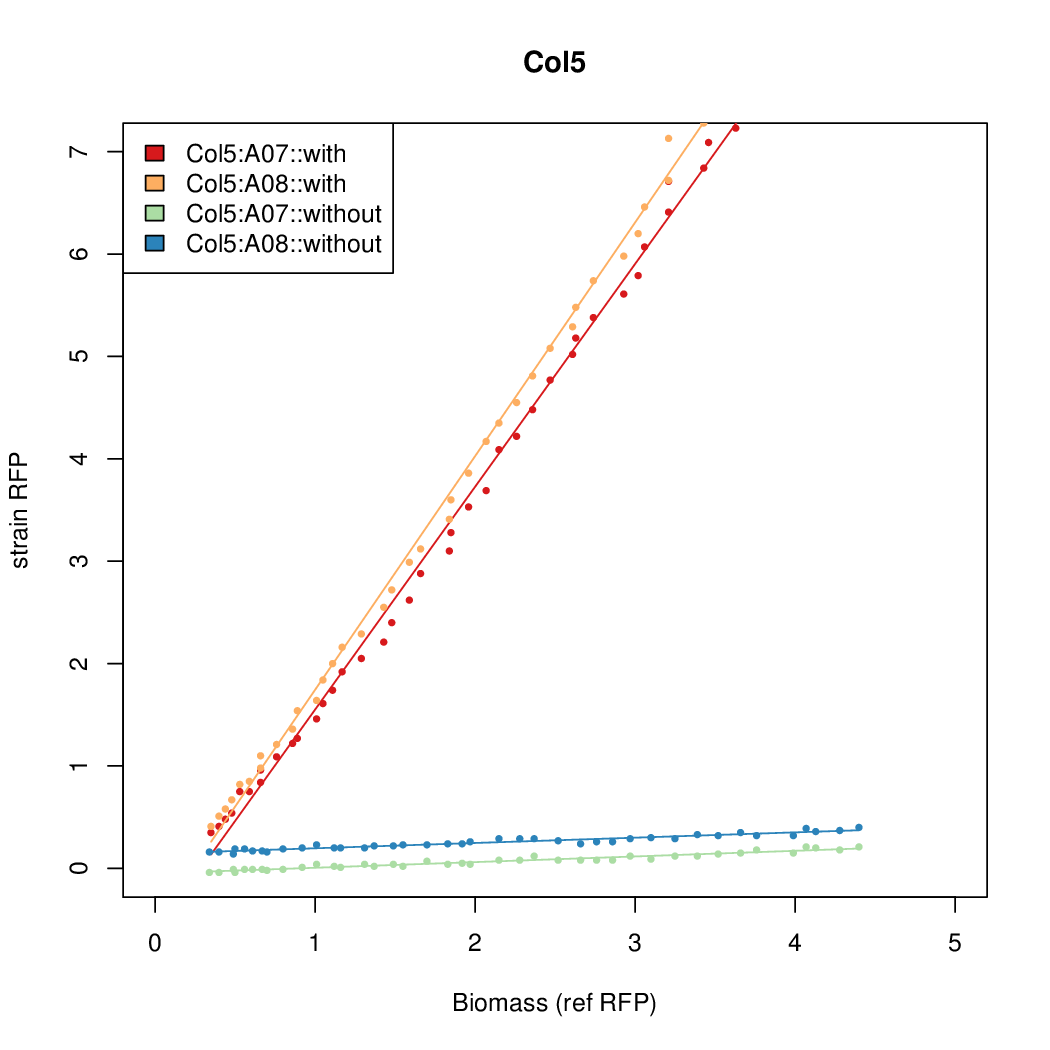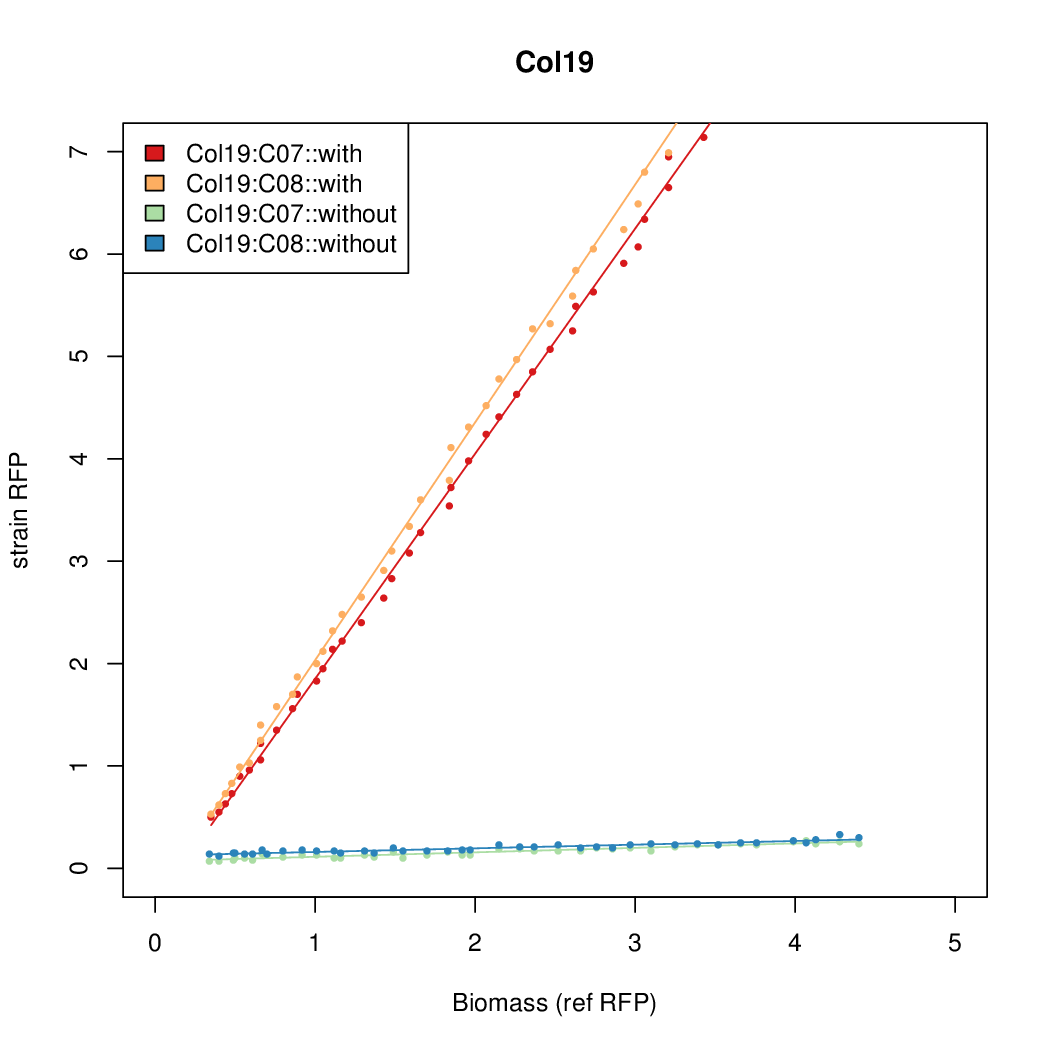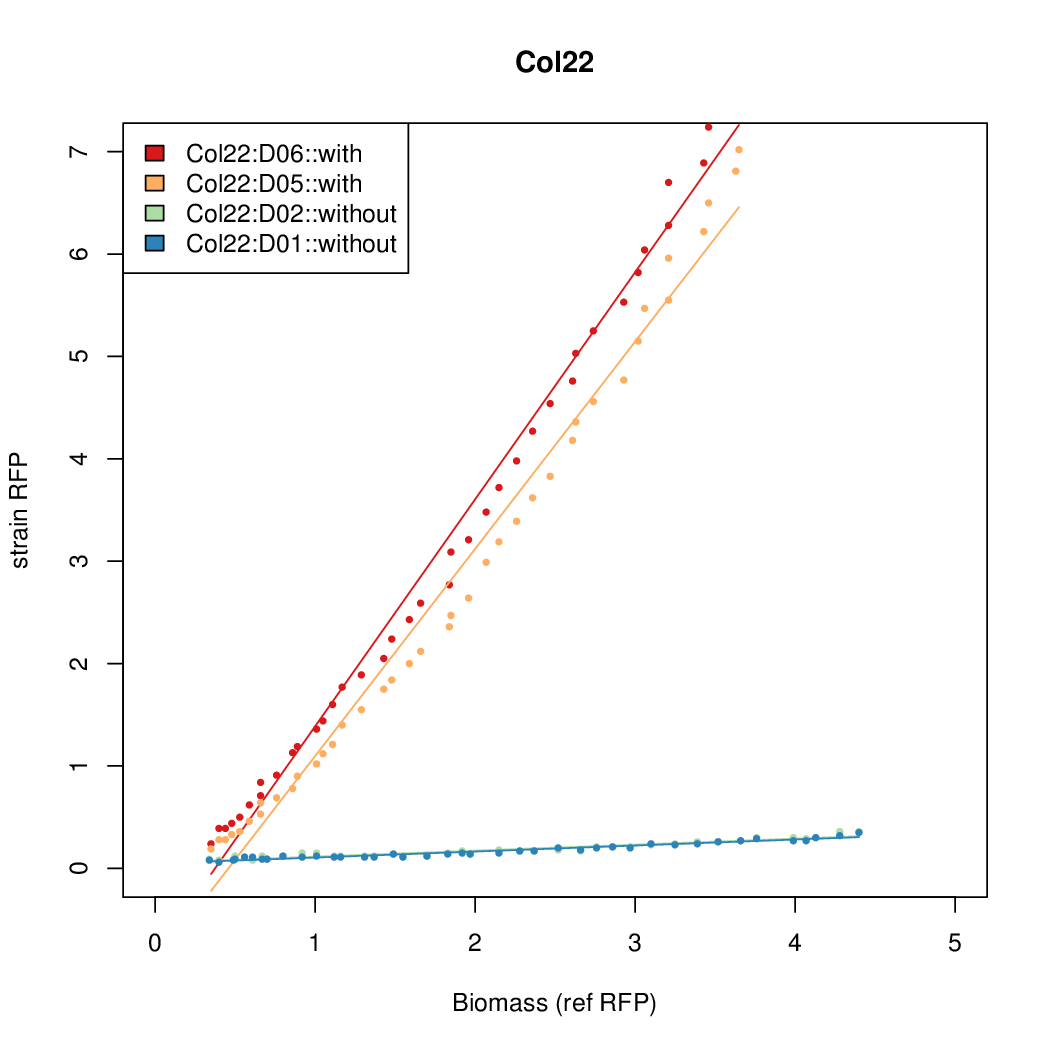Team:DTU-Denmark/pBAD SPL
From 2013.igem.org
(→Details) |
(→Details) |
||
| Line 60: | Line 60: | ||
|- | |- | ||
|align="center"|Col. 3 | |align="center"|Col. 3 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGTAATCATACCGCCGAAAGTATTATCATTA</font> |
|align="center"|16.6 | |align="center"|16.6 | ||
|align="center"|0.426 | |align="center"|0.426 | ||
| Line 66: | Line 66: | ||
|- | |- | ||
|align="center"|Col. 4 | |align="center"|Col. 4 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCGACAACATTGCGTCCTATAAAATGCCGA</font> |
|align="center"|14.2 | |align="center"|14.2 | ||
|align="center"|0.345 | |align="center"|0.345 | ||
| Line 72: | Line 72: | ||
|- | |- | ||
|align="center"|Col. 5 | |align="center"|Col. 5 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGGCCCGCCCCCGATGCGCATAAAATACCAA</font> |
|align="center"|17.7 | |align="center"|17.7 | ||
|align="center"|0.424 | |align="center"|0.424 | ||
| Line 78: | Line 78: | ||
|- | |- | ||
|align="center"|Col. 12 | |align="center"|Col. 12 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCGTATCGCGAGCGGGCGTTATTATACGCA</font> |
|align="center"|13.6 | |align="center"|13.6 | ||
|align="center"|0.414 | |align="center"|0.414 | ||
| Line 84: | Line 84: | ||
|- | |- | ||
|align="center"|Col. 10 | |align="center"|Col. 10 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCCCGAATCAGTAGTATTTATTATCTAGA</font> |
|align="center"|18 | |align="center"|18 | ||
|align="center"|0.462 | |align="center"|0.462 | ||
| Line 90: | Line 90: | ||
|- | |- | ||
|align="center"|Col. 9 | |align="center"|Col. 9 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCACCCCCCCCCCCGGCGTATAATTCCCA</font> |
|align="center"|16.1 | |align="center"|16.1 | ||
|align="center"|0.530 | |align="center"|0.530 | ||
| Line 96: | Line 96: | ||
|- | |- | ||
|align="center"|Col. 8 | |align="center"|Col. 8 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCCTACCGCTCGCCCCCCTATTATACCCA</font> |
|align="center"|13.4 | |align="center"|13.4 | ||
|align="center"|0.211 | |align="center"|0.211 | ||
| Line 102: | Line 102: | ||
|- | |- | ||
|align="center"|Col. 13 | |align="center"|Col. 13 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCCCCCCCGTCCACCCCCTAATATCCCGA</font> |
|align="center"|10.9 | |align="center"|10.9 | ||
|align="center"|0.096 | |align="center"|0.096 | ||
| Line 108: | Line 108: | ||
|- | |- | ||
|align="center"|Col. 15 | |align="center"|Col. 15 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGTGCGCCTGCCGTCCAAAGTAATATCCTTA</font> |
|align="center"|15.5 | |align="center"|15.5 | ||
|align="center"|0.235 | |align="center"|0.235 | ||
| Line 114: | Line 114: | ||
|- | |- | ||
|align="center"|Col. 18 | |align="center"|Col. 18 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCCCGAATCAGTAGTATTTATTATCTAGA</font> |
|align="center"|19.8 | |align="center"|19.8 | ||
|align="center"|0.524 | |align="center"|0.524 | ||
| Line 120: | Line 120: | ||
|- | |- | ||
|align="center"|Col. 19 | |align="center"|Col. 19 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCTCGCACCGGCGACCCGATAATATCCATA</font> |
|align="center"|18.0 | |align="center"|18.0 | ||
|align="center"|0.315 | |align="center"|0.315 | ||
| Line 126: | Line 126: | ||
|- | |- | ||
|align="center"|Col. 31 | |align="center"|Col. 31 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCCCCCCTCCCCCGACCTAAAATATCCAG</font> |
|align="center"|3.8 | |align="center"|3.8 | ||
|align="center"|0.161 | |align="center"|0.161 | ||
| Line 132: | Line 132: | ||
|- | |- | ||
|align="center"|Col. 29 | |align="center"|Col. 29 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCGCTGCACCCGTCCCCCTATTATAGCAA</font> |
|align="center"|14.8 | |align="center"|14.8 | ||
|align="center"|0.301 | |align="center"|0.301 | ||
| Line 138: | Line 138: | ||
|- | |- | ||
