Team:Lethbridge/results
From 2013.igem.org
| Line 109: | Line 109: | ||
<h2>Generating a Pseudoknot Library</h2> | <h2>Generating a Pseudoknot Library</h2> | ||
| - | <p><b>Generating a Library of Pseudoknots with Variable Frameshifting Frequencies</b></p> | + | <p><b>Generating a Library of Pseudoknots with Variable Frameshifting Frequencies</b></p><br> |
| - | <p>To make better use of the frameshifting capability of the pseudoknot, we are also generating a randomized library of pseudoknot sequences that can be characterized for frameshifting frequency. It is our goal to create a library of pseudoknots that have a range of frameshifting frequencies to better facilitate broad application of this part. To do this, the PK401 sequence will be mutagenized using error-prone PCR to generate a library of primers that will be used to amplify the plasmid containing the original sequence. The newly generated plasmids contained mutated PK401 sequences will be transformed into E. coli cells, sequenced, and characterized for their specific frameshifting efficiency using a construct such as BBa_K1210000.</p> | + | <p>To make better use of the frameshifting capability of the pseudoknot, we are also generating a randomized library of pseudoknot sequences that can be characterized for frameshifting frequency. It is our goal to create a library of pseudoknots that have a range of frameshifting frequencies to better facilitate broad application of this part. To do this, the PK401 sequence will be mutagenized using error-prone PCR to generate a library of primers that will be used to amplify the plasmid containing the original sequence. The newly generated plasmids contained mutated PK401 sequences will be transformed into E. coli cells, sequenced, and characterized for their specific frameshifting efficiency using a construct such as BBa_K1210000.</p><br> |
<p><b>Optimizing Conditions for the Error-Prone PCR</b></p> | <p><b>Optimizing Conditions for the Error-Prone PCR</b></p> | ||
| + | <br> | ||
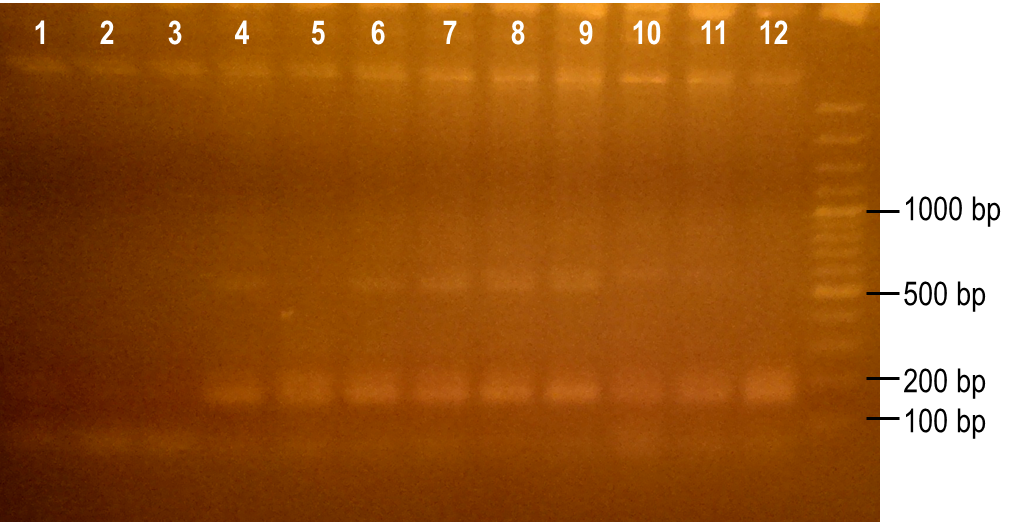
<p>To generate a library with high variability of sequence, conditions for the error-prone PCR had to first be optimized. We wanted to disrupt the PCR experiment enough to produce a large variety of primer sequences, but not too much so that the PCR was no longer successful. To do this, we used a temperature gradient to determine the annealing temperature that would best facilitate the amplification of the expected product. Figure 6 shows the resulting PCR products from the temperature gradient reactions, using temperatures from 41.9-62.1°C. The expected size of the PCR product is 93 bp, which was seen in the first three reaction samples. Various unspecific PCR products were seen in the remaining reactions samples. For this reason, the temperature chosen for the subsequent error-prone PCRs was 43.6°C, the temperature used to produce the PCR product in lane 3.</p> | <p>To generate a library with high variability of sequence, conditions for the error-prone PCR had to first be optimized. We wanted to disrupt the PCR experiment enough to produce a large variety of primer sequences, but not too much so that the PCR was no longer successful. To do this, we used a temperature gradient to determine the annealing temperature that would best facilitate the amplification of the expected product. Figure 6 shows the resulting PCR products from the temperature gradient reactions, using temperatures from 41.9-62.1°C. The expected size of the PCR product is 93 bp, which was seen in the first three reaction samples. Various unspecific PCR products were seen in the remaining reactions samples. For this reason, the temperature chosen for the subsequent error-prone PCRs was 43.6°C, the temperature used to produce the PCR product in lane 3.</p> | ||
<br> | <br> | ||
| - | <center>[[Image:ULeth2013iGEM tempgradient.png| | + | <center>[[Image:ULeth2013iGEM tempgradient.png|500px]]</center> |
| - | <p><b>Figure 6. Determining the ideal temperature for error-prone PCR reactions.</b> PCR reactions were performed using various annealing temperatures from 41.9-62.1°C (lanes 1-12). The primers used in the reaction flank the PK401 pseudoknot, making the expected product size 93 bp. The temperature from the reaction in lane 3 (43.6°C) was selected for subsequent error-prone PCRs.</p> | + | <p><b>Figure 6. Determining the ideal temperature for error-prone PCR reactions.</b> PCR reactions were performed using various annealing temperatures from 41.9-62.1°C (lanes 1-12). The primers used in the reaction flank the PK401 pseudoknot, making the expected product size 93 bp. The temperature from the reaction in lane 3 (43.6°C) was selected for subsequent error-prone PCRs.</p><br> |
