Team:Heidelberg/Templates/DelH week9
From 2013.igem.org
(Created page with " ==24-06 - 30-06-13 == ===Characterization of DelH plasmid 19-06=== ====SDS Page==== * Using NuPAGE Novex 10% Bis-Tris precast gel from Invitrogen with MOPS running buffer : 3NuP...") |
m (→Result) |
||
| (2 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | ==24-06 - 30-06-13 == | + | == 24-06 - 30-06-13 == |
===Characterization of DelH plasmid 19-06=== | ===Characterization of DelH plasmid 19-06=== | ||
====SDS Page==== | ====SDS Page==== | ||
| Line 11: | Line 11: | ||
* Washed in ddH<sub>2</sub>O multiple times | * Washed in ddH<sub>2</sub>O multiple times | ||
====Result==== | ====Result==== | ||
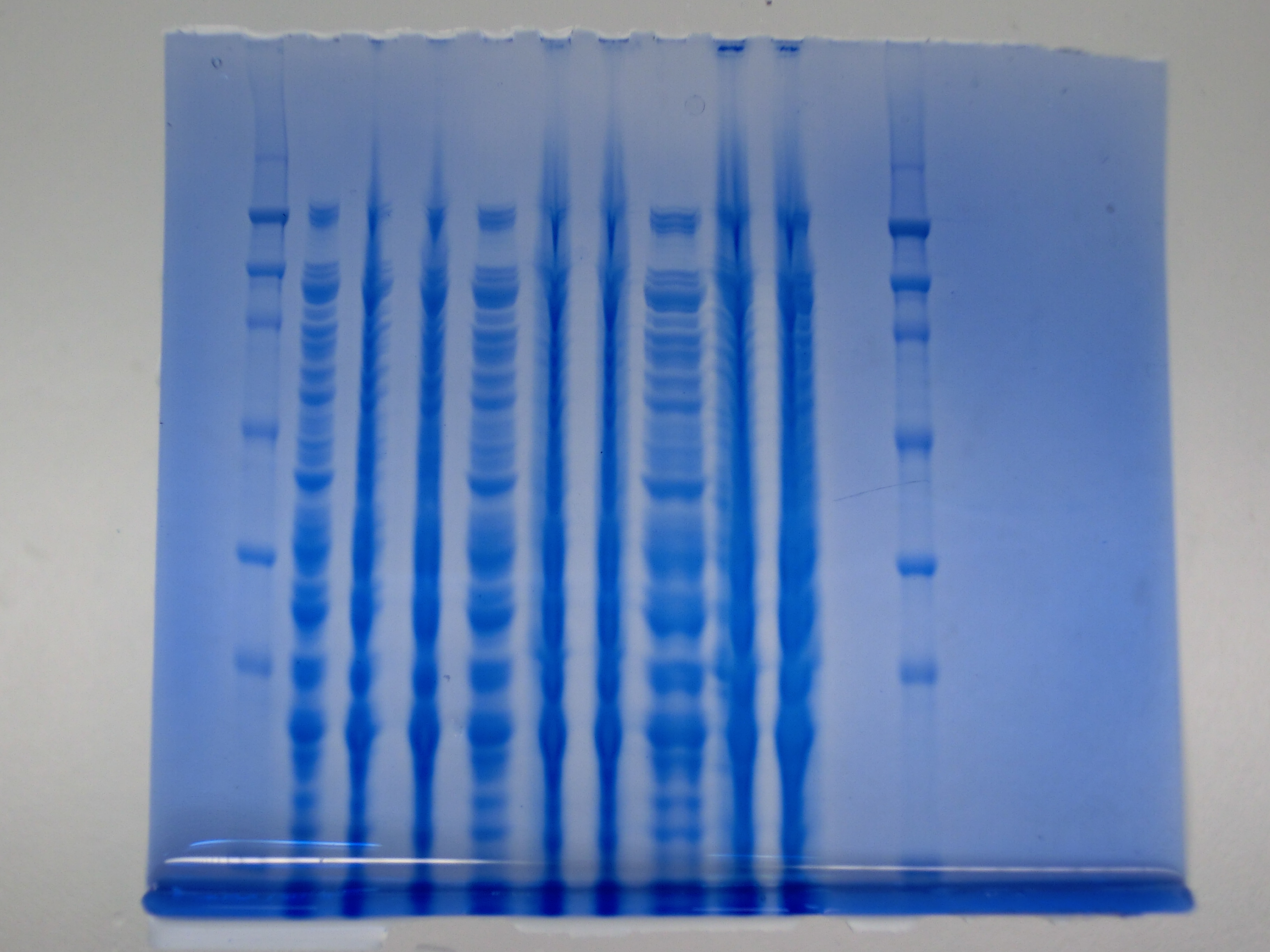
| - | [[File: | + | [[File:Heidelberg_CIMG1018.JPG|thumb|right|'''Fig.9.1''' SDS PAGE <br> ''L1:'' Ladder ''L2:'' 6 µl of Konrad’s ''E. coli'' ''L3:'' 6 µl DelH colony 4 ''L4:'' 6 µl DelH colony 6 ''L5:'' 9 µl of Konrad’s ''E. coli'' ''L6:'' 9 µl DelH colony 4 ''L7:'' 9 µl DelH colony 6 ''L8:'' 15 µl of Konrad’s ''E. coli'' ''L9:'' 15 µl DelH colony 4 ''L10:'' 15 µl DelH colony 6 ''L11:'' empty ''L12:'' Ladder ]] |
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
None clear band at ~600 kDa visible. Maybe in highest amounts, but not reliable. | None clear band at ~600 kDa visible. Maybe in highest amounts, but not reliable. | ||
:=> Probably no DelH expression in analyzed colonies. | :=> Probably no DelH expression in analyzed colonies. | ||
<br/> | <br/> | ||
| + | |||
====Mini Prep==== | ====Mini Prep==== | ||
* Picked colonies from LB Amp plates into 5 ml LB Amp (colonies 4, 6 and 9) | * Picked colonies from LB Amp plates into 5 ml LB Amp (colonies 4, 6 and 9) | ||
| Line 38: | Line 39: | ||
Expected band pattern: 5 kp, 7.3 kp, 13 kp | Expected band pattern: 5 kp, 7.3 kp, 13 kp | ||
<br/> | <br/> | ||
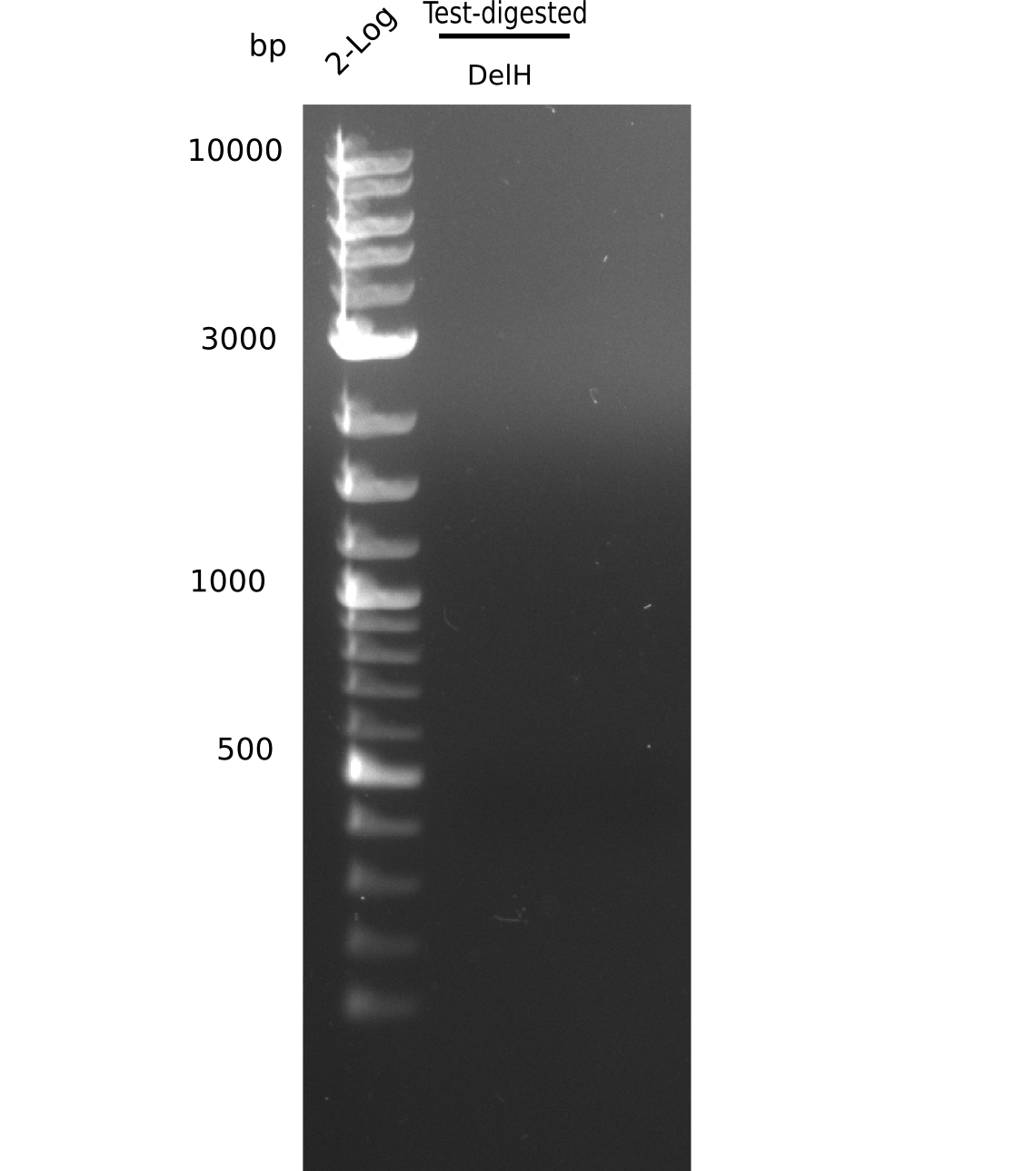
| - | [[File: | + | [[File:Heidelberg_20130626 2log test digested.png|200px|thumb|right|'''Fig.9.2''' Gel of amplified fragments (loaded 20 µL) <br> ''l1:'' 2 log ladder, ''l2:''test digested DelH fragment <br> ''l2:'' no visible band ]] |
| - | + | ||
No clear bands visible. | No clear bands visible. | ||
:=> DNA yield from mini prep was probably too low. | :=> DNA yield from mini prep was probably too low. | ||
| - | + | <div style="clear:both"></div> | |
====DNA Concentration==== | ====DNA Concentration==== | ||
{| class="wikitable" | {| class="wikitable" | ||