|align="center"|Col. 26 | |align="center"|Col. 26 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCGATCCTATCTCCCTATTAAAATTTTTA</font> |
|align="center"|13.8 | |align="center"|13.8 | ||
|align="center"|0.292 | |align="center"|0.292 | ||
| Line 144: | Line 144: | ||
|- | |- | ||
|align="center"|Col. 22 | |align="center"|Col. 22 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCCCGAATCAGTAGTATTTATTATCTAGA</font> |
|align="center"|16.8 | |align="center"|16.8 | ||
|align="center"|0.466 | |align="center"|0.466 | ||
| Line 150: | Line 150: | ||
|- | |- | ||
|align="center"|Col. 33 | |align="center"|Col. 33 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCAGCTTCCTACTTCAAATATAATCCGTA</font> |
|align="center"|14.4 | |align="center"|14.4 | ||
|align="center"|1.01 | |align="center"|1.01 | ||
| Line 156: | Line 156: | ||
|- | |- | ||
|align="center"|Col. 34 | |align="center"|Col. 34 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGTCCCTCTTCCGCGCCCCCTAAAATACCCA</font> |
|align="center"|12.4 | |align="center"|12.4 | ||
|align="center"|0.187 | |align="center"|0.187 | ||
| Line 162: | Line 162: | ||
|- | |- | ||
|align="center"|Col. 35 | |align="center"|Col. 35 | ||
| - | |<font face="Courier" size="+ | + | |<font face="Courier" size="+0.9">CTGACGCCCAACCCGACACAGCGATATAATAATGA</font> |
|align="center"|18.4 | |align="center"|18.4 | ||
|align="center"|0.89 | |align="center"|0.89 | ||
Revision as of 17:26, 4 October 2013
pBAD SPL
Contents |
pBAD synthetic promoter library
A synthetic promoter library (SPL) is a library of cells each having one promoter sequence that differs between them. This promoter is always the same and is usually upstream a fluorescent protein like GFP/RFP. The term was first coined in 1998 and used in Lactococcus lactis [1]. The method was adapted by 2010 DTU iGEM team to make a SPL for E.coli that enabled the modulation of constitutive gene expression with great precision. They even made a new standard for this method with the use of biobricks ([http://dspace.mit.edu/handle/1721.1/60080 RFC 63]).
We used the method to build a non-leaky arabinose inducible promoter as a tool for expressing lethal proteins in E. coli. The reason was that many of the proteins we are working with are membrane bound or integral membrane proteins and will be lethal if expressed in to high quantities.
For testing our pathways we needed to grow the transformed cell to a certain concentration and growth with the gene constitutively expressed was not working very well. We needed a inducible system but the standard arabinose inducible system was way too leaky and we got same stood with the same problem as before. Therefore we needed to either build or buy (if an possible) an inducible system with great tightness able to be induced easily. We choose to build such system.
Methods
Experimental procedure
- Random promoter sequences were ordered matching the sequence CTGACGNNNNNNNNNNNNNNNNNNTAWWATNNNNA.
- USER cloning to add RFP downstream of promoter.
- Colonies were plated.
- After inspection under UV-light the visually non RFP containing colonies where picked.
- Plates were induced by spraying them with an 5% w/v aqueous arabinose solution.
- The plates were again inspected under UV-light and this time the most red florescent cells were picked.
- Colonies were grown in culture tubes and screened in parallel on BioLector. Wells on BioLector plate were loaded with culture by transferring a toothpick from each overnight culture selected and into the wells of the plate. All wells were run in duplicate.
- All duplicate colonies were run twice -- once with arabinose added at t=0, and again without arabinose.
- The pBAD system [http://parts.igem.org/Part:BBa_K808000 BBa_K808000] was used as a reference.
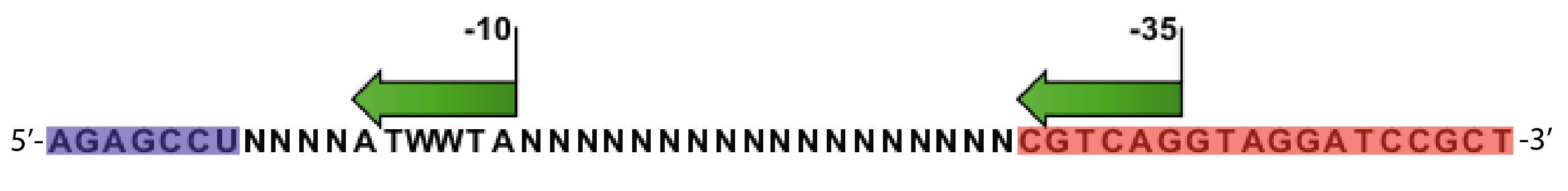
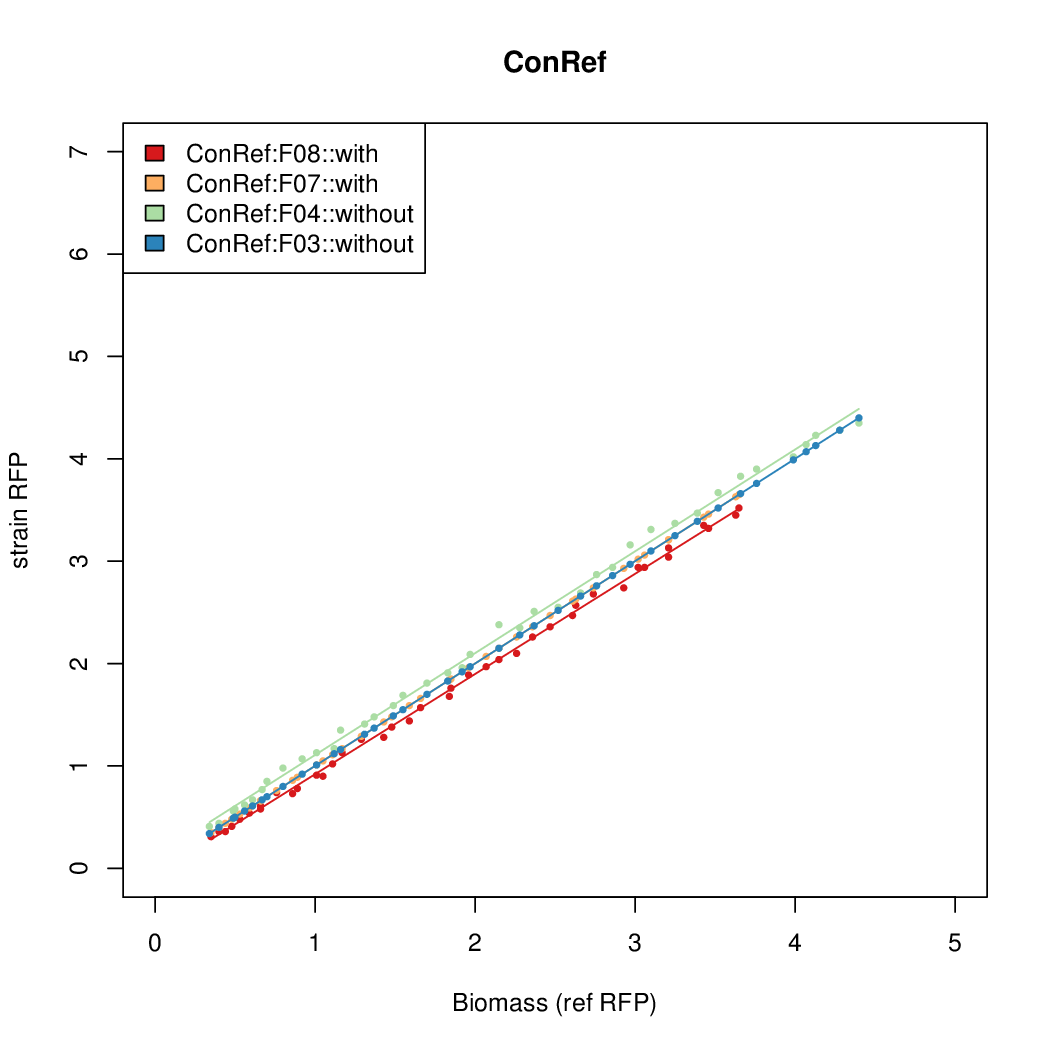
Data analysis
- Data was collected from the Biolector, and analyzed using a series of R scripts written by Chris Workman (unpublished).