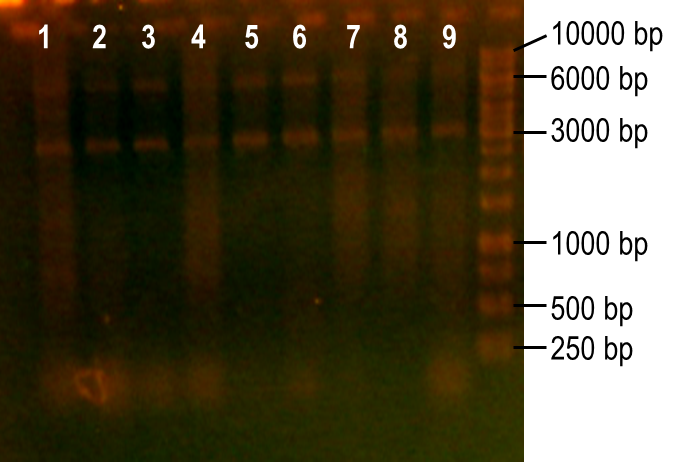
<p>Next, we modulated a few of the typical PCR conditions to induce errors during the reaction. First, we used Taq polymerase, which is known to have higher error rates than higher fidelity polymerases commonly used in PCRs. Second, we added MnCl2 to the reaction to further hinder the accuracy of the polymerase. Finally, different ratios of deoxynucleotides were used in the reactions, either by increasing the dGTP concentration or by using a higher concentration of both dTTP and dCTP. A number of these conditions worked to produce the expected PCR product at 93 bp (Fig. 7, lanes 1-4, 6). The conditions tested in lanes 1-4 included MnCl2 at concentrations of 0-0.45 mM, and lane 6 tested the effect of increasing dGTP concentrations to 1 mM while the other dNTPs were at a concentration of 0.2 mM.</p> | <p>Next, we modulated a few of the typical PCR conditions to induce errors during the reaction. First, we used Taq polymerase, which is known to have higher error rates than higher fidelity polymerases commonly used in PCRs. Second, we added MnCl2 to the reaction to further hinder the accuracy of the polymerase. Finally, different ratios of deoxynucleotides were used in the reactions, either by increasing the dGTP concentration or by using a higher concentration of both dTTP and dCTP. A number of these conditions worked to produce the expected PCR product at 93 bp (Fig. 7, lanes 1-4, 6). The conditions tested in lanes 1-4 included MnCl2 at concentrations of 0-0.45 mM, and lane 6 tested the effect of increasing dGTP concentrations to 1 mM while the other dNTPs were at a concentration of 0.2 mM.</p> | ||
<br> | <br> | ||
| - | <center>[[Image:ULeth2013iGEM_errorpronepcr.png| | + | <center>[[Image:ULeth2013iGEM_errorpronepcr.png|500px]]</center> |
| - | <p><b>Figure 7. Error-prone PCR of PK401 sequence.</b> PCR conditions were adjusted to induce errors in the PK401 sequence. Reactions were performed with the following modified condtions: lanes 1-6 used MnCl2 concentrations from 0-0.65 mM, lanes 4-6 used dGTP concentrations that were increased to 0.7-1.2 mM, lane 7 used increased concentrations of dTTP and dCTP, lane 8 used increased concentrations of dGTP only, and lane 9 used 0.65 mM MnCl2 only.</p> | + | <p><b>Figure 7. Error-prone PCR of PK401 sequence.</b> PCR conditions were adjusted to induce errors in the PK401 sequence. Reactions were performed with the following modified condtions: lanes 1-6 used MnCl2 concentrations from 0-0.65 mM, lanes 4-6 used dGTP concentrations that were increased to 0.7-1.2 mM, lane 7 used increased concentrations of dTTP and dCTP, lane 8 used increased concentrations of dGTP only, and lane 9 used 0.65 mM MnCl2 only.</p><br> |
<p>Now that we have determined the conditions for the error-prone PCR, we will perform a large-scale amplification so that we can purify the resulting PK401 primers by extracting it from an agarose gel. The primers will then be used in a high fidelity PCR reaction to amplify the entire plasmid, which will create the library of PK401 sequences. The final steps will be to transform the final PCR products into E. coli cells, screen them for frameshifting frequency using a similar construct as BBa_K1210000, and sequence the pseudoknot before submitting the parts to the registry to complete the library of frameshifting elements.</p> | <p>Now that we have determined the conditions for the error-prone PCR, we will perform a large-scale amplification so that we can purify the resulting PK401 primers by extracting it from an agarose gel. The primers will then be used in a high fidelity PCR reaction to amplify the entire plasmid, which will create the library of PK401 sequences. The final steps will be to transform the final PCR products into E. coli cells, screen them for frameshifting frequency using a similar construct as BBa_K1210000, and sequence the pseudoknot before submitting the parts to the registry to complete the library of frameshifting elements.</p> | ||
Revision as of 02:41, 28 September 2013

 "
"