| Line 90: | Line 90: | ||
Expected bands: BB (7.3 Kb), DelH F2 (8 Kb), F1a + F1b (10 Kb) | Expected bands: BB (7.3 Kb), DelH F2 (8 Kb), F1a + F1b (10 Kb) | ||
<br/> | <br/> | ||
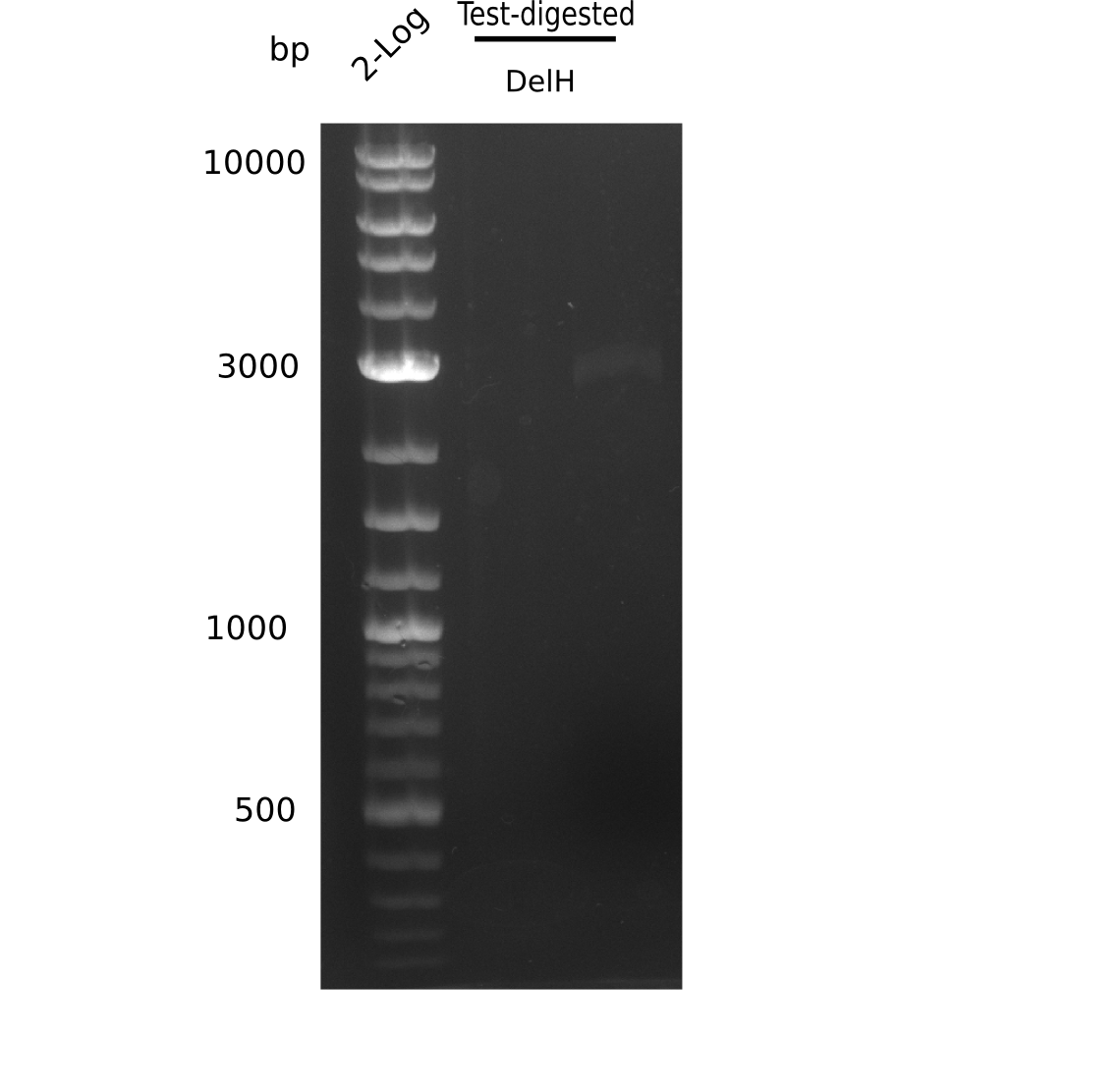
| - | [[File: | + | [[File:Heidelberg_20130627 2log test digested.png|200px|thumb|right|'''Fig.9.3''' miniprep resitriction digested (loaded 20 µL) <br> ''l1:'' 2 log ladder, ''l2:'' DelH-F1a <br> ''l2:'' DelH-F1a shows specific band = cut out ]] |
| - | + | ||
Unexpected bands at 3 Kb, but not desired ones. | Unexpected bands at 3 Kb, but not desired ones. | ||
:=> Colonies do not contain desired plasmid and entire ligation has to be repeated. | :=> Colonies do not contain desired plasmid and entire ligation has to be repeated. | ||
<br/> | <br/> | ||
| - | + | <div style="clear:both"></div> | |
===Amplification of DelH F1b=== | ===Amplification of DelH F1b=== | ||
====PCR Conditions F1b.W9.A==== | ====PCR Conditions F1b.W9.A==== | ||
| Line 133: | Line 133: | ||
Expected band: 5 Kb, Loaded 1 µl of PCR | Expected band: 5 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
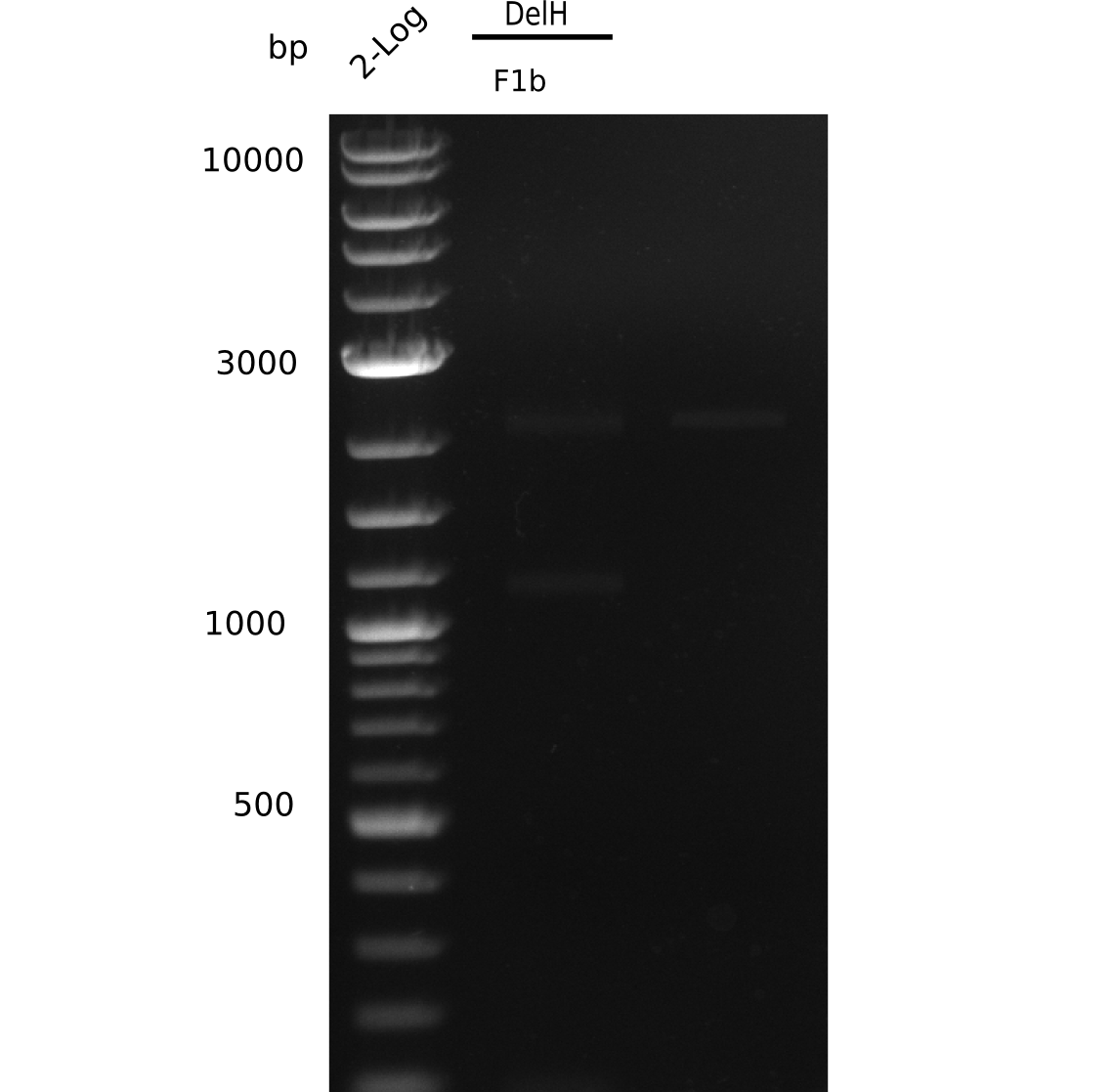
| - | [[File: | + | [[File:Heidelberg_20130627 2log F1b.png|200px|thumb|right|'''Fig.9.4''' gel of amplified DelH 1b - fragment (loaded 20 µL) <br> ''l1:'' 2 log ladder, ''l2:'' DelH-F1b <br> ''l2:'' DelH-F1b shows no band]] |
| - | + | ||
No correct band visible. | No correct band visible. | ||
:=> Why could we not reproduce the previous amplification? | :=> Why could we not reproduce the previous amplification? | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
====PCR Conditions F1b.W9.B==== | ====PCR Conditions F1b.W9.B==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 174: | Line 175: | ||
Expected band: 5 Kb, Loaded 1 µl of PCR | Expected band: 5 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
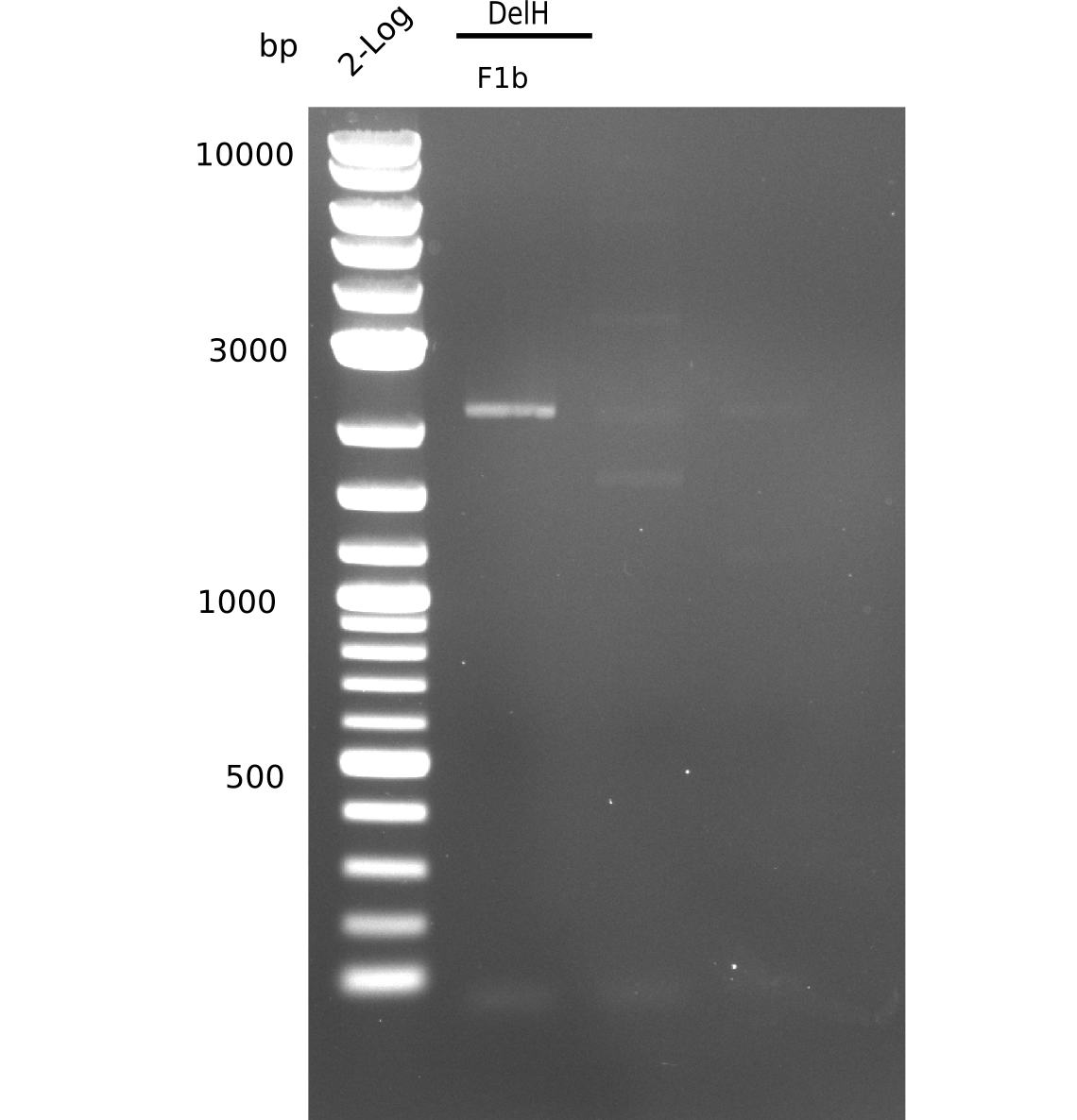
| - | [[File: | + | [[File:Heidelberg_20130627 2log F1bREP.png|200px|thumb|right|'''Fig.9.5''' gel of amplified DelH-1b-fragment (loaded 20 µL) <br> ''l1:'' 2 log ladder, ''l2:'' DelH-F1b <br> ''l2:'' DelH-F1b shows unexpected band at 2.3 Kb]] |
| - | + | ||
Again, expected band not there. | Again, expected band not there. | ||
:=> Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs. | :=> Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
====PCR Conditions F1b.W9.C==== | ====PCR Conditions F1b.W9.C==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 215: | Line 217: | ||