- The maturation and degradation times for mCherry were both assumed to be 40 min.
- The growth rate, mu, was estimated to be 1.28 (from an average of all wells on all plates) since we expect each strain to grow at the same rate.
- A time window representing exponential growth was selected (between 1 and 4.5 hours).
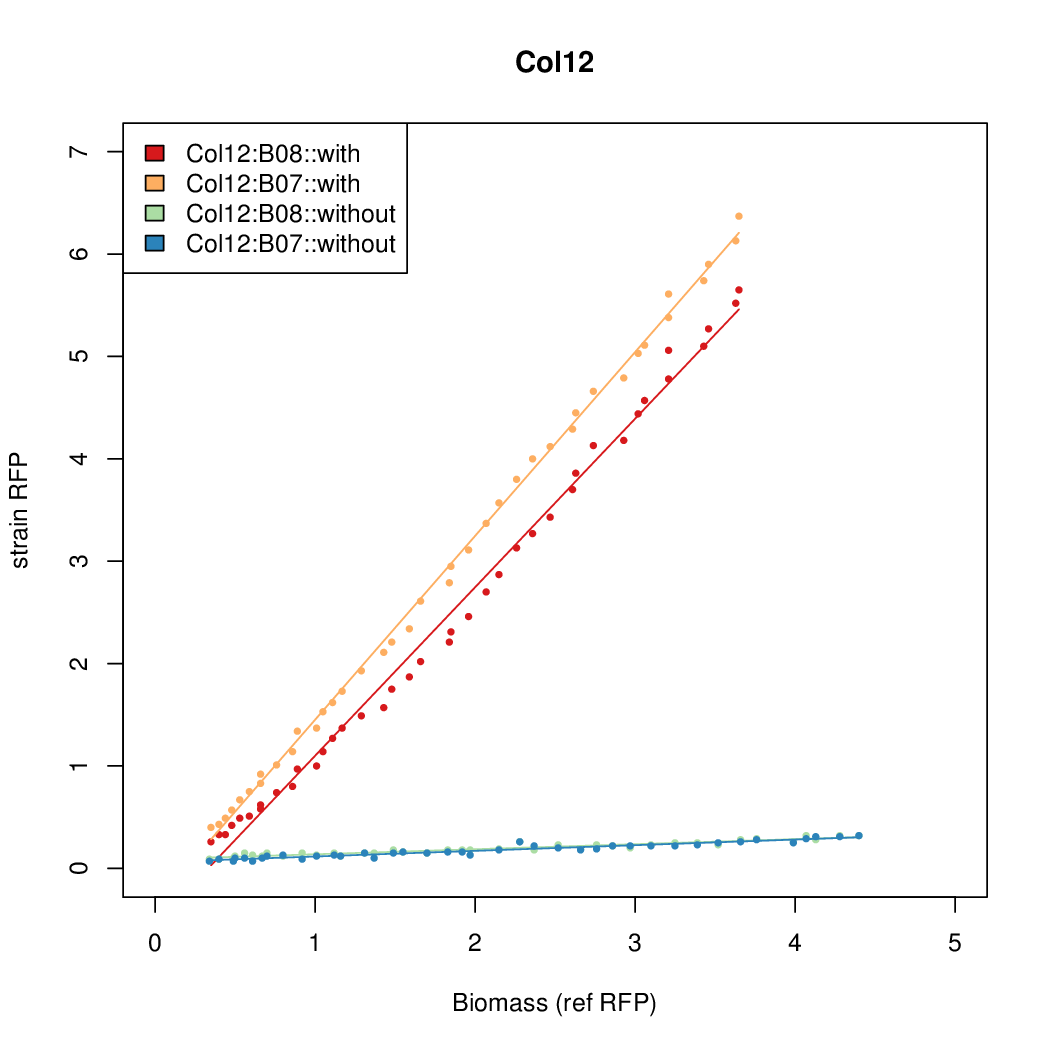
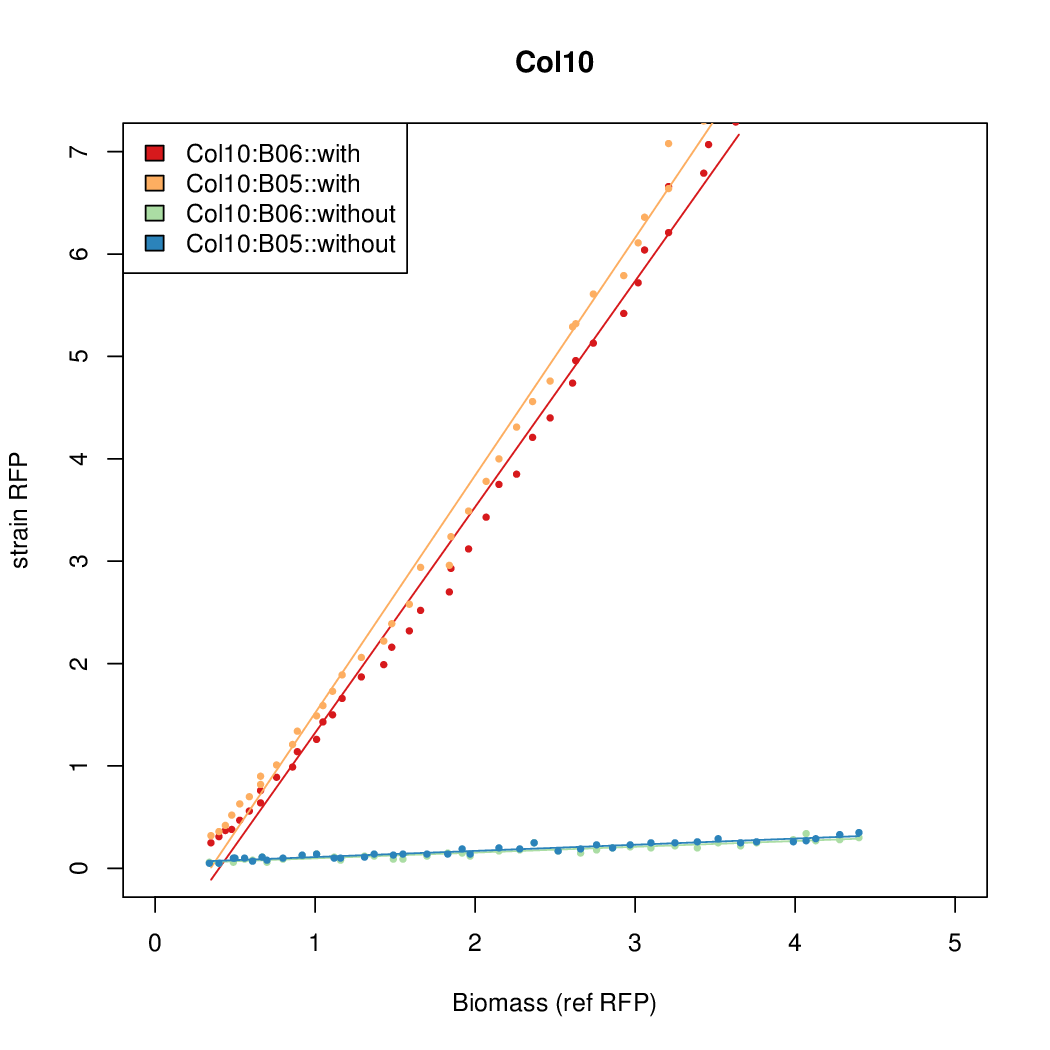
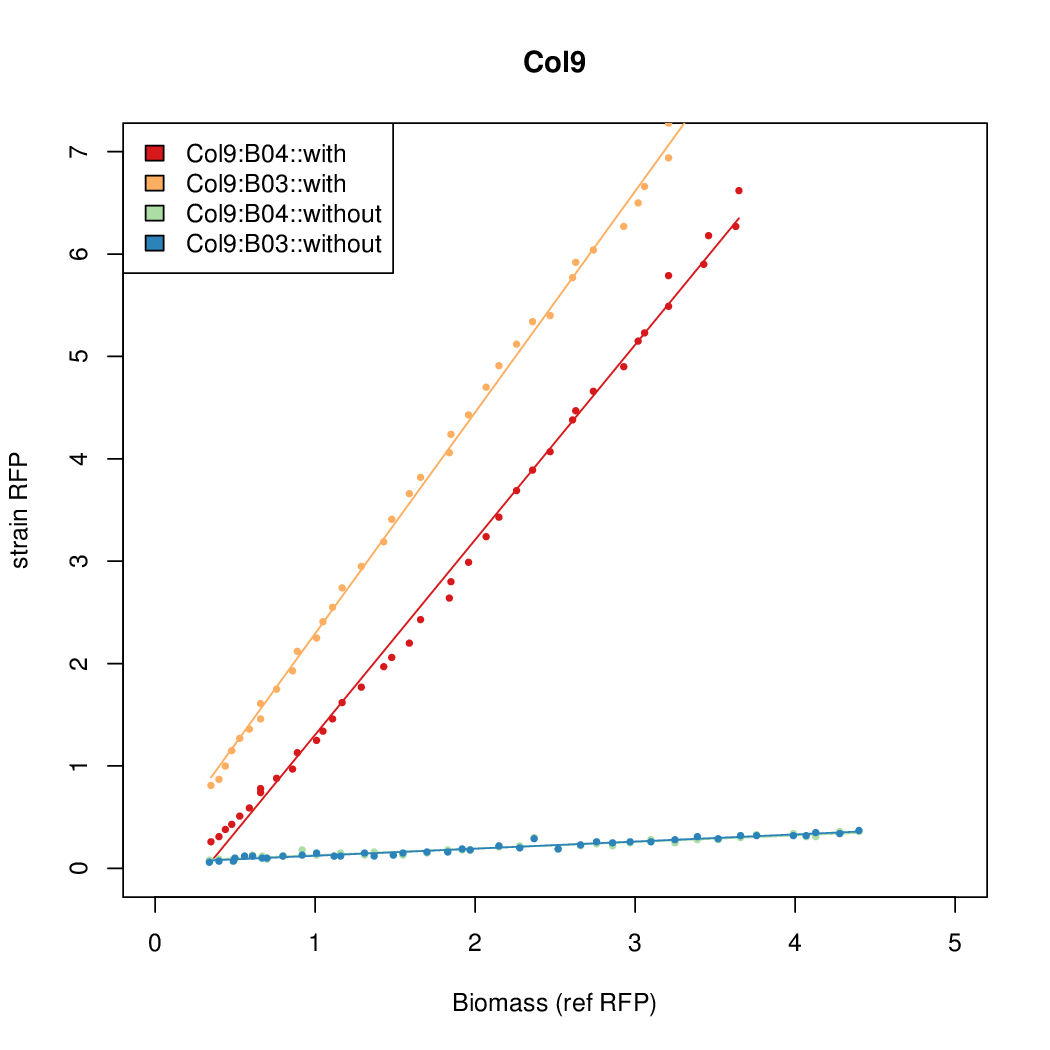
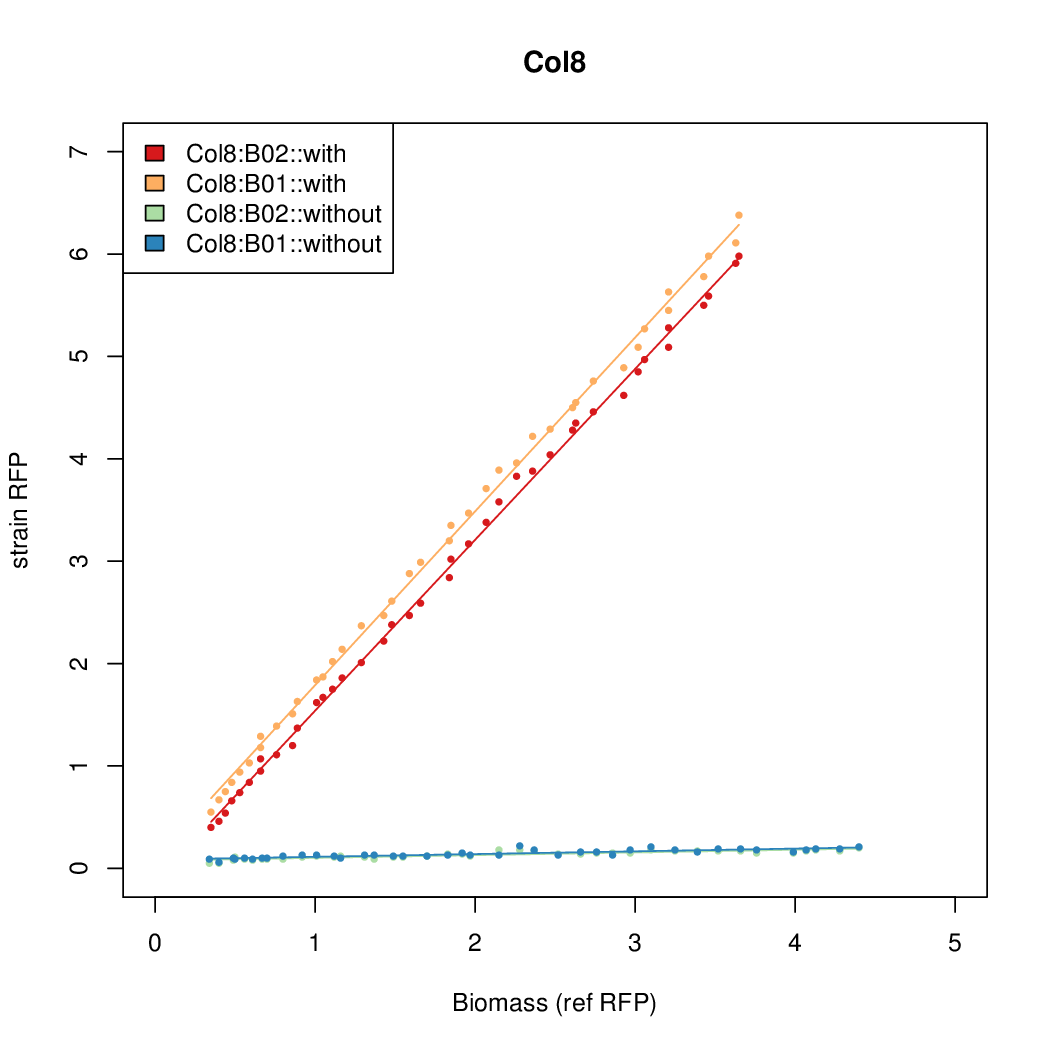
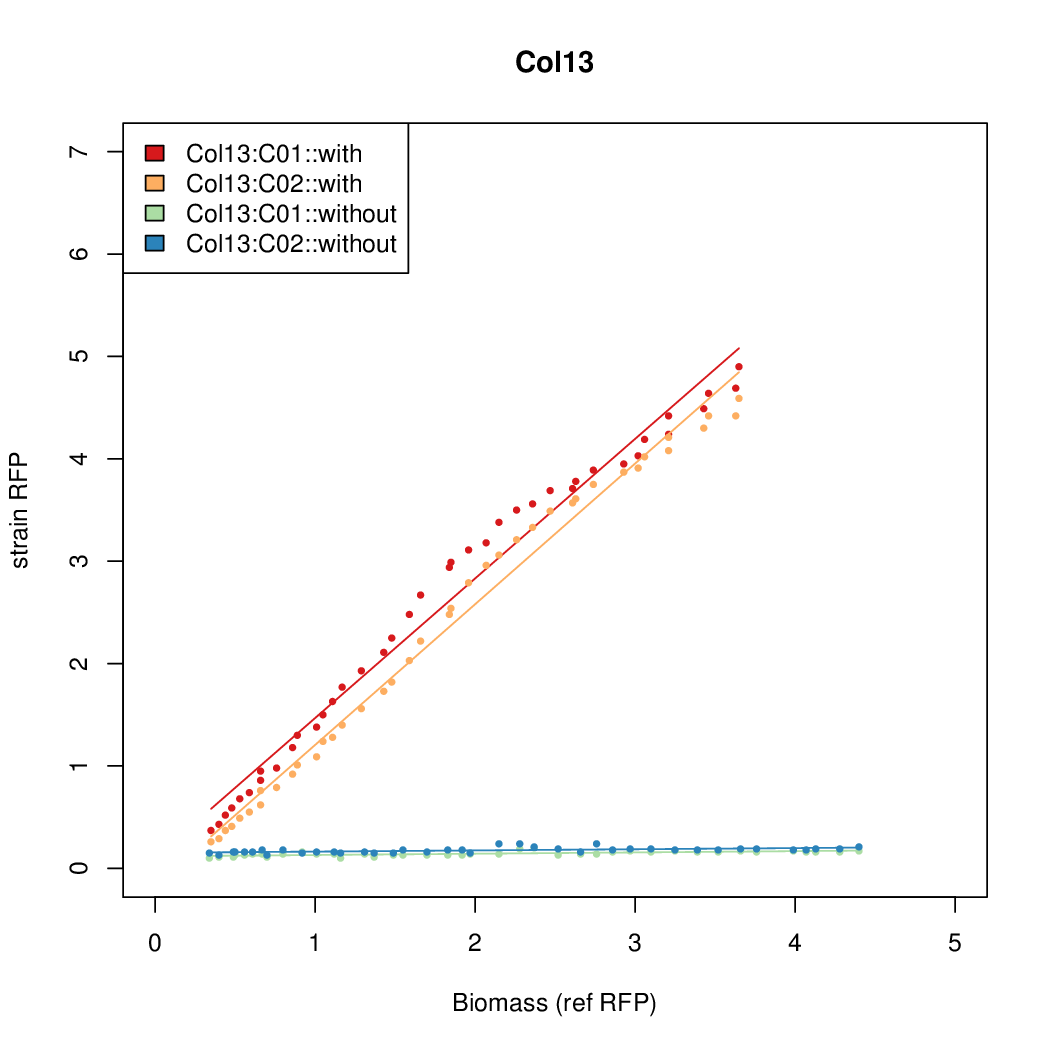
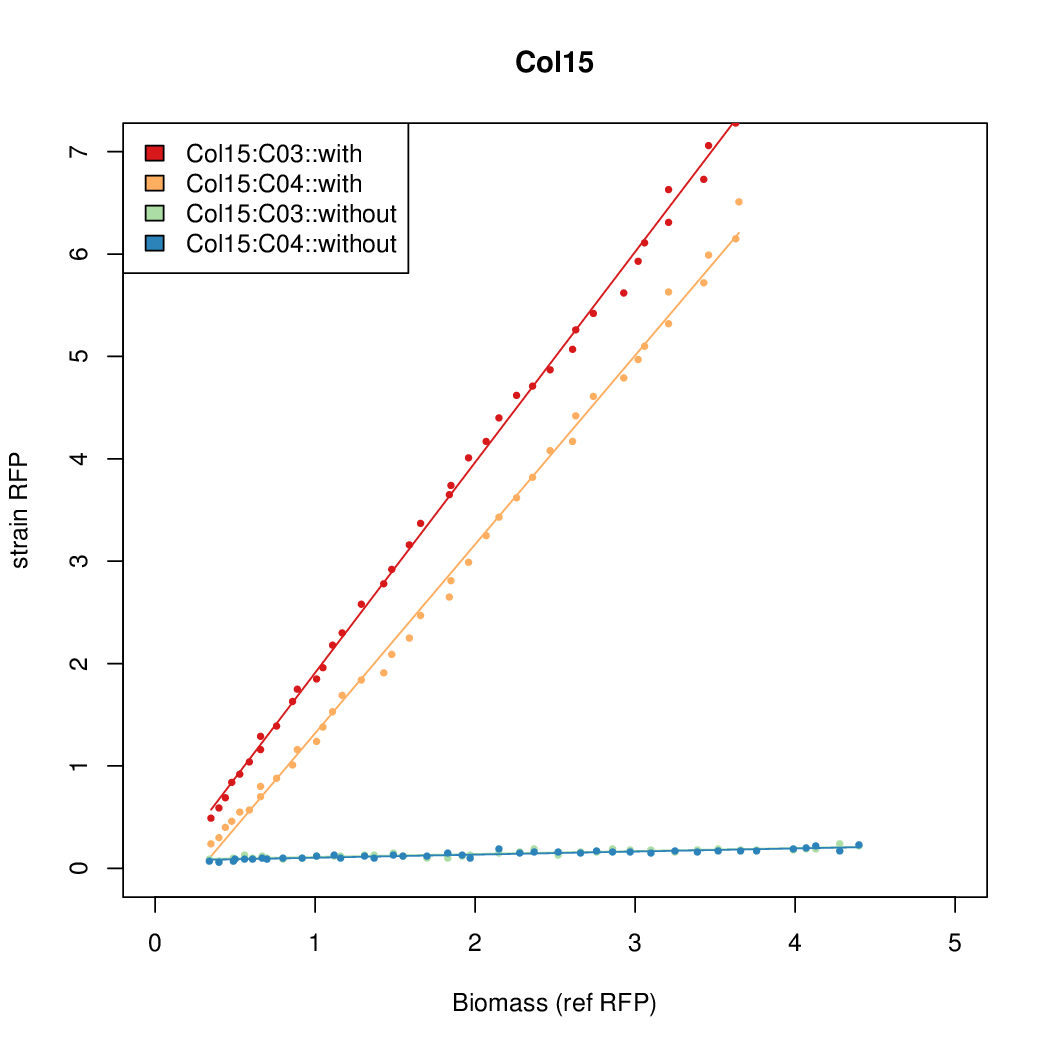
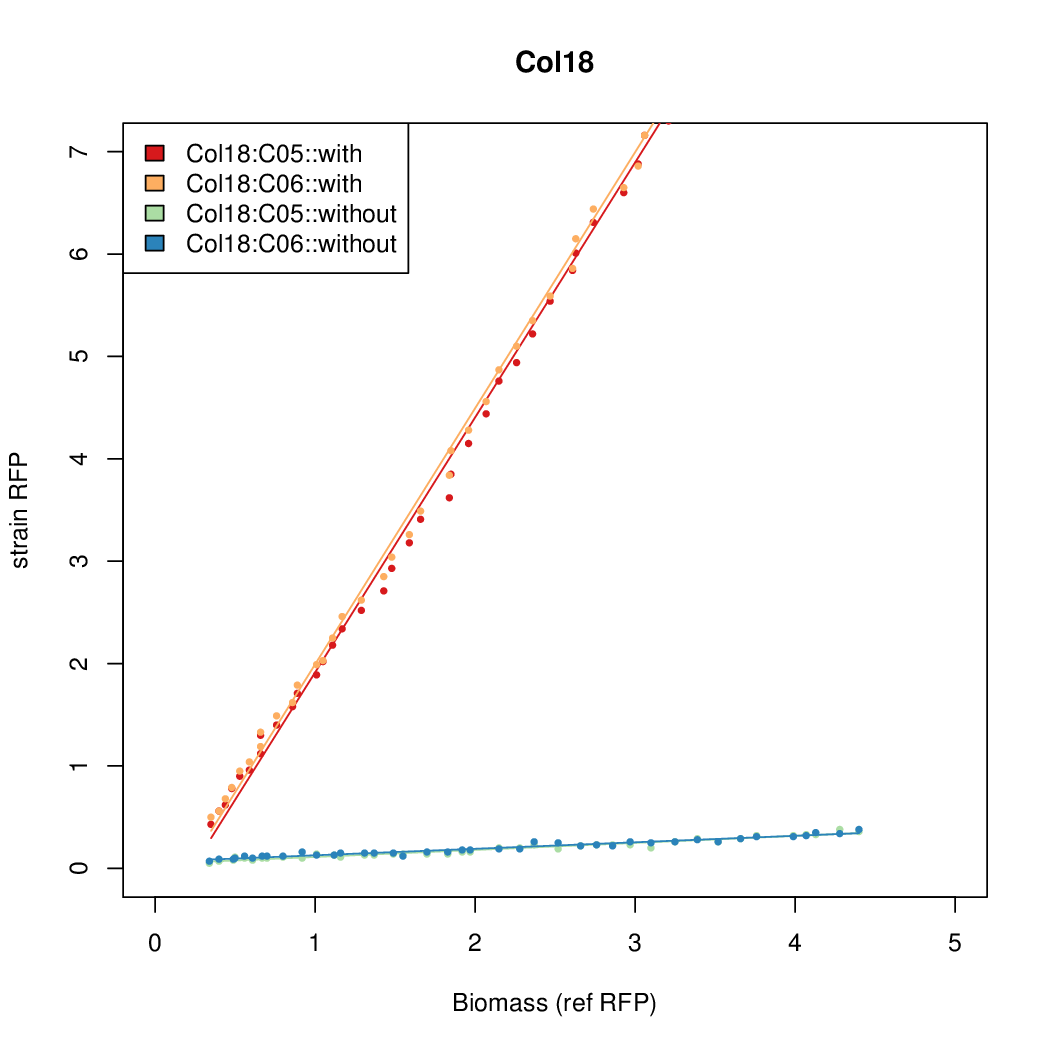
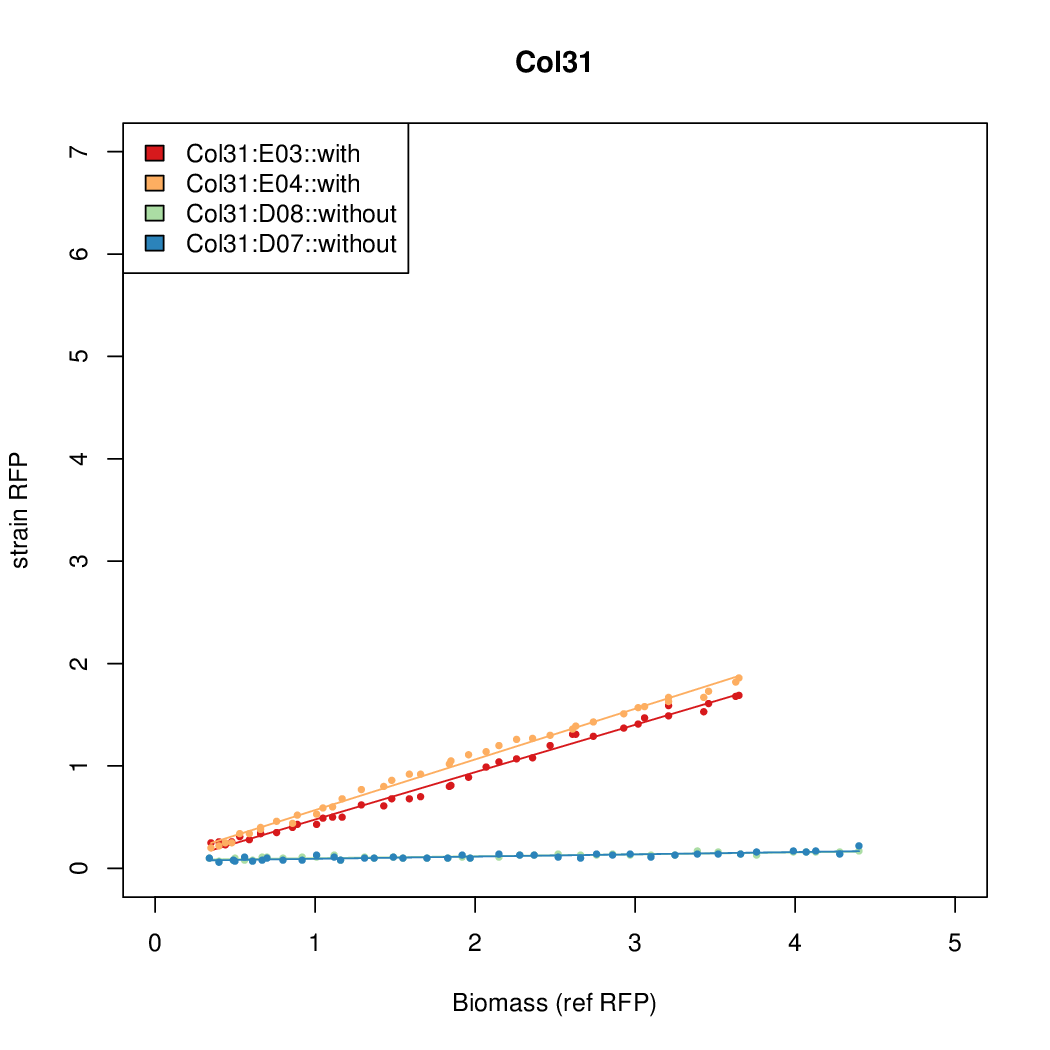
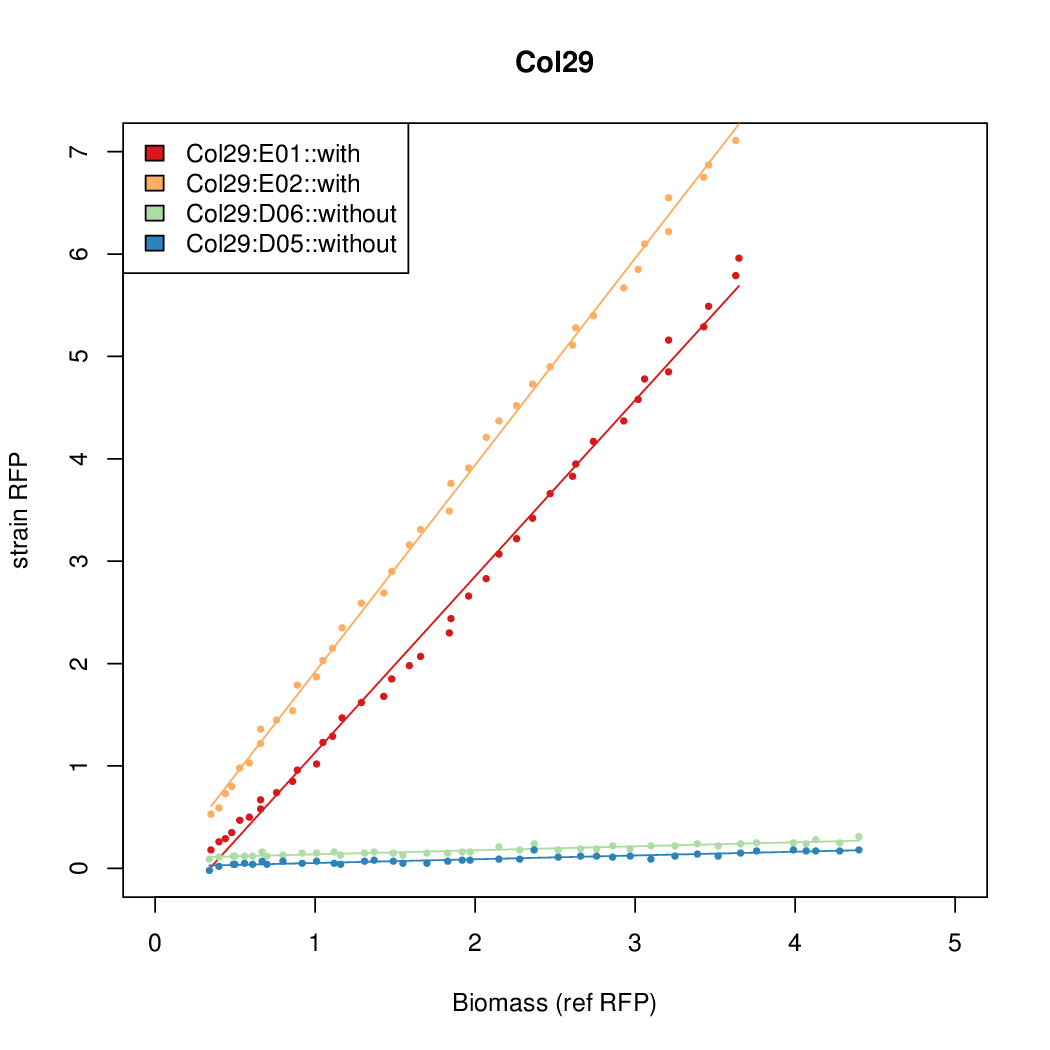
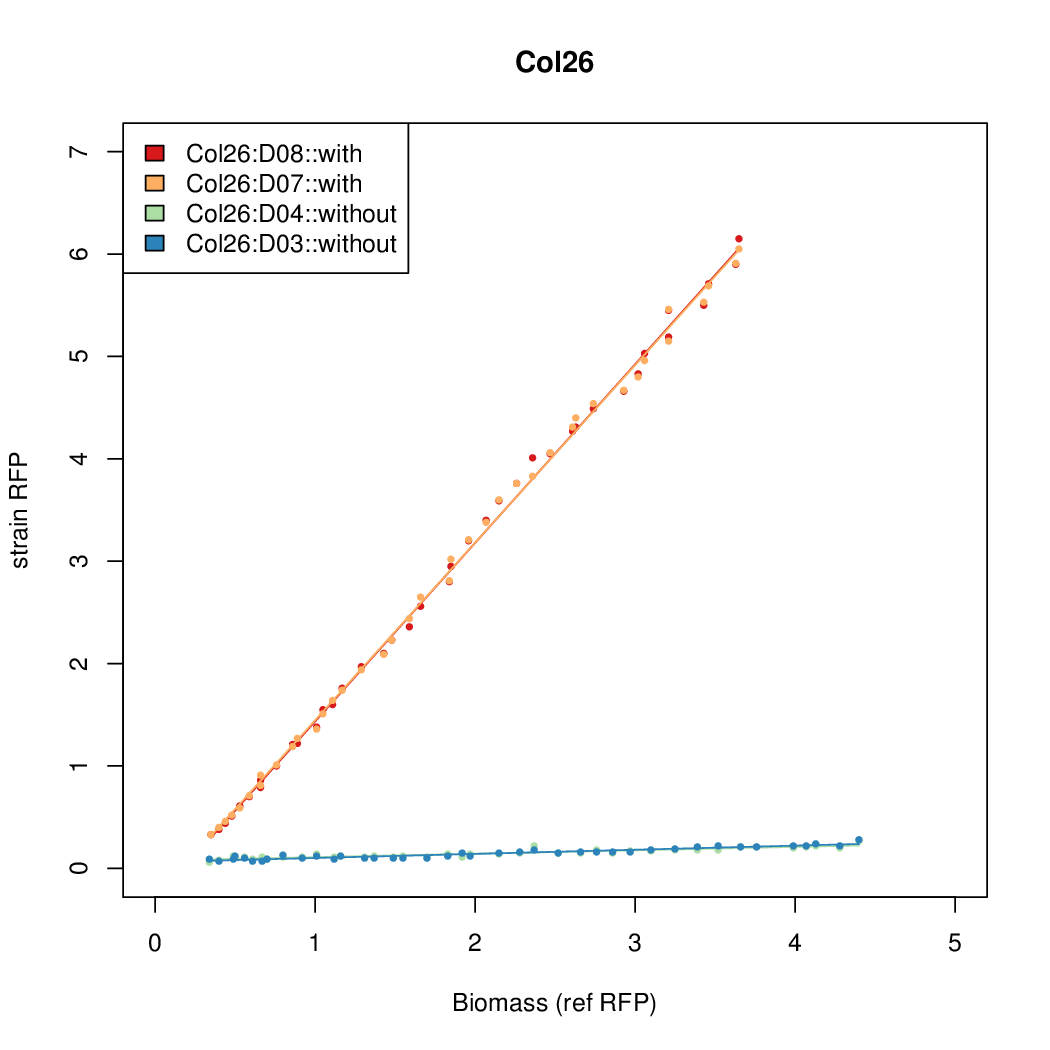
