Team:NCTU Formosa/project
From 2013.igem.org
Introduction
Base pairing offers a powerful way for one RNA to control the activity of another. Both prokaryotes and eukaryotes have many cases which a single-stranded RNA base pairs with a complementary region of an mRNA. As a result, it prevents expression of the mRNA.
RNA interference(RNAi), also called post transcriptional gene silencing, is a process in which RNA molecules inhibit gene expression by destroying specific mRNA molecules. In 2006, Andrew Fire and Craig C.Mello shared the Nobel Prize in Physiology or Medicine because of their study on RNA interference11. This powerful gene silencing tool in eukaryotes has been used in many research. As their counterparts in bacteria, small non-coding RNAs (sRNAs) are important regulatory roles.
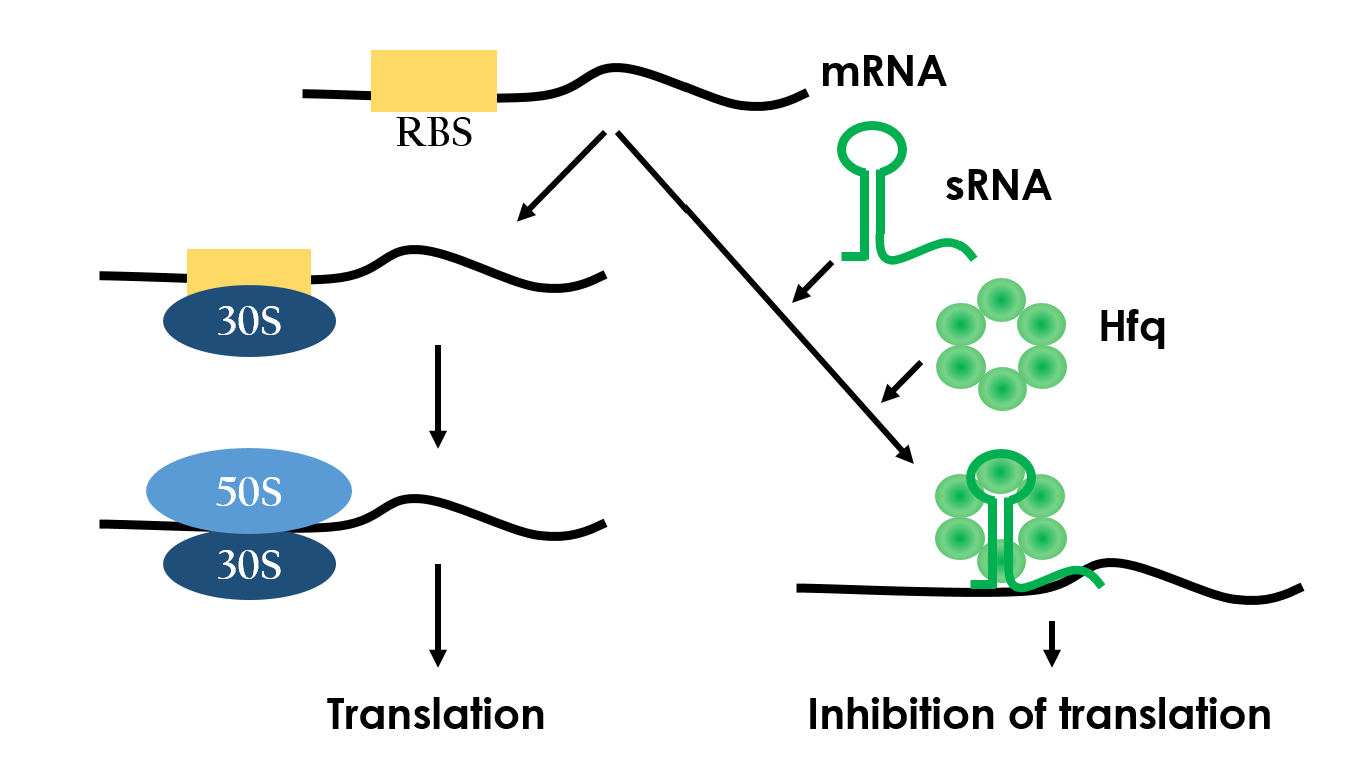
Small RNAs (sRNAs) have become increasingly significant in playing the role of bacterial gene regulation. Most sRNAs interact with the targeted mRNAs by imperfect base pairing, reducing the translation efficiency and recruiting chaperones such as Hfq for translation termination. The sRNAs would bind to the target with its hairpin-like structure which wound by some of it's own sequences, and the chaperons would stuck between the hairpins in order to protect the sRNA-mRNA combination from degrading. In the end, the sRNAs regulate gene expression by forestalling translation. Since this regulated-system has already been employed in vivo, it is necessary to design an artificial sRNA that targets specifically to a desired genes, so that we can prevent the sRNA from affecting other undesired genes.
Unlike the regulated-system popular used in iGEM projects (e.g., Ptet & tetR, Plux & LuxR, etc.), sRNA regulated-system seems more efficient. Because the inhibition works under RNA level, which means the energy wasted in E. coli is less than other system(producing proteins to regulate).
Figure 8 depicts how the small RNA chaperone, Hfq, associates with the small RNA and represses a target mRNA.12
 "
"