Expected band: 5 Kb, Loaded 1 µl of PCR | Expected band: 5 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
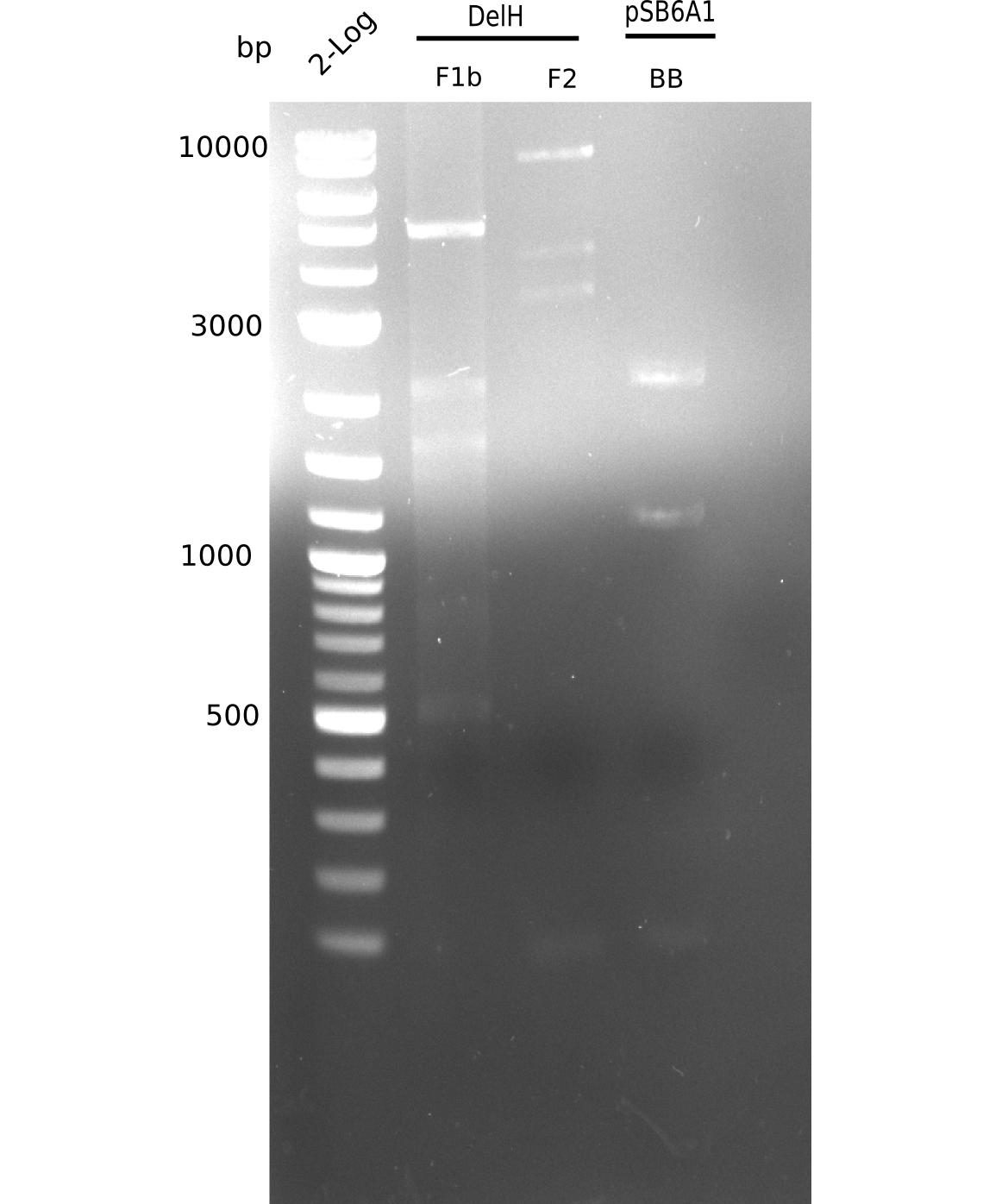
| - | [[File: | + | [[File:Heidelberg_20130628 2log F1b F2 flash all cut.png|200px|thumb|right|'''Fig.9.9''' gel of amplified fragments (loaded 20 µL) <br> ''l1-3:''F1b, ''l4:'' 2log ladder, ''l5-7:'' BB <br> BB was not amplified]] |
| - | + | <div class="tright" style="clear:none">[[File:Heidelberg_20130628 2log F1b F2 flash all.png|200px|thumb|right|'''Fig.9.8''' gel of amplified fragments (loaded 20 µL) <br> ''l1-3:''F1b, ''l4:'' 2log ladder, ''l5-7:'' BB <br> BB was not amplified]]</div> | |
| + | <div class="tright" style="clear:none">[[File:Heidelberg_20130628 2log F1b F2 BB cut.png|200px|thumb|right|'''Fig.9.7''' Gel of amplified DelH-fragments(F1a & F2) and Backbone (pSB6A1-AraC-lacZ) (loaded 20 µL) <br> ''l1:''2log ladder, ''l2:'' F1a, ''l3:''F2, ''l4:'' BB <br> l2: F1a shows specific band at 5 Kb = was cut out]]</div> | ||
| + | <div class="tright" style="clear:none">[[File:Heidelberg_20130628 2log F1b F2 BB.png|200px|thumb|right|'''Fig.9.6''' Gel of amplified DelH-fragments(F1a & F2) and Backbone (pSB6A1-AraC-lacZ) (loaded 20 µL) <br> ''l1:''2log ladder, ''l2:'' F1a, ''l3:''F2, ''l4:'' BB <br> l2: F1a shows specific band at 5 Kb]]</div> | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
Highly specific band at 5 Kb, as well as additional smaller ones. | Highly specific band at 5 Kb, as well as additional smaller ones. | ||
:=> Band was cut. Run gel with remaining sample for gel extraction. | :=> Band was cut. Run gel with remaining sample for gel extraction. | ||
| - | + | ||
| - | + | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
<br/> | <br/> | ||
| Line 261: | Line 265: | ||
Expected band: DelH F1b 5 Kb | Expected band: DelH F1b 5 Kb | ||
<br/> | <br/> | ||
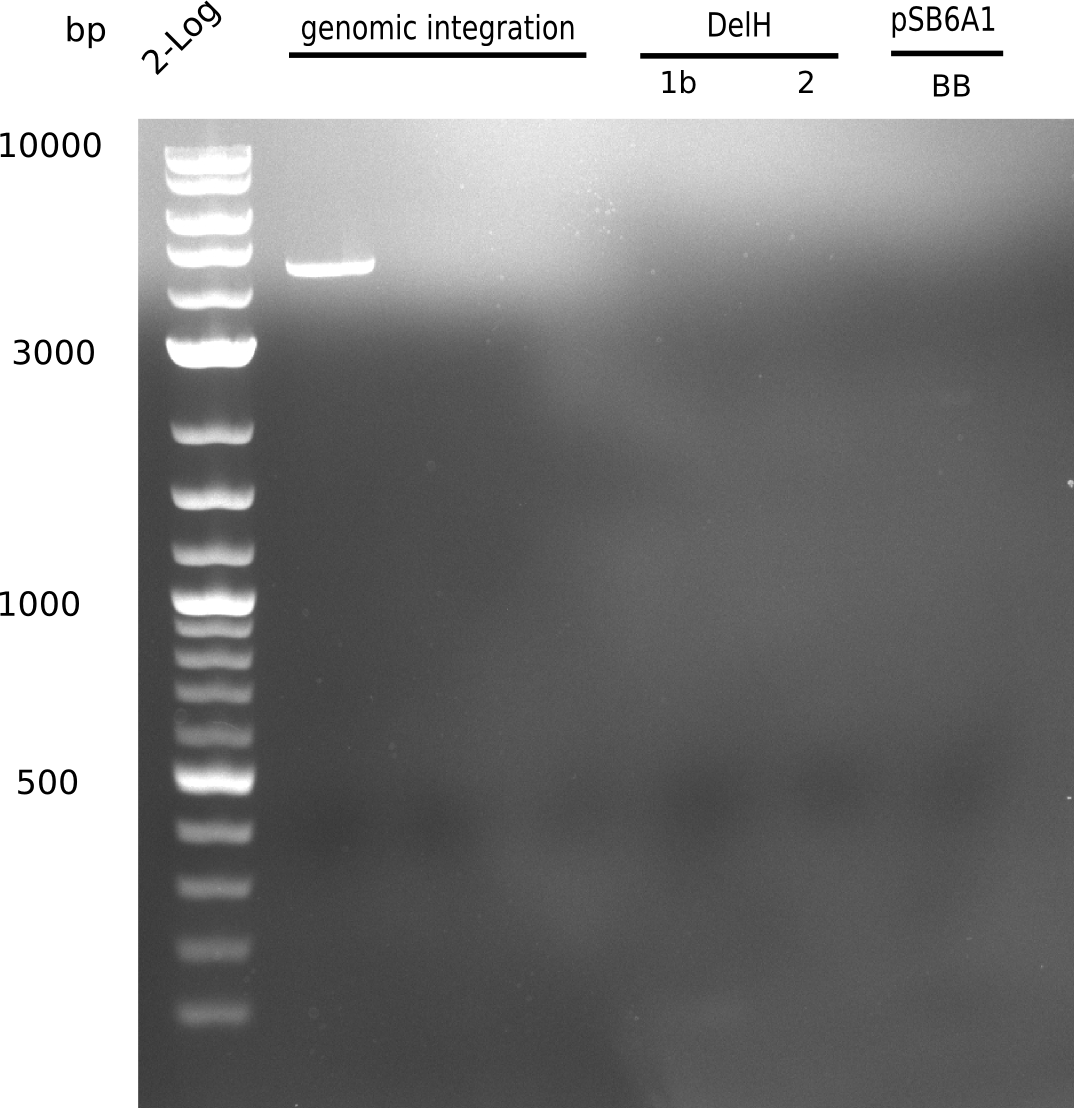
| - | [[File: | + | [[File:Heidelberg_20130628 2log genomicintegration DelH-1 2 BB.png|200px|thumb|right|'''Fig.9.10''' gel of amplified fragments using Q5 (loaded 1 µL) <br> ''l1:''2log ladder, ''l5:'' fragment 1b, ''l6:'' fragment 2, ''l7:'' BB <br> no product]] |
| - | + | ||
Neither of the PCRs yield any product. | Neither of the PCRs yield any product. | ||
:=> Therefore, Q5 is no option for us! | :=> Therefore, Q5 is no option for us! | ||
<br/> | <br/> | ||
| - | + | <div style="clear:both"></div> | |