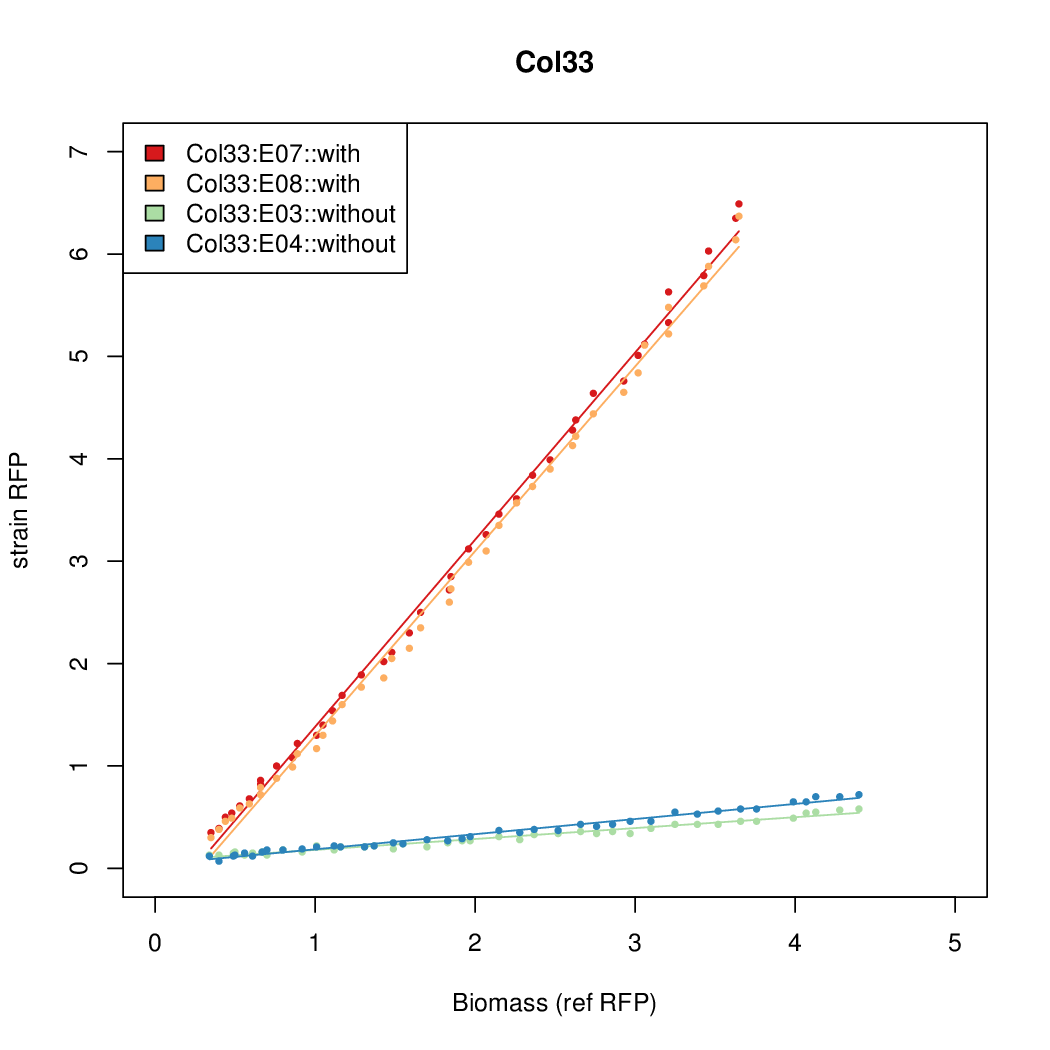
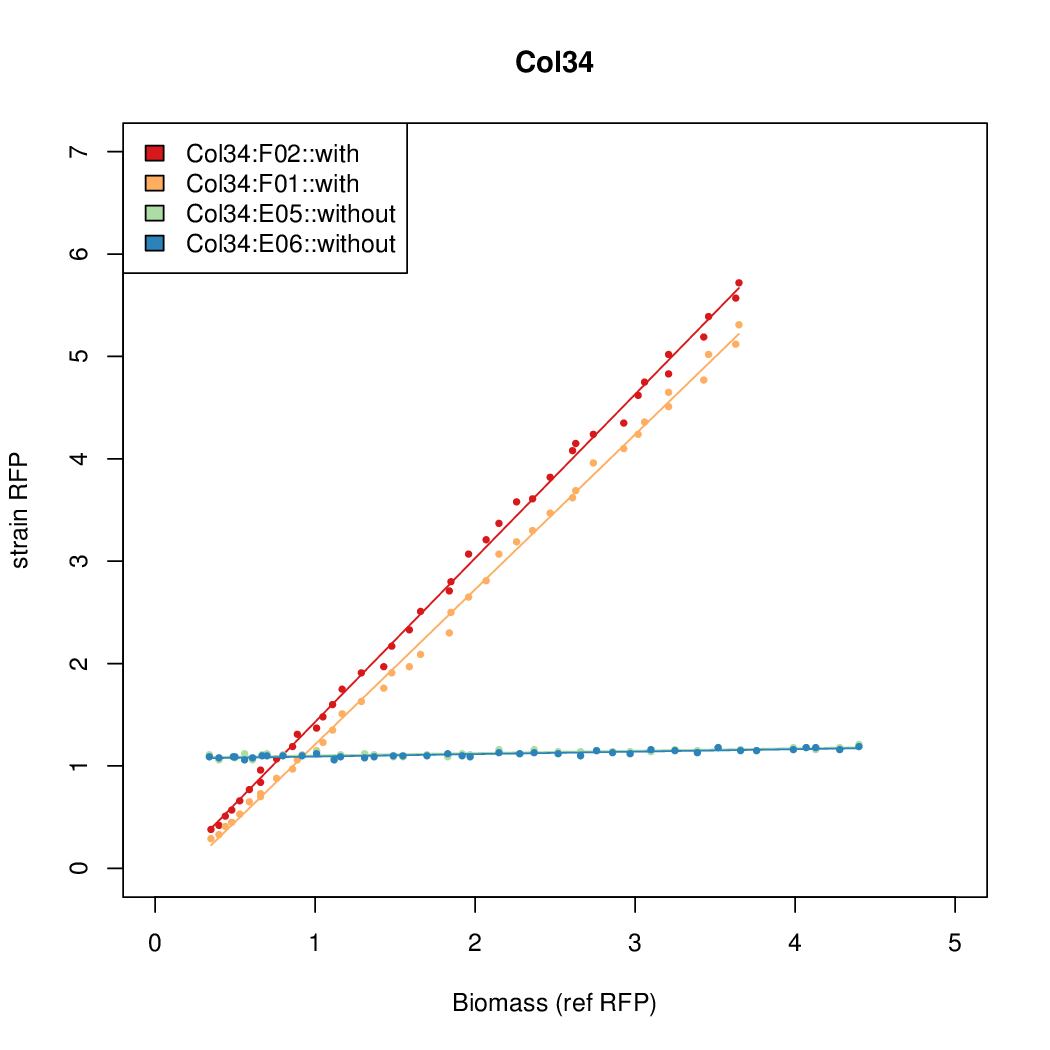
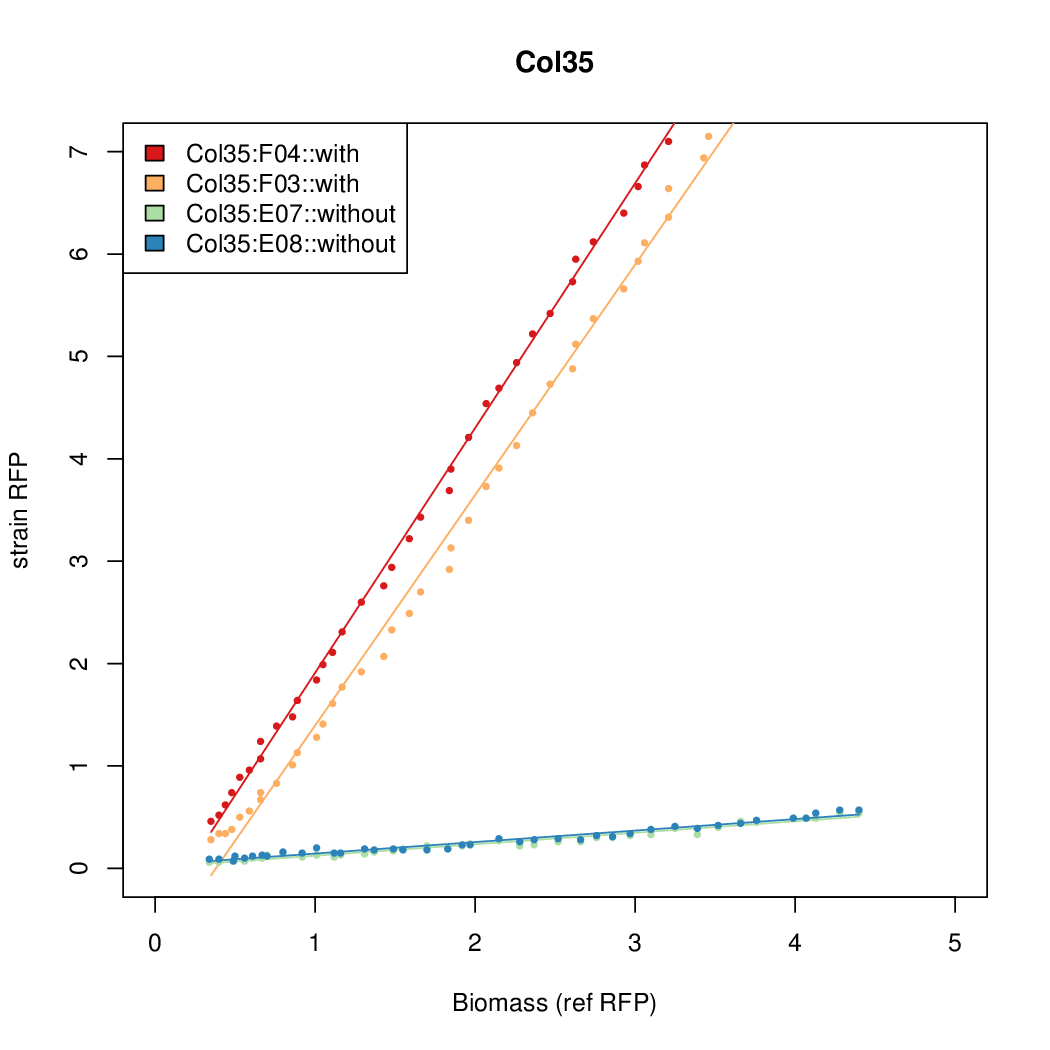
- The RFP measurement for a constitutively expressed strain was used as a standard measure of growth. This is plotted on the x-axis in the detailed plots per colony below.
- Figures were plotted using R.
Results
Summary
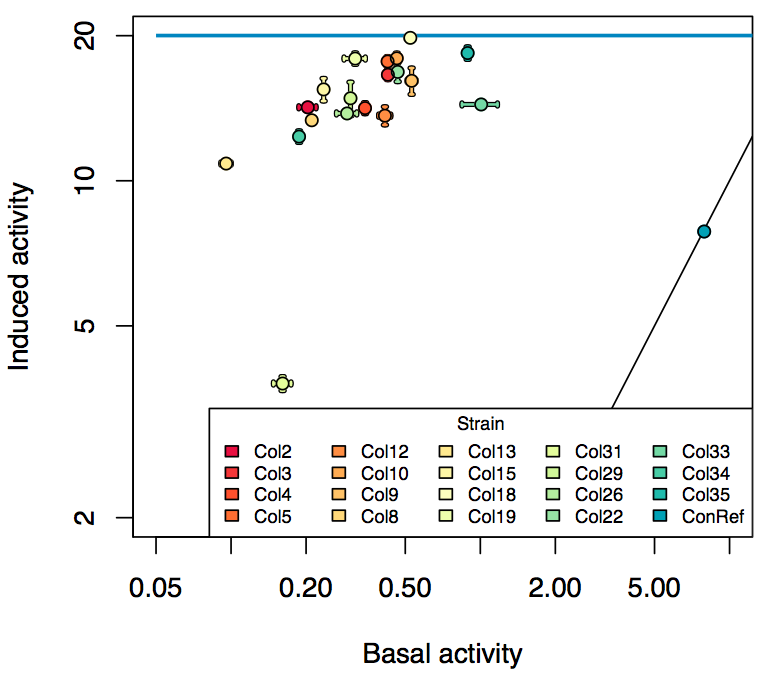
Promoter activity when induced (with arabinose added) plotted vs basal activity (without arabinose; ie leakiness of the promoter). The colonies that we selected all show less activity than the the constitutive promoter, and when induced, show higher activity than the constitutive promoter. The induced activity of the arabinose reference promoter is shown as a band representing two replicates.
