Team:Heidelberg/Templates/MM week21p
From 2013.igem.org
(Difference between revisions)
Nils.kurzawa (Talk | contribs) m (Created page with "== 2013-09-16 == * send pIK8.6, pIK8.9 to sequencing with primer RB43 == 2013-09-17 == * sequences arrived: PIK8.6-pIK8.9-2013-09-16.zip, ...") |
|||
| Line 3: | Line 3: | ||
== 2013-09-17 == | == 2013-09-17 == | ||
| - | * sequences arrived: [[:File: | + | * sequences arrived: [[:File:Heidelberg_PIK8.6-pIK8.9-2013-09-16.zip|PIK8.6-pIK8.9-2013-09-16.zip]], [[:File:Heidelberg_PIK8.6-2013-09-16 RB43.clustal.txt|PIK8.6-2013-09-16 RB43.clustal.txt]], [[:File:Heidelberg_PIK8.9-2013-09-16 RB43.clustal.txt|File:Heidelberg_PIK8.9-2013-09-16 RB43.clustal.txt]] |
* no mutations at primer site | * no mutations at primer site | ||
* sequence with VR primer | * sequence with VR primer | ||
| Line 9: | Line 9: | ||
== 2013-09-18 == | == 2013-09-18 == | ||
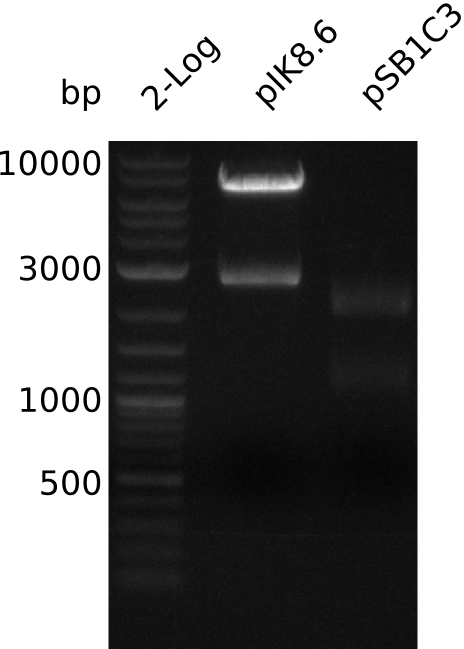
| - | [[File: | + | [[File:Heidelberg_Methylmalonyl-digest2013-09-18.png|300px|thumb|left|Lane 1: NEB 2-log; lane 2: pIK8.6 digested with EcoRI+SpeI, lane 3: pSB1C3 digested with EcoRI+SpeI]] |
| - | * sequences arrived: [[:File: | + | * sequences arrived: [[:File:Heidelberg_PIK8.6-pIK8.9-2013-09-18.zip|PIK8.6-pIK8.9-2013-09-18.zip]], [[:File:Heidelberg_PIK8.6-2013-09-18 VR.clustal.txt|PIK8.6-2013-09-18 VR.clustal.txt]], [[:File:Heidelberg_PIK8.9-2013-09-18 VR.clustal.txt|File:Heidelberg_PIK8.9-2013-09-18 VR.clustal.txt]] |