===Amplification of DelH F2=== | ===Amplification of DelH F2=== | ||
====PCR Conditions F2.W9.A==== | ====PCR Conditions F2.W9.A==== | ||
| Line 304: | Line 308: | ||
Expected band: 8 Kb | Expected band: 8 Kb | ||
<br/> | <br/> | ||
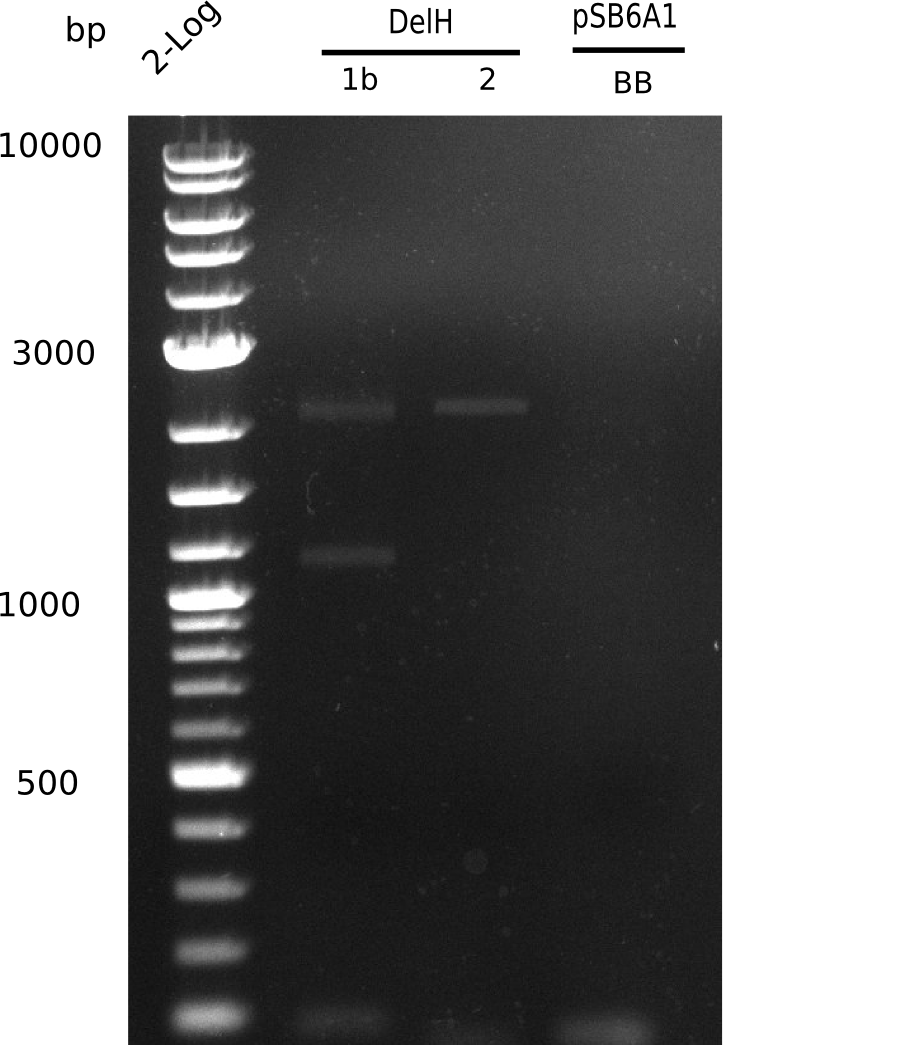
| - | [[File: | + | [[File:Heidelberg_20130627 2log DelH-1b 2 BB 1pcr.png|200px|thumb|right|'''Fig.9.11''' gel of amplified fragments (loaded 1 µL) <br>''l1:'' 2log ladder, ''l2:''F1b, ''l3:'' F2, ''l4:'' BB <br> shows band at 3 Kb]] |
| - | + | ||
Unexpected bands at 3 Kb in lanes of DelH 2 | Unexpected bands at 3 Kb in lanes of DelH 2 | ||
:=> Why could we not reproduce the previous amplification? | :=> Why could we not reproduce the previous amplification? | ||
<br/> | <br/> | ||
| - | + | <div style="clear:both"></div> | |
====PCR Conditions F2.W9.B==== | ====PCR Conditions F2.W9.B==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 352: | Line 356: | ||
Expected band: 8 Kb | Expected band: 8 Kb | ||
<br/> | <br/> | ||
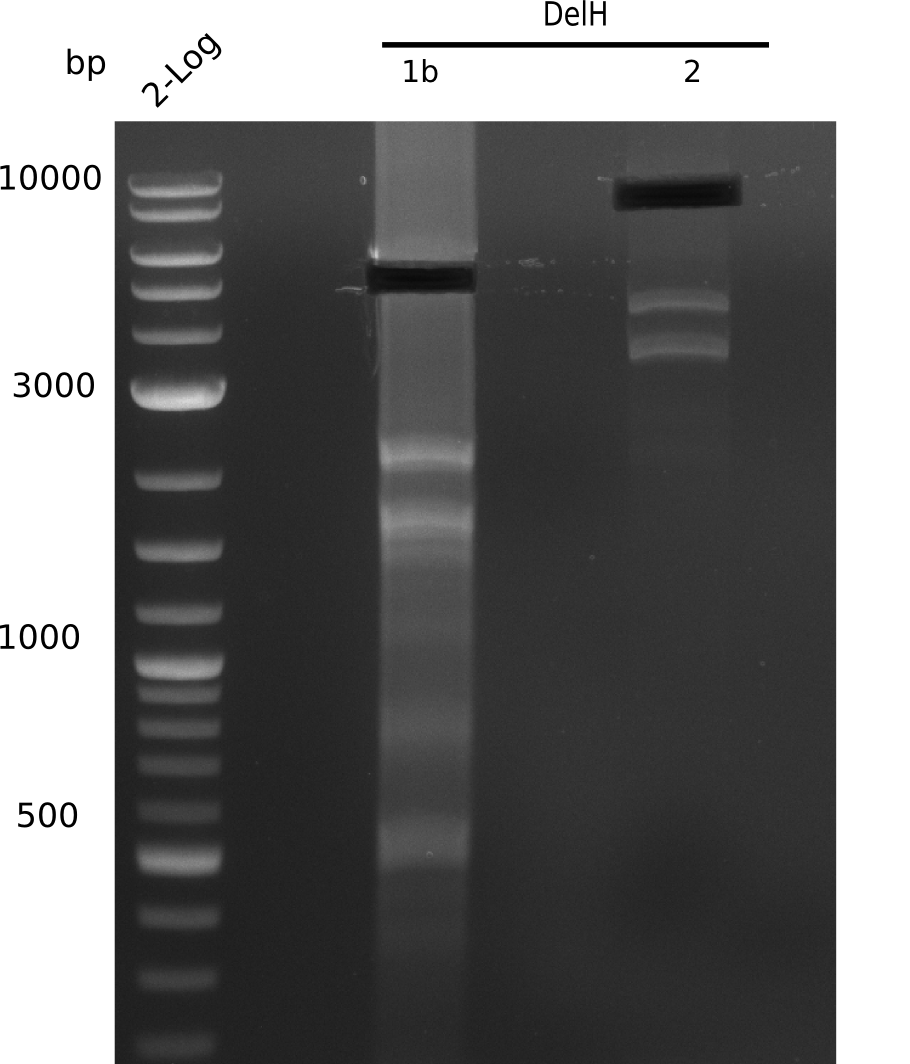
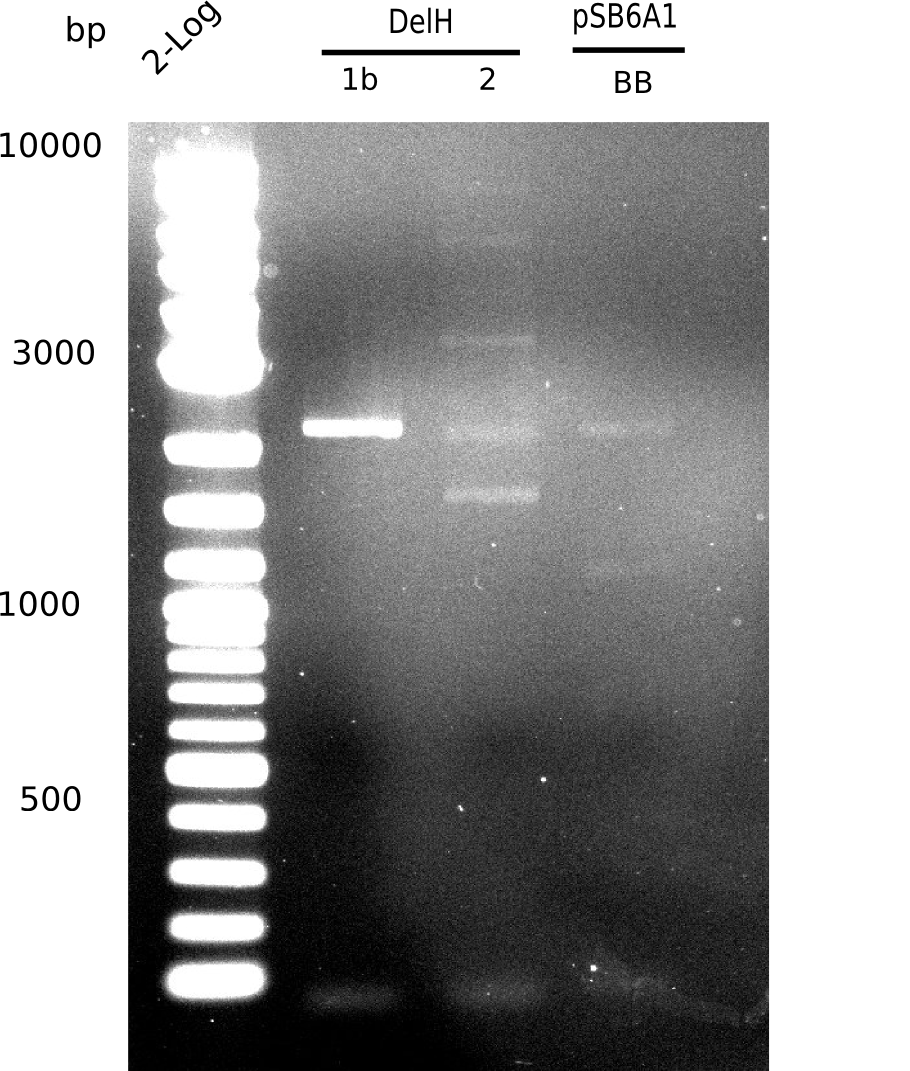
| - | [[File: | + | [[File:Heidelberg_20130627 2log DelH-1b 2 BB.png|200px|thumb|right|'''Fig.9.12''' gel of amplified fragments (loaded 1 µL) <br> ''l1:''2log, ''l2:'' F1b, ''l3:'' F2, ''l4:'' BB <br> F2 shows no specific band]] |
| - | + | ||
Unexpected bands at 1.5 Kb, 2.2 Kb, 4 Kb and 6 Kb. Desired 8 Kb band is present but hardly visible. | Unexpected bands at 1.5 Kb, 2.2 Kb, 4 Kb and 6 Kb. Desired 8 Kb band is present but hardly visible. | ||
:=> Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs. | :=> Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs. | ||
<br/> | <br/> | ||
| - | + | <div style="clear:both"></div> | |
====PCR Conditions F2.W9.C==== | ====PCR Conditions F2.W9.C==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 400: | Line 404: | ||
Expected band: 8 Kb | Expected band: 8 Kb | ||
<br/> | <br/> | ||
| - | [[File: | + | [[File:Heidelberg_20130628 2log F1b F2 flash all cut.png|200px|thumb|right|'''Fig.9.9''' Gel of amplified DelH-fragments(F1a & F2) and Backbone (pSB6A1-AraC-lacZ) (loaded 19 µL) <br> ''l1:''2log ladder, ''l2:'' F1a, ''l3:''F2 <br> l3: F2 shows specific band at 8 Kb and other ones]] |
| - | [[File: | + | <div class="tright" style="clear:none">[[File:Heidelberg_20130628 2log F1b F2 flash all.png|200px|thumb|right|'''Fig.9.8''' Gel of amplified DelH-fragments(F1a & F2) and Backbone (pSB6A1-AraC-lacZ) (loaded 19 µL) <br> ''l1:''2log ladder, ''l2:'' F1a, ''l3:''F2 <br> l3: F2 shows specific band at 8 Kb and other ones]]</div> |