Details
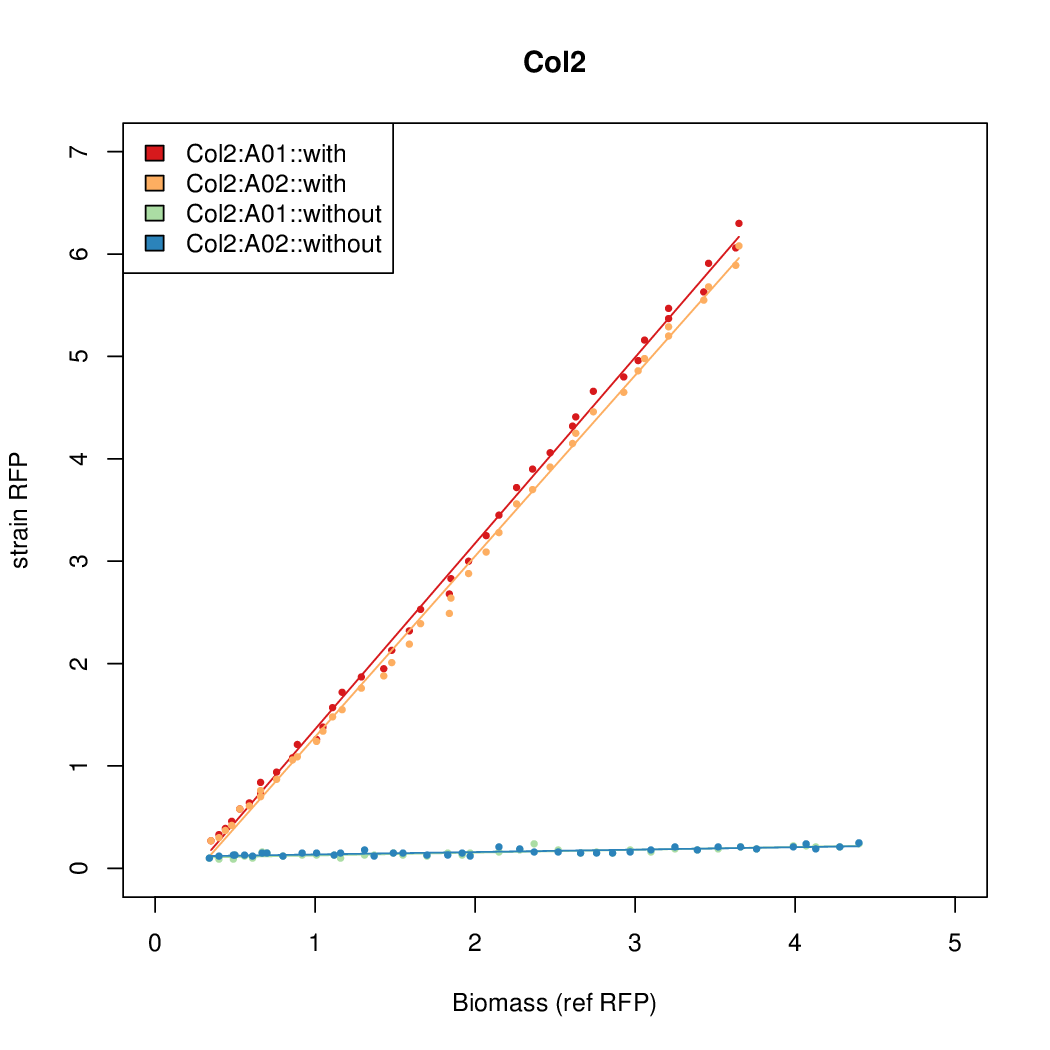
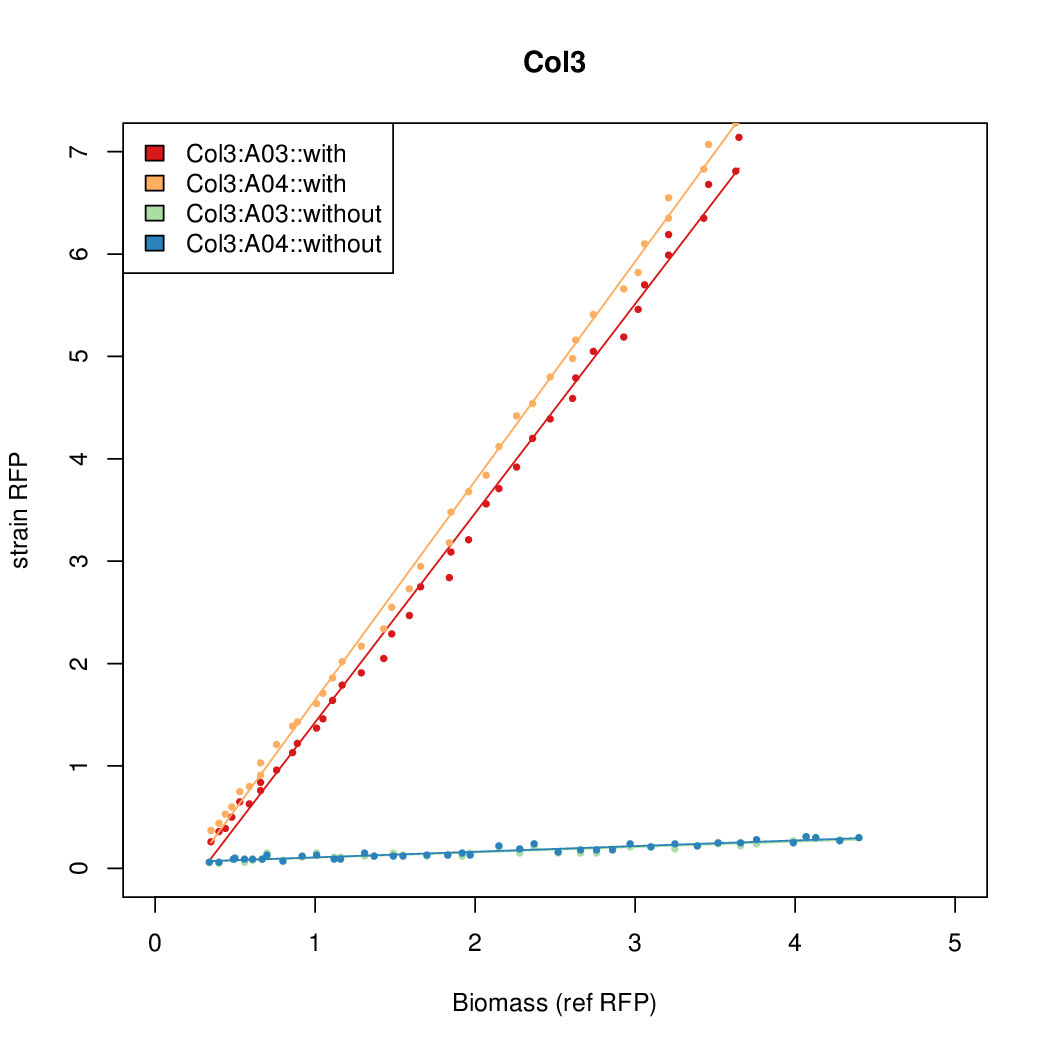
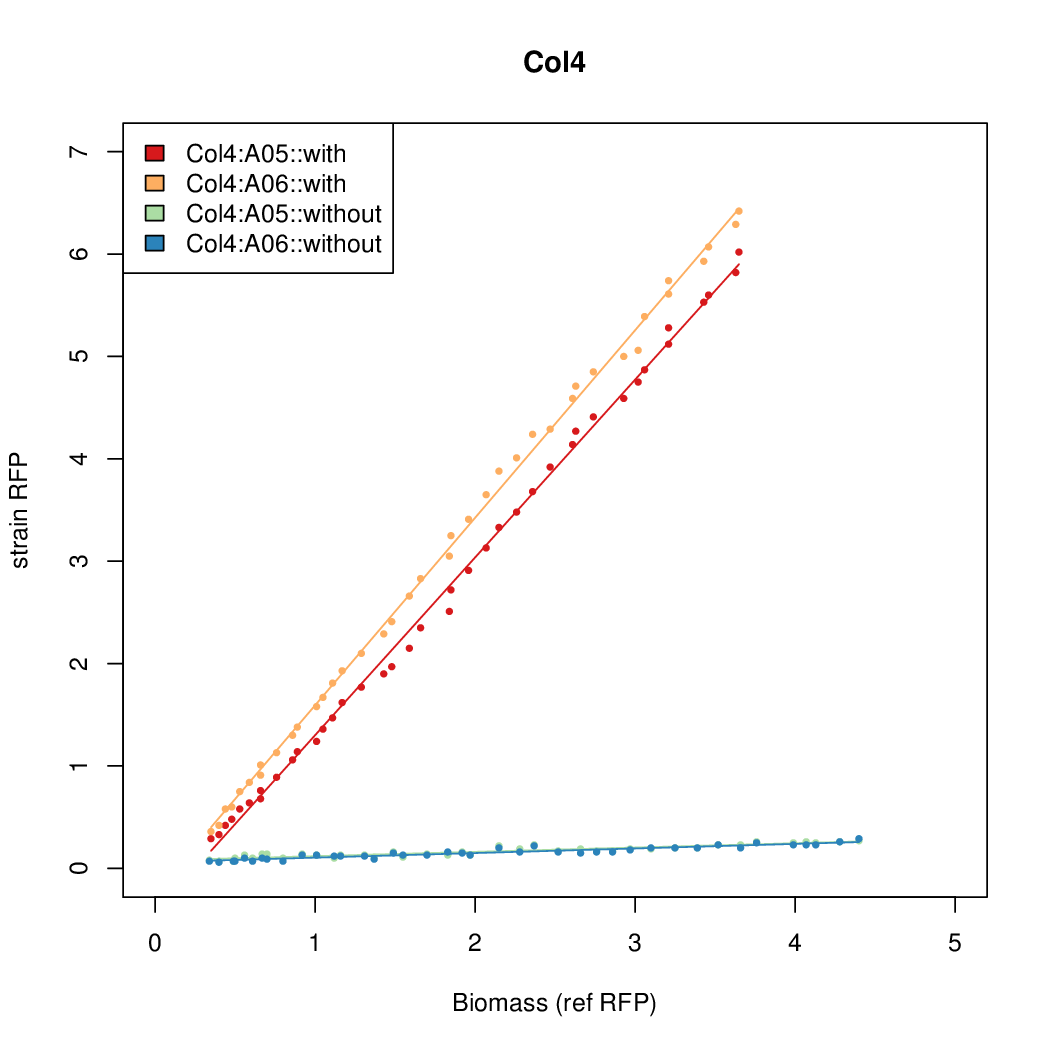
Promoter strengths for two trials of each colony with and without arabinose.
| Identifier | Sequencea | Induced Strength | Leakiness | Inducibilityb | ||
|---|---|---|---|---|---|---|
| Col. 2 | CTGACGACGCCCCTCTCCGCCCCTTAAAATGTCCA | 14.2 | 0.203 | 69.9 | ||
| Col. 3 | CTGACGTAATCATACCGCCGAAAGTATTATCATTA | 16.6 | 0.426 | 39 | ||
| Col. 4 | CTGACGCGACAACATTGCGTCCTATAAAATGCCGA | 14.2 | 0.345 | 41 | ||
| Col. 5 | CTGACGGCCCGCCCCCGATGCGCATAAAATACCAA | 17.7 | 0.424 | 41.6 | ||