* sequences look good | * sequences look good | ||
* make miniPrep of pIK8.6 -> 269 ng/µl in 38 µl | * make miniPrep of pIK8.6 -> 269 ng/µl in 38 µl | ||
| Line 35: | Line 35: | ||
== 2013-09-19 == | == 2013-09-19 == | ||
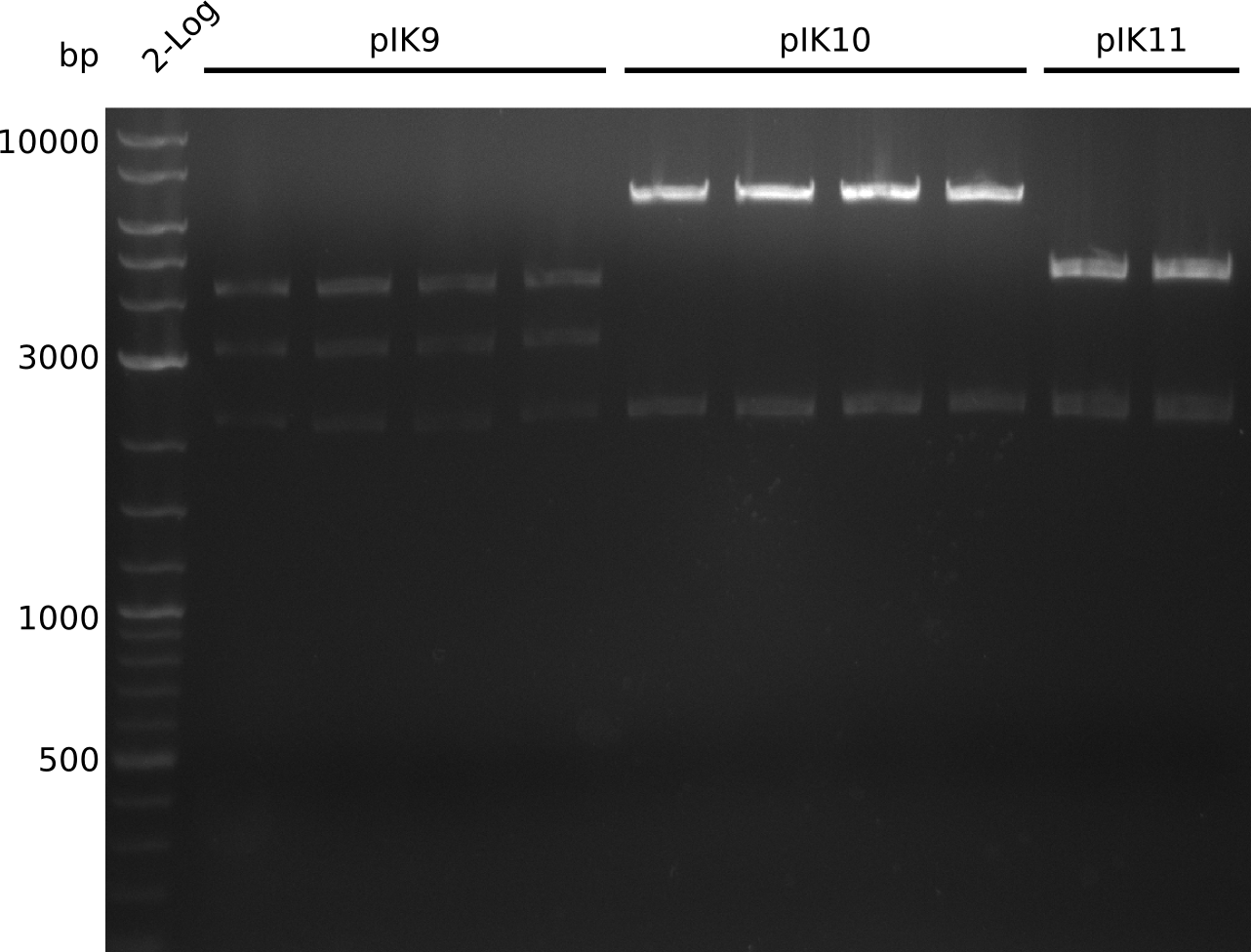
| - | [[File: | + | [[File:Heidelberg_Methylmalonyl-colony-PCR2013-09-19.png|300px|thumb|right|Lane 1: NEB 2-log; lanes 2-5: pIK9; lanes 6-9: pIK10; lanes 10-13: pIK11]] |
* colonies present for all ligations | * colonies present for all ligations | ||
* colonies for pIK10 extremely small -> permeability device toxic | * colonies for pIK10 extremely small -> permeability device toxic | ||
| Line 64: | Line 64: | ||
* to distinguish between pSB3C5 and pSB1C3: digest pIK10, pIK11 with XhoI (expected: 1 band for pSB3C5, 3 bands for pSB1C3) | * to distinguish between pSB3C5 and pSB1C3: digest pIK10, pIK11 with XhoI (expected: 1 band for pSB3C5, 3 bands for pSB1C3) | ||
<gallery widths=300px heights=300px> | <gallery widths=300px heights=300px> | ||
| - | File: | + | File:Heidelberg_Methylmalonyl-digest2013-09-20_1.png|All digested with BamHI+HindIII. Lane 1: NEB 2-log; lanes 2-5: pIK9; lanes 6-9: pIK10; lanes 10-11: pIK11 |
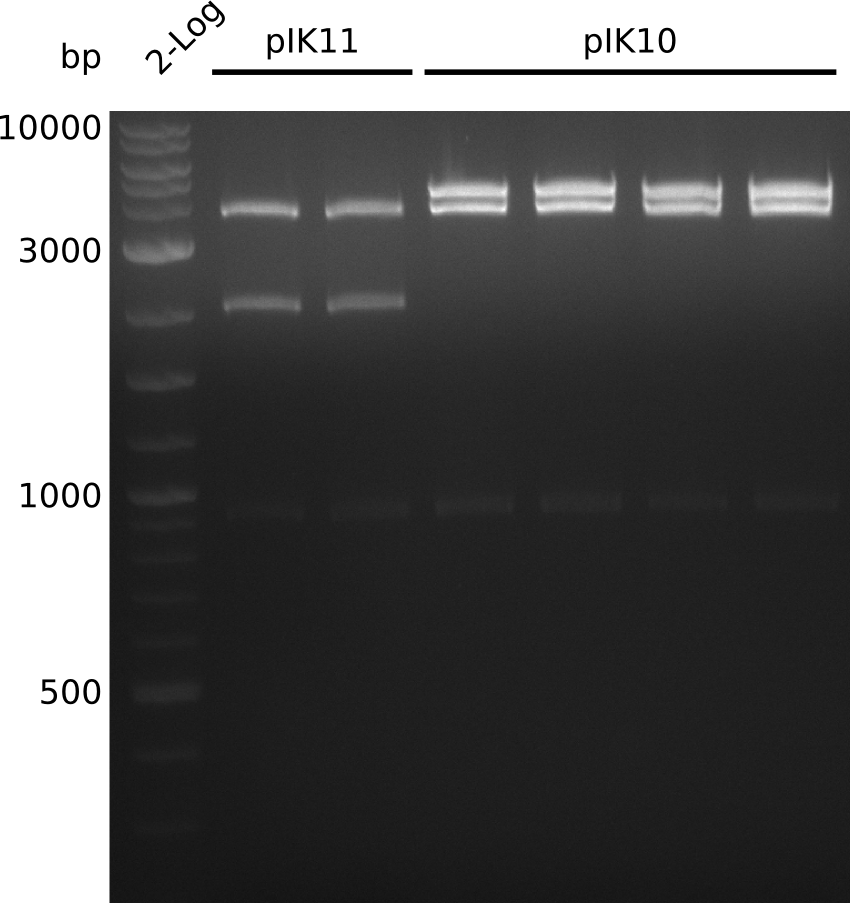
| - | File: | + | File:Heidelberg_Methylmalonyl-digest2013-09-20_2.png|All digested with XhoI. Lane 1: NEB 2-log; lanes 2-3: pIK11; lanes 4-7: pIK10. |
