaine (5M) || - || 10uL (1M) || - || - || - || - || - | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | |||
| + | Repeated reaction mix number 1 and ran on same program to get more backbone. | ||
==Results== | ==Results== | ||
| Line 41: | Line 90: | ||
* 1 kb ladder | * 1 kb ladder | ||
| - | + | [[File:2013-08-20 screen big.jpg|600px]] | |
* 1 kb ladder | * 1 kb ladder | ||
| Line 59: | Line 108: | ||
* 1 kb ladder | * 1 kb ladder | ||
| - | + | [[File:2013-08-20 screen small.jpg|600px]] | |
| + | |||
| + | gel on today's PCR to linearize pZA21::ara | ||
| + | |||
| + | * 1 kb ladder | ||
| + | * sample 1 | ||
| + | * sample 2 | ||
| + | * sample 3 | ||
| + | * sample 4 | ||
| + | * negative controll | ||
| + | * 1 kb ladder | ||
| + | |||
| + | [[File:2013-08-20 test gel pza.jpg|600px]] | ||
| + | |||
| + | Two samples were lost because the lids opened during the PCR reaction and sample evaporated. | ||
| + | |||
| + | ===purification gels=== | ||
| + | |||
| + | Pooled Nir2 and Nir1 samples for Morten Nørholm that gave bands. | ||
| + | |||
| + | * 1 kb ladder | ||
| + | * pZA21::RFP::AraBAD | ||
| + | * pZA21::RFP::AraBAD | ||
| + | * pZA21::RFP::AraBAD | ||
| + | * Nir2 | ||
| + | * Nir2 | ||
| + | * Nir2 | ||
| + | * Nir1 | ||
| + | * Nir1 | ||
| + | * Nir1 | ||
| + | * Nir1 | ||
| + | * 1 kb ladder | ||
| + | |||
| + | [[File:2013-08-20 puri lin pza21ara nir2 nir1.jpg|600px]] | ||
| + | |||
| + | Second purification | ||
| + | |||
| + | * 1 kb ladder | ||
| + | * pZA21::RFP::AraBAD, sample 1 (from todays PCR) | ||
| + | * Nir1 | ||
| + | * Nir1 | ||
| + | * Nir2 | ||
| + | * Nir2 | ||
| + | * Nir2 | ||
| + | * 1 kb ladder | ||
| + | |||
| + | [[File:2013-08-20 puri2 pza nir.jpg|600px]] | ||
==Conclusion== | ==Conclusion== | ||
<hr/> | <hr/> | ||
| + | |||
| + | Something is definitely wrong with our gels. It seems the gradient PCR did not give the right product. Amplification of the linearized pZA21::ara worked. | ||
| + | |||
Navigate to the [[Team:DTU-Denmark/Notebook/19_August_2013|Previous]] or the [[Team:DTU-Denmark/Notebook/21_August_2013|Next]] Entry | Navigate to the [[Team:DTU-Denmark/Notebook/19_August_2013|Previous]] or the [[Team:DTU-Denmark/Notebook/21_August_2013|Next]] Entry | ||
{{:Team:DTU-Denmark/Templates/EndPage}} | {{:Team:DTU-Denmark/Templates/EndPage}} | ||
Latest revision as of 17:46, 28 September 2013
20 August 2013
Contents |
Lab 208
Main purpose
- PCR to linearize pZA21::RFP::araBAD
Who was in the lab
Kristian, Helen, Henrike, Julia
Procedure
PCR to linearize pZA21::RFP::araBAD
- Template: pZA21::RFP::araBAD
- Primers: 55a and 55b
- Program:
| temperature | time | cycles |
|---|---|---|
| 98C | 2:00 | - |
| 98C | 0:10 | 36 |
| 65C | 1:00 | 36 |
| 72C | 3:00 | 36 |
| 72C | 5:00 | - |
| 10C | hold | - |
The table shows the composition of each reaction:
| component (per reaction) | 1 | 2 | 3 | 4 | 5 | 6 | Neg |
|---|---|---|---|---|---|---|---|
| dNTPs | 1uL | 1uL | 1uL | 1uL | 1uL | 1uL | - |
| buffer (GC/HF + volume) | HF + 10uL | HF + 10uL | HF + 10uL | GC + 10uL | HF + 10uL | HF + 10uL | HF + 10uL |
| X7 polymerase | 0.5uL | 0.5uL | 0.5uL | 0.5uL | 0.5uL | 0.5uL | 0.5uL |
| MilliQ water | 29uL | 21.5uL | 31.5uL | 29uL | 31.5uL | 31.5uL | 32.5uL |
| template | 1uL | 1uL | 1uL | 1uL | 1uL | 1uL | - |
| FW primer 55a | 3uL | 3uL | 3uL | 3uL | 3uL | 3uL | 3uL |
| RV primer 55b | 3uL | 3uL | 3uL | 3uL | 3uL | 3uL | 3uL |
| DMSO (100%) | 2.5uL (5%) | - | - | 2.5uL (5%) | - | - | - |
| Betaine (5M) | - | 10uL (1M) | - | - | - | - | - |
Repeated reaction mix number 1 and ran on same program to get more backbone.