| + | <div class="tright" style="clear:none">[[File:Heidelberg_20130628 2log F1b F2 BB cut.png|200px|thumb|right|'''Fig.9.7''' Gel of amplified DelH-fragments(F1a & F2) and Backbone (pSB6A1-AraC-lacZ) (loaded 1 µL) <br> ''l1:''2log ladder, ''l2:'' F1a, ''l3:''F2, ''l4:'' BB <br> l2: F1a shows specific band at 8 Kb = was cut out]]</div> | ||
| + | <div class="tright" style="clear:none">[[File:Heidelberg_20130628 2log F1b F2 BB.png|200px|thumb|right|'''Fig.9.6''' Gel of amplified DelH-fragments(F1a & F2) and Backbone (pSB6A1-AraC-lacZ) (loaded 1 µL) <br> ''l1:''2log ladder, ''l2:'' F1a, ''l3:''F2, ''l4:'' BB <br> l3: F2 shows specific band at 8 Kb and other ones]]</div> | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
F2 shows specific band at 8 Kb and additional ones. | F2 shows specific band at 8 Kb and additional ones. | ||
:=> Band was cut. Run gel with remaining sample for gel extraction. | :=> Band was cut. Run gel with remaining sample for gel extraction. | ||
| - | |||
| - | |||
| - | |||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
<br/> | <br/> | ||
| Line 453: | Line 456: | ||
Expected band: 8 Kb, Loaded 1 µl of PCR | Expected band: 8 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
| - | [[File: | + | [[File:Heidelberg_20130628 2log genomicintegration DelH-1 2 BB.png|200px|thumb|right|'''Fig.9.10''' gel of amplified fragments using Q5 (loaded 1 µL) <br> ''l1:''2log ladder, ''l5:'' fragment 1b, ''l6:'' fragment 2, ''l7:'' BB <br> no product]] |
| - | + | ||
Neither of the PCRs yield any product. | Neither of the PCRs yield any product. | ||
:=> Therefore, Q5 is no option for us! | :=> Therefore, Q5 is no option for us! | ||
<br/> | <br/> | ||
| - | + | <div style="clear:both"></div> | |
===Amplification of Backbone pSB6A1-AraC-lacZ=== | ===Amplification of Backbone pSB6A1-AraC-lacZ=== | ||
====PCR Conditions BB.W9.A==== | ====PCR Conditions BB.W9.A==== | ||
| Line 497: | Line 500: | ||
Expected band: 7.3 Kb, Loaded 1 µl of PCR | Expected band: 7.3 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
| - | [[File: | + | [[File:Heidelberg_20130627 2log DelH-1b 2 BB 1pcr.png|200px|thumb|right|'''Fig.9.11''' Gel of amplified fragments of DelH and BB(loaded 1 µL) <br> ''l1:''2log ladder, ''l5:'' F1b, ''l6:'' F2, ''l7:'' BB <br> no product]] |
| - | + | ||