</gallery> | </gallery> | ||
* send pIK10.1, pIK10.3, pIK10.3, pIK11.1, pIK11.2 to sequencing with primers VF2, VR | * send pIK10.1, pIK10.3, pIK10.3, pIK11.1, pIK11.2 to sequencing with primers VF2, VR | ||
| Line 72: | Line 72: | ||
** add sterile filter paper, pipet 8 µl, 4 µl, 2 µl, 1 µl of [[Bacitracin stock solution|Bacitracin]] on filter paper discs | ** add sterile filter paper, pipet 8 µl, 4 µl, 2 µl, 1 µl of [[Bacitracin stock solution|Bacitracin]] on filter paper discs | ||
** grow at 37°C | ** grow at 37°C | ||
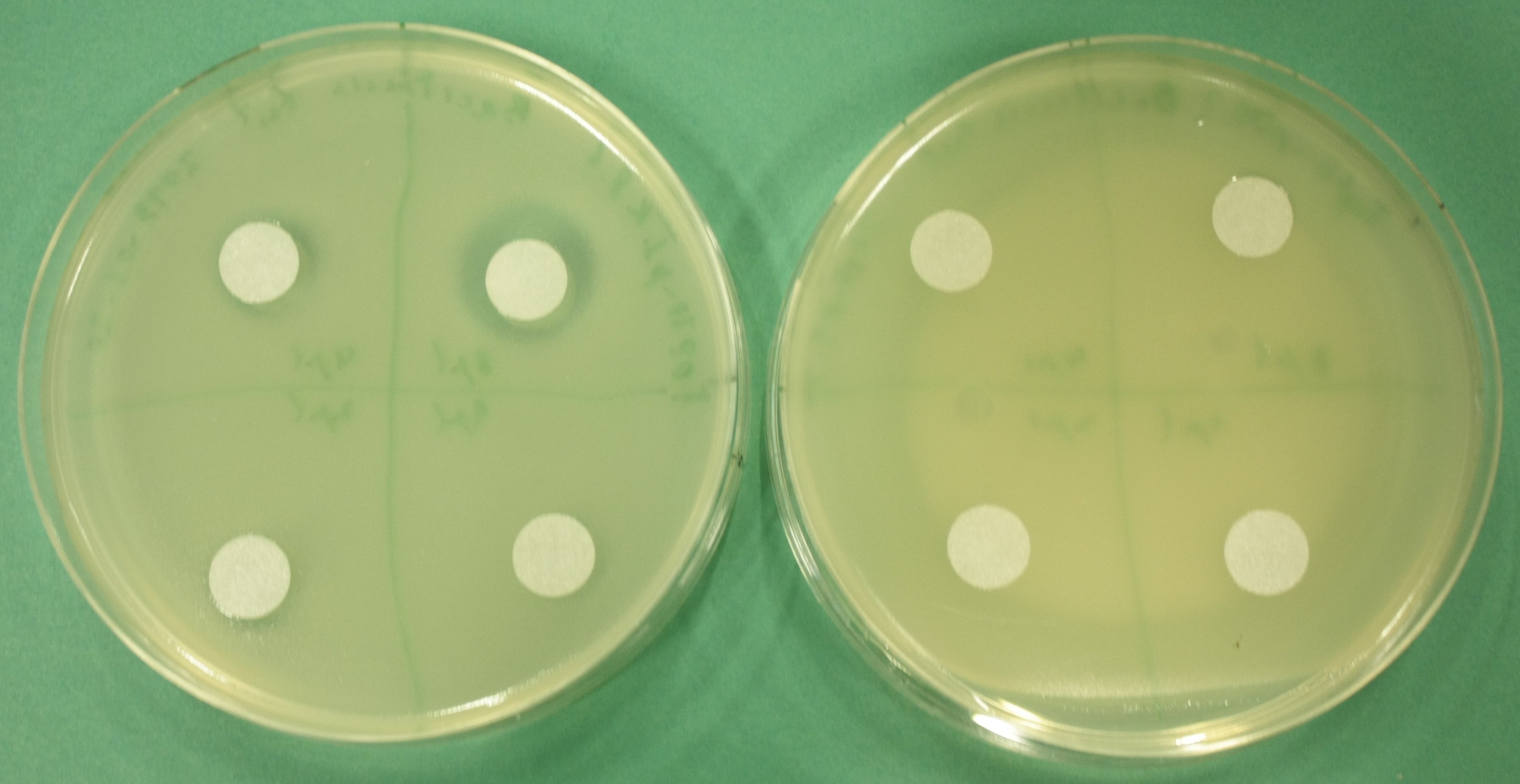
| - | [[File: | + | [[File:Heidelberg_Methylmalonyl Bacitracintest 2013-09-20.jpg|600px|thumb|center|Left: TOP10-pIK8.6, right: TOP10-pIK8.1 (negative control)<br/>counterclockwise, starting top right: 8 µl, 4 µl, 2 µl, 1 µl bacitracin]] |
* Co-transform TOP10 with 25 ng pIK9.1, 10 ng pRB22.4, plate on Cm+Kan, grow at 37°C | * Co-transform TOP10 with 25 ng pIK9.1, 10 ng pRB22.4, plate on Cm+Kan, grow at 37°C | ||
Revision as of 23:44, 4 October 2013
Contents |
2013-09-16
- send pIK8.6, pIK8.9 to sequencing with primer RB43
2013-09-17
- sequences arrived: PIK8.6-pIK8.9-2013-09-16.zip, PIK8.6-2013-09-16 RB43.clustal.txt, File:Heidelberg_PIK8.9-2013-09-16 RB43.clustal.txt
- no mutations at primer site
- sequence with VR primer
- inoculate 2x 6 ml 2xYT+Cm with DH10ß-pIK8.6
