Team:Greensboro-Austin/Standard Proposal
From 2013.igem.org
Mhammerling (Talk | contribs) |
Mhammerling (Talk | contribs) |
||
| Line 33: | Line 33: | ||
As the field of Synthetic Biology emerged, arguments for the standardizing the assembly methods involved in constructing plasmids also emerged. The archetypical example of such standardization is BioBrick RFC[10], introduced in 2003 by Tom Knight at MIT. BioBricks are stored on a standard plasmid, pSB1C3, which contains prefix and suffix sequences flanking the part. These sequences contain two pairs of 6 bp restriction sites (EcoRI+XbaI and SpeI+PstI), which can be used for both part assembly and quality control. BioBricks are intended to be well-characterized biological parts, such as genes or promoters, that are predictable, ready to use, and can be combined in unique ways. The rules of this assembly method also require that none of these sites are present in the parts themselves. This requirement can be an onerous imposition for iGEM teams using large, novel parts such as genes and operons obtained from environmental samples. | As the field of Synthetic Biology emerged, arguments for the standardizing the assembly methods involved in constructing plasmids also emerged. The archetypical example of such standardization is BioBrick RFC[10], introduced in 2003 by Tom Knight at MIT. BioBricks are stored on a standard plasmid, pSB1C3, which contains prefix and suffix sequences flanking the part. These sequences contain two pairs of 6 bp restriction sites (EcoRI+XbaI and SpeI+PstI), which can be used for both part assembly and quality control. BioBricks are intended to be well-characterized biological parts, such as genes or promoters, that are predictable, ready to use, and can be combined in unique ways. The rules of this assembly method also require that none of these sites are present in the parts themselves. This requirement can be an onerous imposition for iGEM teams using large, novel parts such as genes and operons obtained from environmental samples. | ||
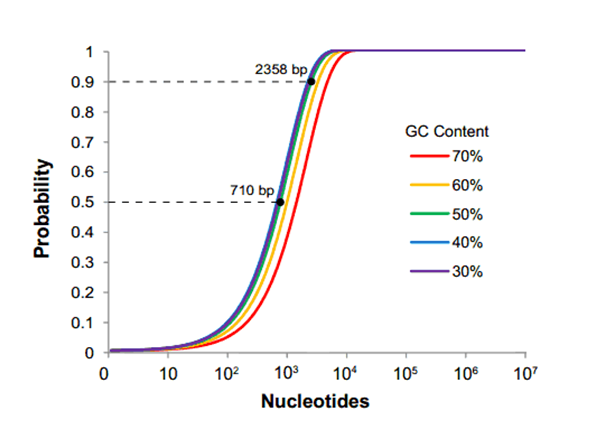
[[File:larry4.png|frame|left|500px|||Fig. 1: The probability of at least one restriction site appearing in a sequence. The length at which a sequence will have a 50% and 90% likelihood of containing a site are displayed for the 50% GC content line.]] | [[File:larry4.png|frame|left|500px|||Fig. 1: The probability of at least one restriction site appearing in a sequence. The length at which a sequence will have a 50% and 90% likelihood of containing a site are displayed for the 50% GC content line.]] | ||
| + | Despite the problems with the current standard, it is clear that having a repository of genetic parts stored in a standard plasmid is useful to the field, especially for those with limited resources. Once these constructs are submitted into a database it is very easy for others to incorporate those parts into their own constructs because of the standardized submission and assembly methods; thus making collaboration much simpler. | ||
| - | + | We concluded that the problem with RFC[10] lies in the fact that it requires everyone who wants to submit a part to the database to accommodate a highly restrictive assembly method. In an era when many fast, cheap, and scarless assembly methods are available, mandating a single assembly method no longer benefits the community. It is easily demonstrable that as the size of your construct increases toward the size of an average eukaryotic gene, the chance of one of the four restriction sites popping up in your sequence is rapidly surpasses 50%. We concluded from this analysis that restriction site mutagenesis likely burdens a majority of teams wishing to submit novel parts to the registry. | |
| - | + | Fixing restriction sites requires both time and money because in order for anyone to utilize your part using the same standard you need to mutate away those unwanted restriction sites in your construct. For IGEM teams who have both limited time and funding, this is a real problem and nonetheless an obstacle for any research group trying to share their work using the Biobrick database. Furthermore, it draws resources away from better characterization of parts - a well-known deficiency of constructs submitted to the registry. Due to these major drawbacks, we speculated that other teams might be having the same issues with regards to part submission and illegal restriction sites. To find out we made a survey asking other teams if they ever come across the same sorts of problems as our team. | |