No band visible, amplification failed. | No band visible, amplification failed. | ||
:=> Why could we not reproduce the previous amplification? | :=> Why could we not reproduce the previous amplification? | ||
<br/> | <br/> | ||
| - | + | <div style="clear:both"></div> | |
====PCR Conditions BB.W9.B==== | ====PCR Conditions BB.W9.B==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 546: | Line 549: | ||
Expected band: 7.3 Kb, Loaded 1 µl of PCR | Expected band: 7.3 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
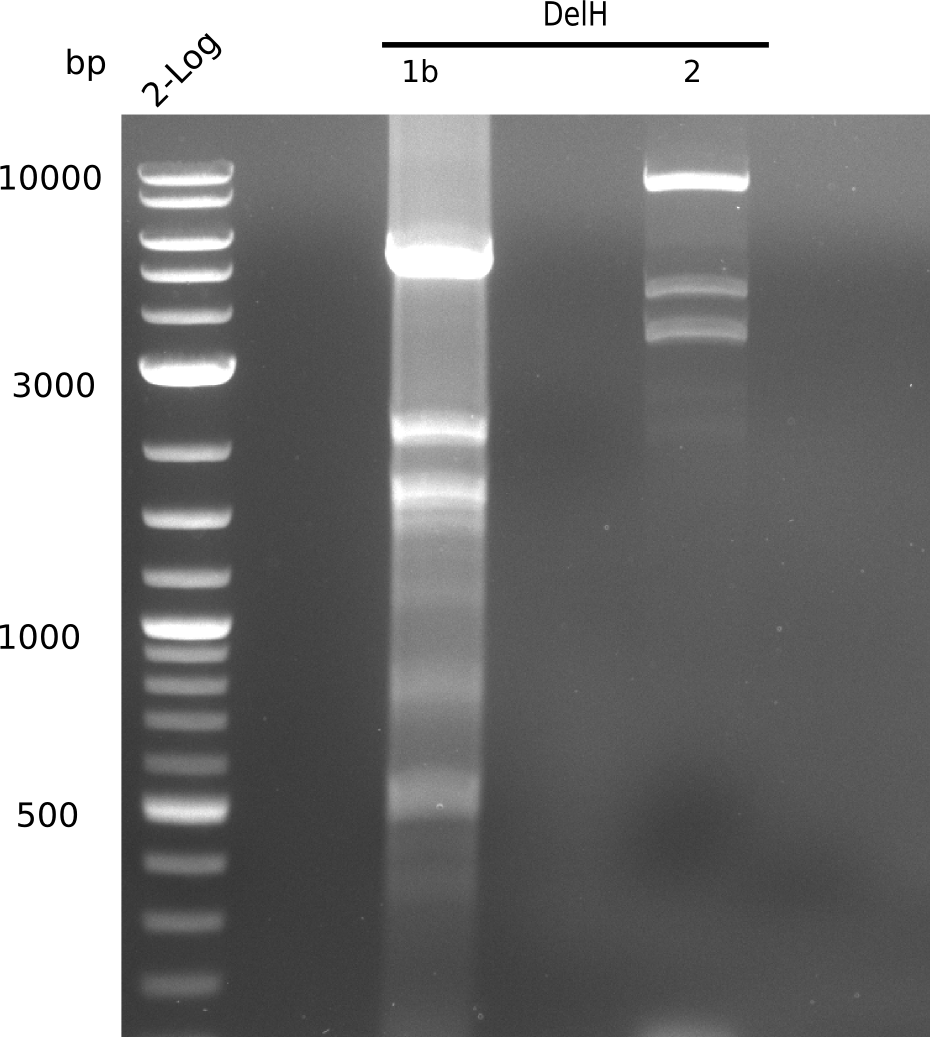
| - | [[File: | + | [[File:Heidelberg_20130627 2log DelH-1b 2 BB.png|200px|thumb|'''Fig. 9.12''' Gel of amplified DelH-fragments and BB ''l1:''2log, ''l2:'' F1b,''l3:''F2, ''l4:''BB]] |
| - | + | ||
Unexpected band at 2.2 Kb | Unexpected band at 2.2 Kb | ||
:=> Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs. | :=> Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs. | ||
<br/> | <br/> | ||
| - | + | <div style="clear:both"></div> | |
====PCR Conditions BB.W9.C==== | ====PCR Conditions BB.W9.C==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 594: | Line 597: | ||
Expected band: 7.3 Kb, Loaded 1 µl of PCR | Expected band: 7.3 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
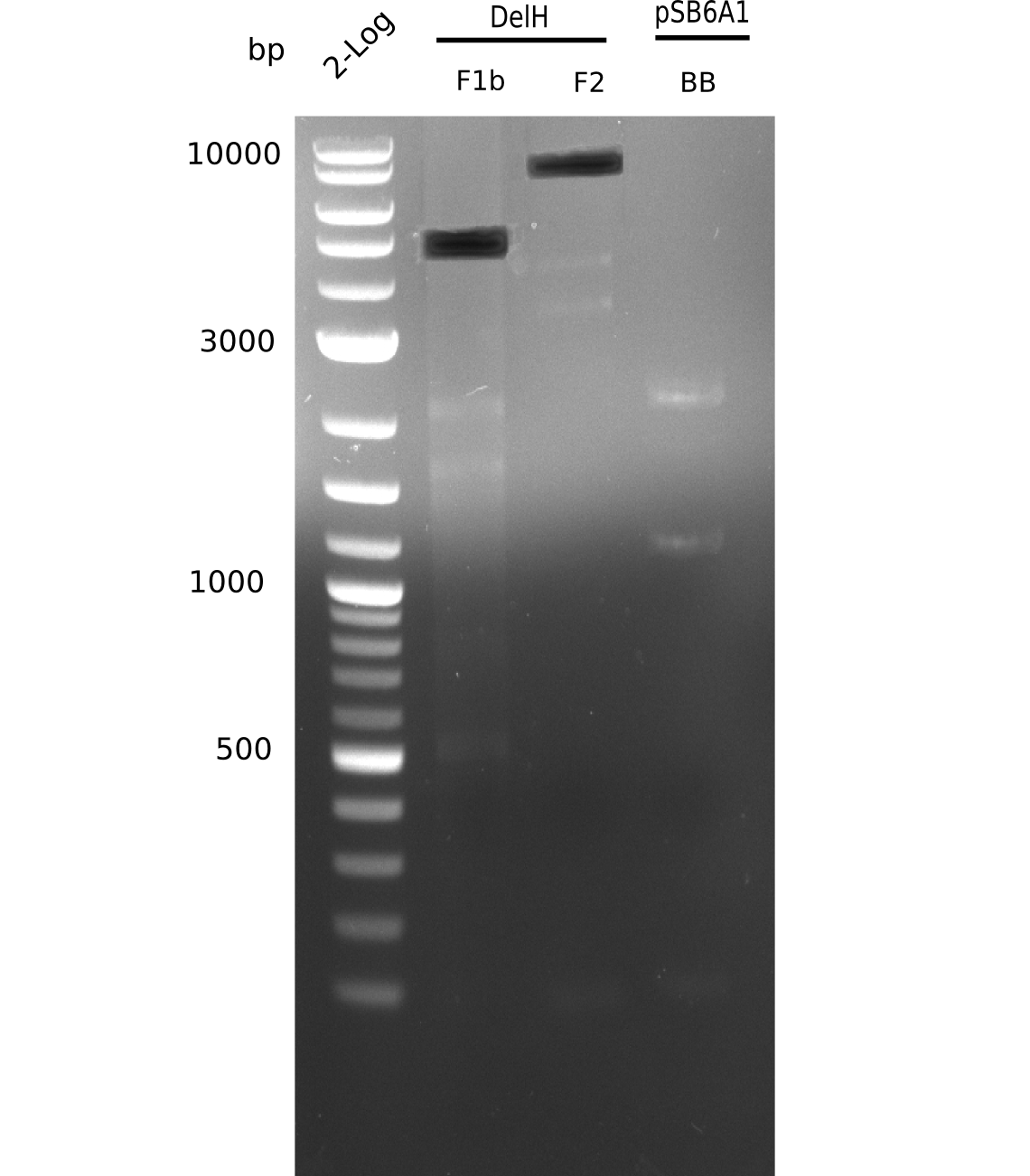
| - | [[File: | + | [[File:Heidelberg_20130628 2log F1b F2 BB.png|200px|thumb|right|'''Fig.9.6''' Gel of amplified DelH-fragments(F1a & F2) and Backbone (pSB6A1-AraC-lacZ) (loaded 20 µL) <br> ''l1:''2log ladder, ''l2:'' F1a, ''l3:''F2, ''l4:'' BB <br> l4: BB shows no expected band]] |
| - | + | ||
Gel does not show expected fragment. | Gel does not show expected fragment. | ||
:=> Make sure right template was used. Is the restricted and purified fragment suitable? | :=> Make sure right template was used. Is the restricted and purified fragment suitable? | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
====PCR Conditions BB.W9.D==== | ====PCR Conditions BB.W9.D==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 641: | Line 645: | ||
Expected band: 7.3 Kb, Loaded 1 µl of PCR | Expected band: 7.3 Kb, Loaded 1 µl of PCR | ||
<br/> | <br/> | ||
| - | [[File: | + | [[File:Heidelberg_20130628 2log genomicintegration DelH-1 2 BB.png|200px|thumb|right|'''Fig.9.10''' gel of amplified fragments using Q5 (loaded 1 µL) <br> ''l1:''2log ladder, ''l5:'' fragment 1b, ''l6:'' fragment 2, ''l7:'' BB <br> no product]] |
| - | + | ||