| Col. 12 | CTGACGCGTATCGCGAGCGGGCGTTATTATACGCA | 13.6 | 0.414 | 32.9 | ||
| Col. 10 | CTGACGCCCCGAATCAGTAGTATTTATTATCTAGA | 18 | 0.462 | 38.8 | ||
| Col. 9 | CTGACGCCACCCCCCCCCCCGGCGTATAATTCCCA | 16.1 | 0.530 | 30.4 | ||
| Col. 8 | CTGACGCCCTACCGCTCGCCCCCCTATTATACCCA | 13.4 | 0.211 | 63.4 | ||
| Col. 13 | CTGACGCCCCCCCCGTCCACCCCCTAATATCCCGA | 10.9 | 0.096 | 113.6 | ||
| Col. 15 | CTGACGTGCGCCTGCCGTCCAAAGTAATATCCTTA | 15.5 | 0.235 | 65.8 | ||
| Col. 18 | CTGACGCCCCGAATCAGTAGTATTTATTATCTAGA | 19.8 | 0.524 | 37.8 | ||
| Col. 19 | CTGACGCTCGCACCGGCGACCCGATAATATCCATA | 18.0 | 0.315 | 57 | ||
| Col. 31 | CTGACGCCCCCCCTCCCCCGACCTAAAATATCCAG | 3.8 | 0.161 | 23.6 | ||
| Col. 29 | CTGACGCCGCTGCACCCGTCCCCCTATTATAGCAA | 14.8 | 0.301 | 49.2 | ||
