Team:Evry/Sensor
From 2013.igem.org
Iron Sensor

Construction of the iron-responsive biosensors
E. coli's genome is composed of many Fur binding site. Based on a genome study, we identified 4 promoters which are controled by the FUR protein.
- AceB promoter - (BBa_K1163102)
- Fes promoter - (BBa_K1163108)
- FepA promoter - (BBa_K1163105)
- yncE promoter - (BBa_K1163111)
Using PCR on E. coli genome, we extracted these four promoters. We constructed iron-responsive biosensors by combining 3 genetic parts: an E. coli promoter with a Ferric Uptake Regulator (Fur) binding site, a fluorescent reporter (sfGFP), and a transcriptional terminator (see Figure 1 below). Promoter-reporter fusions were made with flanking restriction sites that are compatible with Biobrick-based cloning.


Fig. 1 Construction of an iron-responsive genetic element by fusing a Fur-regulated promoter with a reporter gene.
We put all the parts together using Golden Gate assembly method. In order to combine our part in the right way, we designed specific overhang as shown in the table 1.
| Name | Figure | Description |
|---|---|---|
|
E. coli promoter with Fur binding site |

|
iron-Fur complex binds promoter to repress expression |
|
sfGFP |

|
Fluorescent reporter gene |
|
Terminator |

|
terminator to stop transcription |
|
Plasmid |

|
Biobrick-compatible plasmid backbone |
Table 1 Genetic elements used to make iron-responsive sensors.
These biosensors respond to ambient iron by using the Fur system to repress the reporter gene placed downstream the promoter. After several experiments we determined that the promoter AceB was the best candidate to create our iron sensitive biosensor.
Caracterisation of the iron-responsive biosensors
As shown in figure 2, in our construction, sfGFP is placed downstream the Fur binding site. It means that in iron starvation sfGFP should be expressed and in high concentration of iron it should be repressed.
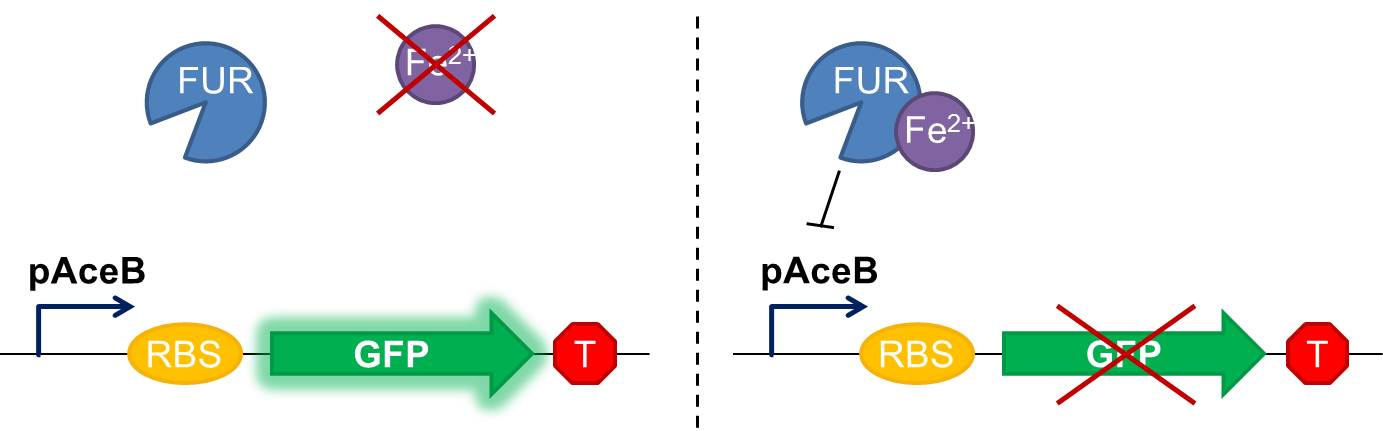
Fig. 2 Diagram of our genetic iron sensor. Iron binds the Ferric Uptake Regulator (Fur) to form a complex with high affinity for the Fur box in the promoter, here shown as the aceB promoter. Once the iron-Fur complex is bound to the promoter, it represses transcription of the target gene GFP. GFP expression is thus negatively correlated with iron availability.
We characterised our construction growing our bacteria in different iron concentrations (0.1 µM, 1 µM and 10 µM). Using 96-wells plate reader, we measured O.D. (600 nm) and GFP intensity (530 nm) each 10 minutes of the bacterial growth.

Fig. 3 The blank is used to calibrate the O.D. measurement. pAceB-LacI is used as a growth control and as a standard for the GFP measurement.
1st step of the analysis
In a first step we checked that our growth was not affected by the difference in iron concentration. As shown in the figure 4, bacterial growth was weakly affected by the iron variation.

Fig. 4 O.D. measurement of bacteria transformed by our system pAceB-GFP, in different concentration of iron.
2nd step of the analysis
In a second step we measured the GFP intensity from our construction, pAceB-GFP, that we standardized using the GFP intensity measured from our second construction, pAceB-LacI. As shown in the figure 5, GFP expression downstream the promoter AceB is expressed in low concentration of iron and its expression repressed with the increase of iron.

Fig. 5 GFP expression is decreased at higher iron concentrations.
3rd step of the analysis
The GFP expression must be compute with the O.D. in order to compare the amount of GFP expressed depending on the iron concentration. Then, we obtained the following results (Figure 6) which show that the increasing of iron leads to a repression of the GFP downstream the promoter AceB.

Fig. 6 GFP expression of the pAceB-GFP Biobrick BBa_K1163102 is repressed at higher iron concentrations. This construct thus functions an an iron-responsive biosensor to repress expression of the reporter gene GFP.
 "
"