2013-09-18
- sequences arrived: PIK8.6-pIK8.9-2013-09-18.zip, PIK8.6-2013-09-18 VR.clustal.txt, File:Heidelberg_PIK8.9-2013-09-18 VR.clustal.txt
- sequences look good
- make miniPrep of pIK8.6 -> 269 ng/µl in 38 µl
- to verify functionality of permeability device: need to co-transform with indigoidine synthetase => need to clone into pSB3K3 -> pIK9
- need to clone into pSB1C3 for submission (pIK10, pIK11): digest 465 ng pSB1C3 (1 µl of Ralf's midiPrep) with EcoRI+SpeI, treat with antarctic phosphatase (37°C, 60 min)
- digest pIK8.6 with EcoRI+SpeI (3 µl DNA, 0.5 µl enzyme, 20 µl total volume)
- gel-purify pIK8.6 insert (20 ng / µl in 20 µl), pSB1C3 (18 µl, concentration not measured)
- ligate for 1 h at RT:
| pIK9 | pIK10 | pIK11 | |
|---|---|---|---|
| backbone | 9 µl pSB3K3 ES from 2013-09-27 | 9 µl pSB1C3 ES | 9 µl pSB1C3 ES |
| insert | 8 µl pIK8.6 ES | 8 µl pIK8.6 ES | 4.5 µl pIK2.6 ES (rest from 2013-08-24) |
| T4 ligase | 1 µl | 1 µl | 1 µl |
| T4 ligase buffer | 2 µl | 2 µl | 2 µl |
| H2O | 0 µl | 0 µl | 3.5 µl |
- transform TOP10 with 10 µl of each ligation, plate on LB+Kan(pIK9), LB+Cm(pIK10, pIK11)
2013-09-19
- colonies present for all ligations
- colonies for pIK10 extremely small -> permeability device toxic
- run colony-PCR using primers VF2+IK25 (expected: 1 kb, OneTaq, 20 µl total volume):
| Cycles | temperature [°C] | Time [s] |
|---|---|---|