| - | + | ||
| - | + | ||
==Survey Results== | ==Survey Results== | ||
Revision as of 20:20, 27 September 2013
Contents |
Introduction and Motivation
As the field of Synthetic Biology emerged, arguments for the standardizing the assembly methods involved in constructing plasmids also emerged. The archetypical example of such standardization is BioBrick RFC[10], introduced in 2003 by Tom Knight at MIT. BioBricks are stored on a standard plasmid, pSB1C3, which contains prefix and suffix sequences flanking the part. These sequences contain two pairs of 6 bp restriction sites (EcoRI+XbaI and SpeI+PstI), which can be used for both part assembly and quality control. BioBricks are intended to be well-characterized biological parts, such as genes or promoters, that are predictable, ready to use, and can be combined in unique ways. The rules of this assembly method also require that none of these sites are present in the parts themselves. This requirement can be an onerous imposition for iGEM teams using large, novel parts such as genes and operons obtained from environmental samples.
Despite the problems with the current standard, it is clear that having a repository of genetic parts stored in a standard plasmid is useful to the field, especially for those with limited resources. Once these constructs are submitted into a database it is very easy for others to incorporate those parts into their own constructs because of the standardized submission and assembly methods; thus making collaboration much simpler.
We concluded that the problem with RFC[10] lies in the fact that it requires everyone who wants to submit a part to the database to accommodate a highly restrictive assembly method. In an era when many fast, cheap, and scarless assembly methods are available, mandating a single assembly method no longer benefits the community. It is easily demonstrable that as the size of your construct increases toward the size of an average eukaryotic gene, the chance of one of the four restriction sites popping up in your sequence is rapidly surpasses 50%. We concluded from this analysis that restriction site mutagenesis likely burdens a majority of teams wishing to submit novel parts to the registry.
Fixing restriction sites requires both time and money because in order for anyone to utilize your part using the same standard you need to mutate away those unwanted restriction sites in your construct. For IGEM teams who have both limited time and funding, this is a real problem and nonetheless an obstacle for any research group trying to share their work using the Biobrick database. Furthermore, it draws resources away from better characterization of parts - a well-known deficiency of constructs submitted to the registry. Due to these major drawbacks, we speculated that other teams might be having the same issues with regards to part submission and illegal restriction sites. To find out we made a survey asking other teams if they ever come across the same sorts of problems as our team.
Survey Results
Notable Responses
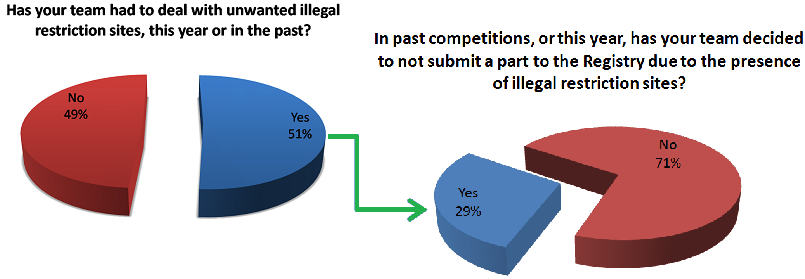
As you can see a significant number of our respondents are experiencing the problem of unwanted restriction sites occurring in their parts and thus affecting the number of new parts that are entered into the registry every year. Some notable responses are also shown below.
"I strongly agree with this statement. Having to perform site-directed mutagenesis in order to submit parts to the Registry is a complete waste of time"
"We need to train our students on the way science is being done now, not back in 1980."
"We started our experiments using restriction-ligation but all of these failed. Next we tried Gibson assembly and were almost immediately successful"
"Taking into account the dropping costs of chemical synthesis of DNA, molecular cloning will not be used for biotechnological purposes in a couple of years. Biobrick-based molecular cloning is obsolete."