Neither of the PCRs yield any product. | Neither of the PCRs yield any product. | ||
:=> Therefore, Q5 is no option for us! | :=> Therefore, Q5 is no option for us! | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
Latest revision as of 08:37, 24 October 2013
Contents |
24-06 - 30-06-13
Characterization of DelH plasmid 19-06
SDS Page
- Using NuPAGE Novex 10% Bis-Tris precast gel from Invitrogen with MOPS running buffer
- 3NuPAGE Novex 3-8% Tris-Acetate precast gel would have been more suitable considering expected 600 kDa DelH, but needed Tris-Acetate SDS buffer was not available.
- Resuspended pellets in 100 µl SDS buffer (from Linda)
- Boiled for 10 min @98°C
- Ran for 90 min, 180 V
- Stained for 40 min in Coomassie Buffer (50 ml acetic acid 100% + 225 ml ddH2O, 1.5 g Coomassie in 225 ml methanol, mix both)
- Washed in ddH2O multiple times
Result
None clear band at ~600 kDa visible. Maybe in highest amounts, but not reliable.
- => Probably no DelH expression in analyzed colonies.
Mini Prep
- Picked colonies from LB Amp plates into 5 ml LB Amp (colonies 4, 6 and 9)
- Incubation ON at 37°C
- 5 ml ON cultures were mini prepared and resuspended in 50 µl
Test Restriction Digest
- 5 µl of the mini prep were test digested
| Reagent | Volume [µl] |
|---|---|
| Miniprep DNA | 5 |
| CutSmart buffer | 2 |
| EcoRI-HF, PacI, KpnI-HF | 0.5 each |
| ddH2O | 10.75 |
- Incubation at 37°C for 1 h
Result
Expected band pattern: 5 kp, 7.3 kp, 13 kp
No clear bands visible.
- => DNA yield from mini prep was probably too low.
DNA Concentration
| Colony | DNA concentration [ng/µl] |
|---|---|
| 1 | 124.5 |
| 4 | 3.9 |
| 6 | 1.6 |
| 9 | 10.2 |
DNA measurement confirmed suspicion from test digest.
- =>Thus, mini prep and test digest have to be repeated
Repetion of Mini Prep
- Picked colonies from LB Amp plates into 5 ml LB Amp (colonies 4, 6 and 9) or from remaining ON culture (colony 1)
- Incubation ON at 37°C
- Colony 1 did not grow (also not on LB Amp plate), obviously lost plasmid
- Mini preped remaining 3, eluted in 35 µl ddH2O
DNA Concentration
| Colony | DNA concentration [ng/µl] |
|---|---|
| 4 | 38.5 |
| 6 | 42.0 |
| 9 | 49.0 |
Test Restriction Digest
- Digested entire mini prep (using remaining from first mini for colony 1) for 2 h at37°C
| Reagent | Volume [µl] |
|---|---|
| Mini prep | 42 |
| CutSmart | 5 |
| EcoRI-HF, PacI, KpnI-HF | 1 each |
Result
Expected bands: BB (7.3 Kb), DelH F2 (8 Kb), F1a + F1b (10 Kb)
Unexpected bands at 3 Kb, but not desired ones.