| 1 | 95 | 300 |
| 35 | 95 | 30 |
| 54 | 30 | |
| 72 | 90 | |
| 1 | 72 | 600 |
| 1 | 10 | inf |
- grow in TB+Kan (pIK9) / TB+Cm (pIK10, pIK11)
- evening: make miniPreps of pIK9, pIK11 (pIK10 grew very slowly) -> ca. 50-60 ng/µl (pIK9), 100-300 ng / µl (pIK11) in 38 µl
- inoculate 2xYT+Cm with pIK8.1, pIK8.6
2013-09-20
- make miniPreps of pIK10 -> 400-600 ng/µl in 38 µl
- digest everything with BamHI+HindIII, expected: 2.1 kb + 3.1 kb + 4.1 kb (pIK9), 2.1 kb + 6.6 kb (pIK10), 2.1 kb + 4.3 kb (pIK11) (20 µl total volume, 0.5 µl enzyme, using 0.5 µl (pIK10), 1 µl (pIK11), 2 µl (pIK9) DNA)
- to distinguish between pSB3C5 and pSB1C3: digest pIK10, pIK11 with XhoI (expected: 1 band for pSB3C5, 3 bands for pSB1C3)
- send pIK10.1, pIK10.3, pIK10.3, pIK11.1, pIK11.2 to sequencing with primers VF2, VR
- test permeability device:
- inoculate 3 ml LB+Cm soft agar with 100 µl of pIK8.1, pIK8.6 ON culture, pour on LB+Cm plates
- add sterile filter paper, pipet 8 µl, 4 µl, 2 µl, 1 µl of Bacitracin on filter paper discs
- grow at 37°C
- Co-transform TOP10 with 25 ng pIK9.1, 10 ng pRB22.4, plate on Cm+Kan, grow at 37°C
2013-09-21
- co-transformats: lots of extremely small, blue-ish colonies, several big white colonies -> possibly mutated indigoidine synthetase, several big black colonies -> possibly mutated permeability device
- pick one little blue-ish colony, grow in 2xYT+Cm+Kan at 37°C
 "
"