The Open Sequence Initiative
After many meetings and brainstorming sessions we decided to propose a standard via an RFC that is intended to replace the current Biobrick standard RFC10 in order to reduce restrictions on parts submitted to the Parts Registry. Crucially, this RFC removes the sequence restrictions imposed by RFC10 regarding the removal of the restriction enzymes sites EcoRI, XbaI, SpeI, and PstI from parts before they are eligible for inclusion in the Parts Registry.
Quality Control and Submission Standard Format
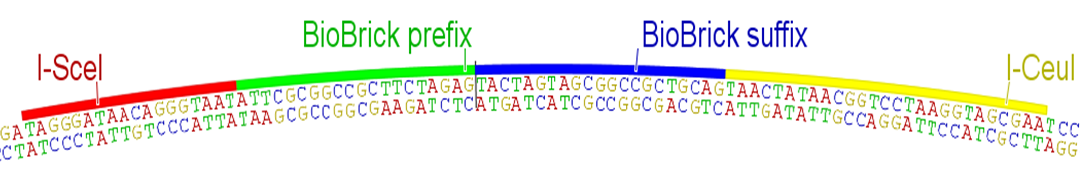
Quality control is critical for any serious engineering endeavor. While teams SHOULD sequence verify their parts to detect point mutations and small indels, the BioBricks Foundation needs a simple and rapid method for determining that parts are at least the correct size when submitted to The Registry. Currently this quality control is achieved through the use of the restriction sites present in the BioBrick prefix and suffix. While using these sites for quality control would technically work with this RFC, internal cut sites would result in complex bands when the digest is ran on a gel. Therefore, we propose using the homing endonucleases I-SceI and I-CeuI and their corresponding restriction enzyme sites for quality control purposes. These sites MUST NOT be contained within the part itself, though the probability of this is negligible for gene-sized or even operon-sized parts
Fig. 2: Proposed order of BioBrick prefix, suffix, and homing endonucleases, as they would appear on the new standard plasmid. The rest of the pSB1C3 backbone remains the same.
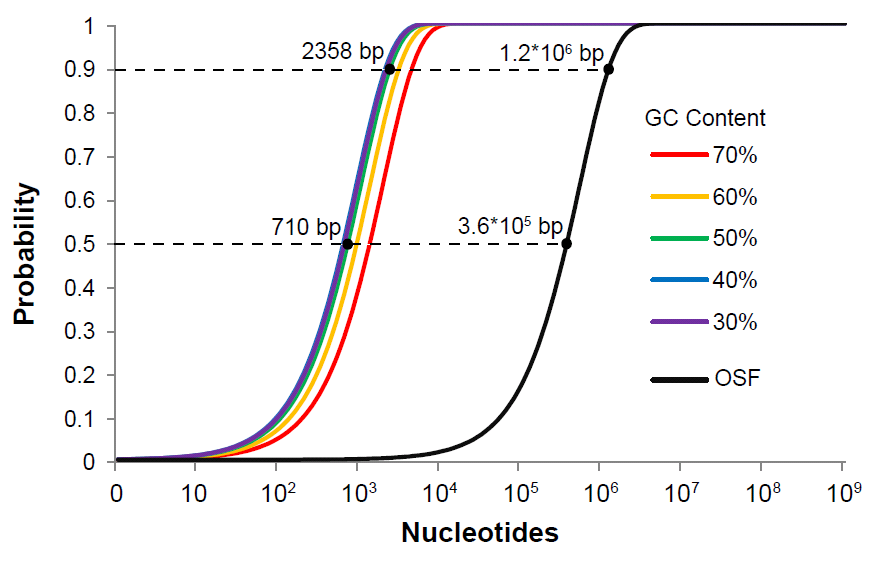
Unlike typical 6 bp restriction sites, these homing endonuclease sites are 18 bp and 27 bp in length, with a tolerance of sequence degeneracy corresponding to a normal restriction site 10 to 12 bp long. However, the probability of even a 10 bp sequence randomly occurring in a submitted part is orders of magnitude lower than a 6 bp site, thus effectively eliminating the problem of illegal sites for the purposes of rapid quality control checking:
 "
"