- => Colonies do not contain desired plasmid and entire ligation has to be repeated.
Amplification of DelH F1b
PCR Conditions F1b.W9.A
| Reagent | DelH F1b |
|---|---|
| Template | 1 µl D. acidovorans stock 29-04-13 |
| Primer fw 10 µM | 0.5 µl DelH_EcoRI_fw 100 µM |
| Primer rev 10 µM | 0.5 µl DelH_f1_SalI_rev 100 µM |
| Phusion Ready Mix | 25 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 30 s |
| 35 | 98 | 15 s |
| 68 | 5 s | |
| 72 | 3:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 5 Kb, Loaded 1 µl of PCR
No correct band visible.
- => Why could we not reproduce the previous amplification?
PCR Conditions F1b.W9.B
| Reagent | DelH F1b |
|---|---|
| Template | 1 µl D. acidovorans stock 29-04-13 |
| Primer fw 10 µM | 0.5 µl DelH_EcoRI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl DelH_f1_SalI_rev 10 µM |
| Phusion Ready Mix | 25 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 30 s |
| 30 | 98 | 15 s |
| 68 | 15 s | |
| 72 | 3:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 5 Kb, Loaded 1 µl of PCR
Again, expected band not there.
- => Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs.
PCR Conditions F1b.W9.C
| Reagent | DelH F1b |
|---|---|
| Template | 1 µl DelH F1b Fragment |
| Primer fw 10 µM | 0.5 µl DelH_EcoRI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl DelH_f1_SalI_rev 10 µM |
| Phusion Flash Ready Mix | 10 µl |
| ddH2O | 7 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 10 s |
| 30 | 98 | 1 s |
| 68 | 5 s | |
| 72 | 120 | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 5 Kb, Loaded 1 µl of PCR
Highly specific band at 5 Kb, as well as additional smaller ones.
- => Band was cut. Run gel with remaining sample for gel extraction.
PCR Conditions F1b.W9.D
| Reagent | DelH F1b |
|---|---|
| Template | 1 µl D. acidovorans stock 29-04-13 1:10 |
| Primer fw 10 µM | 0.5 µl DelH_EcoRI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl DelH_f1_SalI_rev 10 µM |
| Q5 Ready Mix | 10 µl |
| ddH2O | 7 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 10 s |
| 30 | 98 | 1 s |
| 68 | 5 s | |
| 72 | 2:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: DelH F1b 5 Kb
Neither of the PCRs yield any product.
- => Therefore, Q5 is no option for us!
Amplification of DelH F2
PCR Conditions F2.W9.A
| Reagent | DelH 2 |
|---|---|
| Template | 1 µl D. acidovorans stock 29-04-13 |
| Primer fw 10 µM | 0.5 µl DelH_f2_SalI_fw 100 µM |
| Primer rev 10 µM | 0.5 µl DelH_f2_KpnI_rev 100 µM |
| Phusion Ready Mix | 25 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 30 s |
| 35 | 98 | 15 s |
| 66 | 5 s | |
| 72 | 3:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 8 Kb
Unexpected bands at 3 Kb in lanes of DelH 2
- => Why could we not reproduce the previous amplification?
PCR Conditions F2.W9.B
| Reagent | DelH 2 |
|---|---|
| Template | 1 µl D. acidovorans stock 29-04-13 |
| Primer fw 10 µM | 0.5 µl DelH_f2_SalI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl DelH_f2_KpnI_rev 10 µM |
| Phusion Ready Mix | 25 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 30 s |
| 12 | 98 | 15 s |
| 66 decrease by 0.5 | 15 s | |
| 72 | 3:30 min | |
| 18 | 98 | 15 s |
| 66 | 15 s | |
| 72 | 3:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 8 Kb
Unexpected bands at 1.5 Kb, 2.2 Kb, 4 Kb and 6 Kb. Desired 8 Kb band is present but hardly visible.
- => Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs.
PCR Conditions F2.W9.C
| Reagent | DelH 2 |
|---|---|
| Template | 1 µl D. acidovorans stock 29-04-13 1:10 |
| Primer fw 10 µM | 0.5 µl DelH_f2_SalI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl DelH_f2_KpnI_rev 10 µM |
| Phusion Flash Ready Mix | 10 µl |
| ddH2O | 7 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 10 s |
| 12 | 98 | 1 s |
| 66 decrease by 0.5 | 5 s | |
| 72 | 2:30 min | |
| 18 | 98 | 1 s |
| 66 | 5 s | |
| 72 | 2:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 8 Kb
F2 shows specific band at 8 Kb and additional ones.
- => Band was cut. Run gel with remaining sample for gel extraction.
PCR Conditions F2.W9.D
| Reagent | DelH 2 |
|---|---|
| Template | 1 µl D. acidovorans stock 29-04-13 1:10 |
| Primer fw 10 µM | 0.5 µl DelH_f2_SalI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl DelH_f2_KpnI_rev 10 µM |
| Q5 Ready Mix | 10 µl |
| ddH2O | 7 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 10 s |
| 12 | 98 | 1 s |
| 66 decrease by 0.5 | 5 s | |
| 72 | 2:30 min | |
| 18 | 98 | 1 s |
| 66 | 5 s | |
| 72 | 2:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 8 Kb, Loaded 1 µl of PCR
Neither of the PCRs yield any product.
- => Therefore, Q5 is no option for us!
Amplification of Backbone pSB6A1-AraC-lacZ
PCR Conditions BB.W9.A
| Reagent | Backbone |
|---|---|
| Template | 1 µl Backbone (pSB6A1-AraC-lacZ c = 24 ng/µl) |
| Primer fw 10 µM | 0.5 µl AraCbb_KpnI_fw 100 µM |
| Primer rev 10 µM | 0.5 µl AraCbbPacI_rev2 100 µM |
| Phusion Ready Mix | 25 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 30 s |
| 35 | 98 | 15 s |
| 66 | 5 s | |
| 72 | 3:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 7.3 Kb, Loaded 1 µl of PCR
No band visible, amplification failed.
- => Why could we not reproduce the previous amplification?
PCR Conditions BB.W9.B
| Reagent | BB |
|---|---|
| Template | 1 µl Backbone (pSB6A1-AraC-lacZ c = 24 ng/µl) |
| Primer fw 10 µM | 0.5 µl AraCbb_KpnI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl AraCbbPacI_rev2 10 µM |
| Phusion Ready Mix | 25 µl |
| ddH2O | 23 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 30 s |
| 12 | 98 | 15 s |
| 66 decrease by 0.5 | 15 s | |
| 72 | 3:30 min | |
| 18 | 98 | 15 s |
| 66 | 15 s | |
| 72 | 3:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 7.3 Kb, Loaded 1 µl of PCR
Unexpected band at 2.2 Kb
- => Adapt PCR conditions and ask for polymerase used last time, which is the one from the lab upstairs.
PCR Conditions BB.W9.C
| Reagent | DelH 2 |
|---|---|
| Template | 1 µl pSB6A1-AraC-LacZ digested and purified 1:10 |
| Primer fw 10 µM | 0.5 µl AraCbb_KpnI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl AraCbbPacI_rev2 10 µM |
| Phusion Flash Ready Mix | 10 µl |
| ddH2O | 7 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 10 s |
| 12 | 98 | 1 s |
| 66 decrease by 0.5 | 5 s | |
| 72 | 2:30 min | |
| 18 | 98 | 1 s |
| 66 | 5 s | |
| 72 | 2:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 7.3 Kb, Loaded 1 µl of PCR
Gel does not show expected fragment.
- => Make sure right template was used. Is the restricted and purified fragment suitable?
PCR Conditions BB.W9.D
| Reagent | DelH 2 |
|---|---|
| Template | 1 µl pSB6A1-AraC-LacZ digested and purified 1:10 |
| Primer fw 10 µM | 0.5 µl AraCbb_KpnI_fw 10 µM |
| Primer rev 10 µM | 0.5 µl AraCbbPacI_rev2 10 µM |
| Q5 Ready Mix | 10 µl |
| ddH2O | 7 µl |
| Cycles | Temperature [°C] | Time |
|---|---|---|
| 1 | 98 | 10 s |
| 12 | 98 | 1 s |
| 66 decrease by 0.5 | 5 s | |
| 72 | 2:30 min | |
| 18 | 98 | 1 s |
| 66 | 5 s | |
| 72 | 2:30 min | |
| 1 | 72 | 10 min |
| 1 | 4 | inf |
- Using hot start at 98°C
Result
Expected band: 7.3 Kb, Loaded 1 µl of PCR
Neither of the PCRs yield any product.
- => Therefore, Q5 is no option for us!
 "
"