Results
Gels
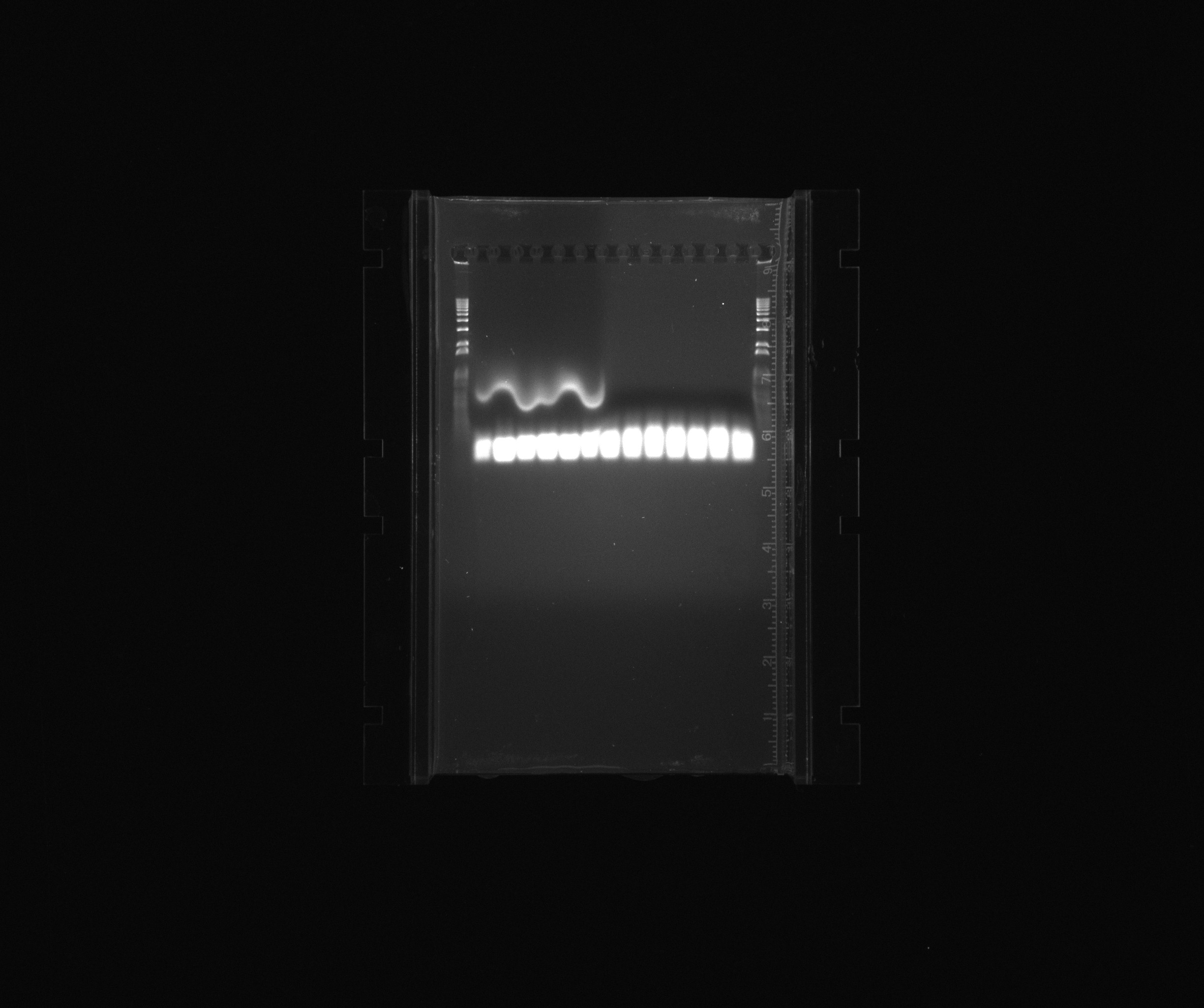
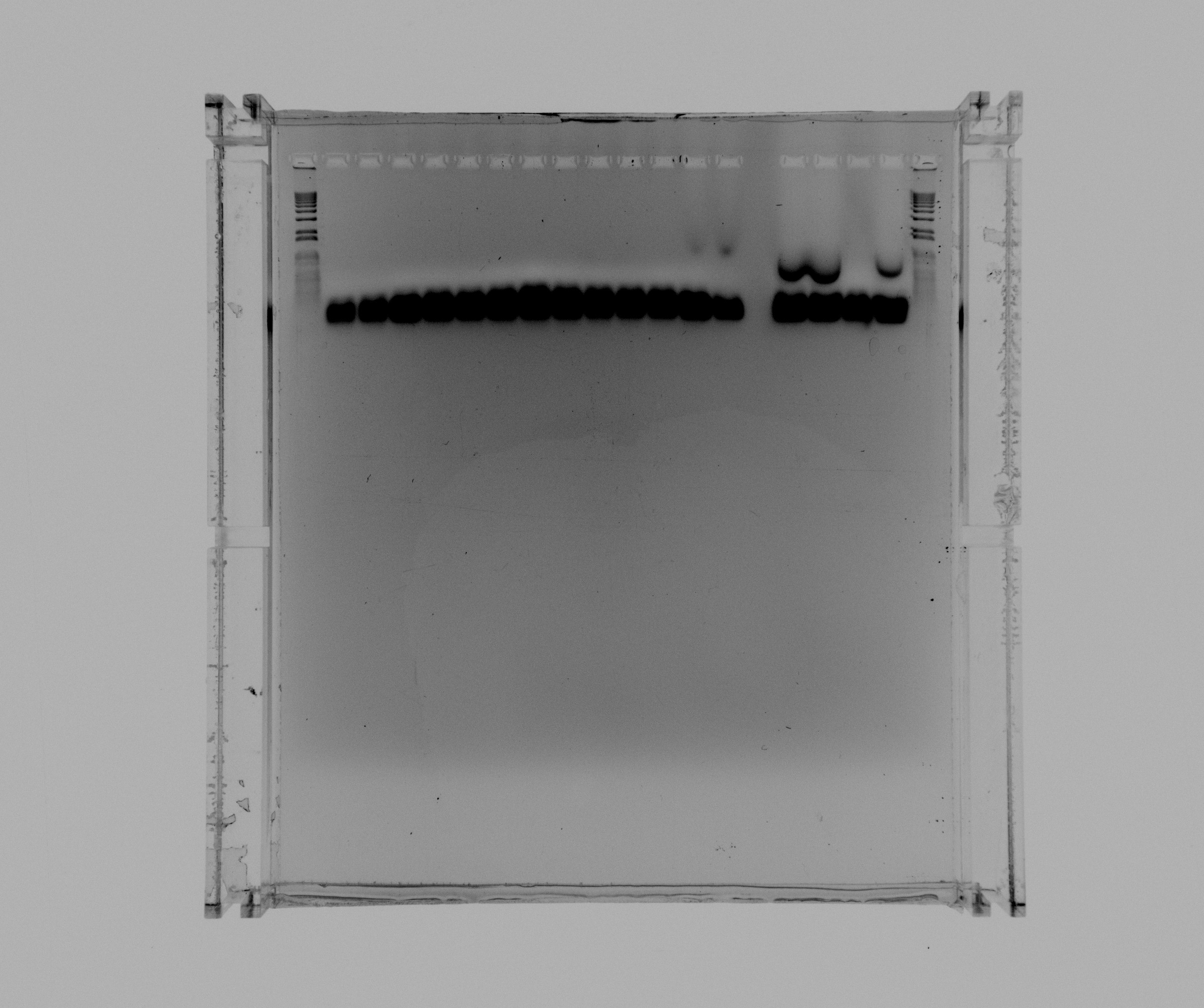
loaded more samples from the screening PCR for BioBrick creation yesterday
- 1 kb ladder
- screening D12
- screening D11
- screening D10
- screening D9
- screening D8
- screening D7
- screening D6
- screening D5
- screening D4
- screening D3
- screening D2
- screening D1
- screening C12
- screening C11
- screening C10
- screening C9
- screening C8
- 1 kb ladder
- 1 kb ladder
- screening C7
- screening C6
- screening C5
- screening C4
- screening C3
- screening C2
- screening C1
- screening B12
- screening B11
- screening B10
- screening B9
- screening B8
- screening B7
- 1 kb ladder
gel on today's PCR to linearize pZA21::ara
- 1 kb ladder
- sample 1
- sample 2
- sample 3
- sample 4
- negative controll
- 1 kb ladder
Two samples were lost because the lids opened during the PCR reaction and sample evaporated.
purification gels
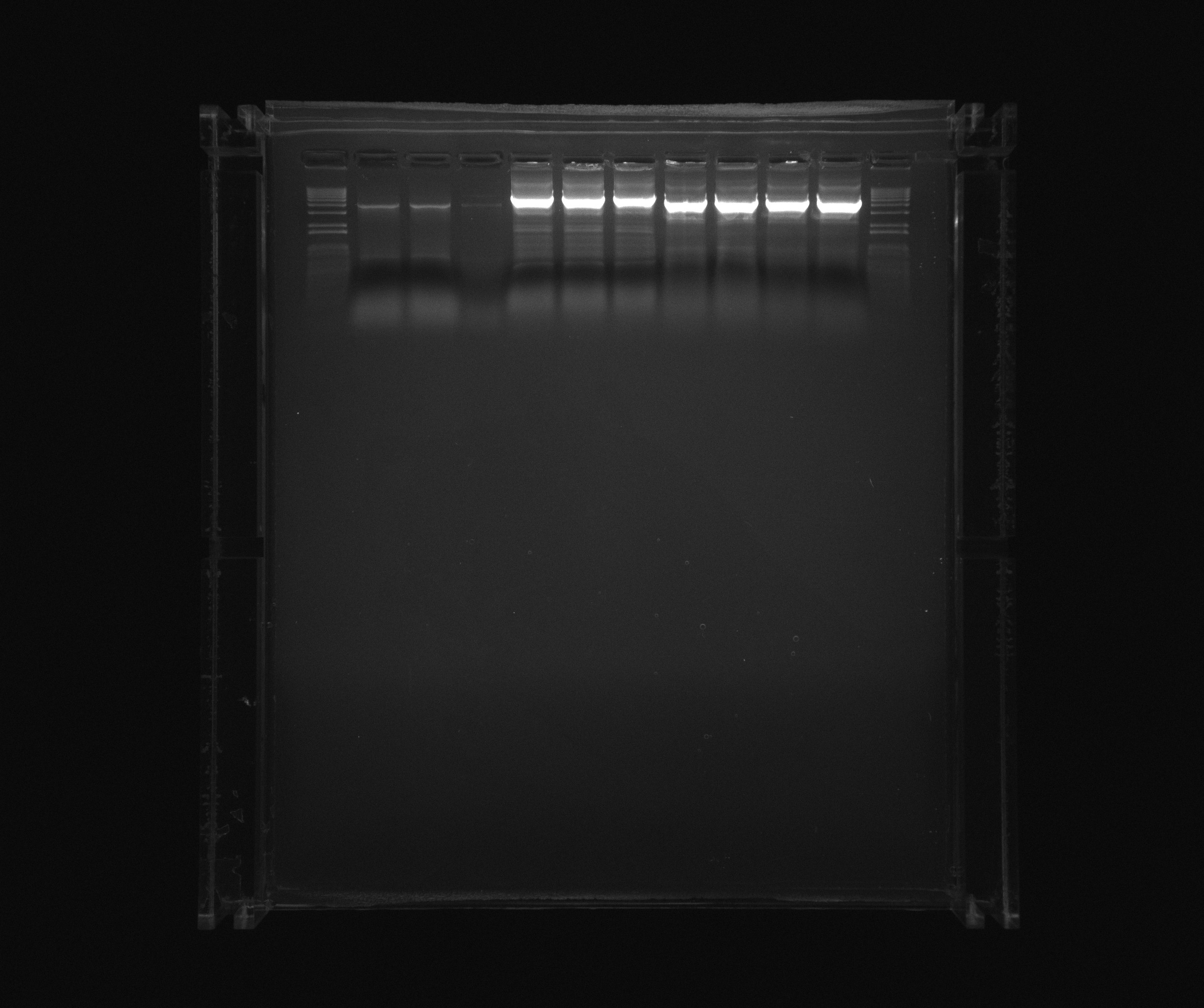
Pooled Nir2 and Nir1 samples for Morten Nørholm that gave bands.
- 1 kb ladder
- pZA21::RFP::AraBAD
- pZA21::RFP::AraBAD
- pZA21::RFP::AraBAD
- Nir2
- Nir2
- Nir2
- Nir1
- Nir1
- Nir1
- Nir1
- 1 kb ladder
Second purification
- 1 kb ladder
- pZA21::RFP::AraBAD, sample 1 (from todays PCR)
- Nir1
- Nir1
- Nir2
- Nir2
- Nir2
- 1 kb ladder
Conclusion
Something is definitely wrong with our gels. It seems the gradient PCR did not give the right product. Amplification of the linearized pZA21::ara worked.
Navigate to the Previous or the Next Entry
 "
"