| Col. 26 | CTGACGCCGATCCTATCTCCCTATTAAAATTTTTA | 13.8 | 0.292 | 47.2 | ||
| Col. 22 | CTGACGCCCCGAATCAGTAGTATTTATTATCTAGA | 16.8 | 0.466 | 36.1 | ||
| Col. 33 | CTGACGCCAGCTTCCTACTTCAAATATAATCCGTA | 14.4 | 1.01 | 14.3 | ||
| Col. 34 | CTGACGTCCCTCTTCCGCGCCCCCTAAAATACCCA | 12.4 | 0.187 | 65.9 | ||
| Col. 35 | CTGACGCCCAACCCGACACAGCGATATAATAATGA | 18.4 | 0.89 | 20.7 | ||
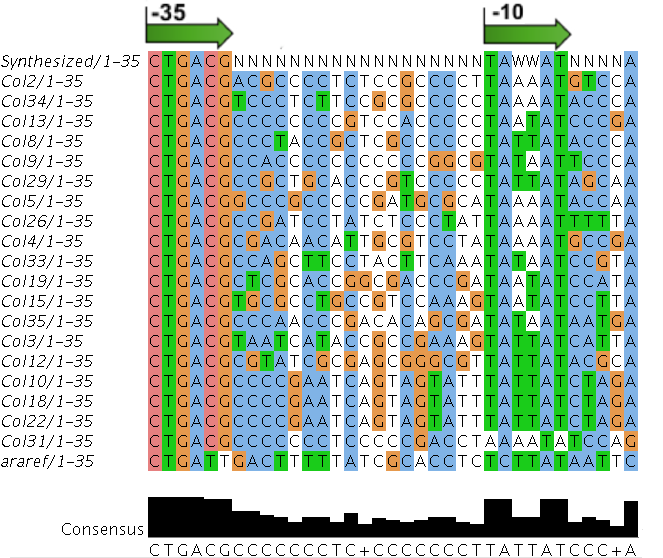
Sequences
We sequenced and aligned the promoters that are shown above. The sequences show conservation in the -10 and -35 regions by design. Within the -10 region, we allowed for two random weak (A or T) bases. We do not see a strong preference for A over T or vice versa. Within the regions where we allowed any base, we see a preference for C over the other bases. This could be due to bias during synthesis.
Example of use
The tight inducible pBAD promoter was used in our "Hello World project" to regulate the expression of GFP SF, which was tagged with a signal peptide to direct it into the periplasm. Production and folding of GFP SF is faster than the transport system of E. coli, which leads to undesired accumulation of GFP SF in the cytoplasm. Only when using a promoter with low leakiness it is possible to translocate a significant fraction of GFP SF after its production has been switched off. Thereby we get a clear signal from the periplasm with low interference from the cytoplasm.
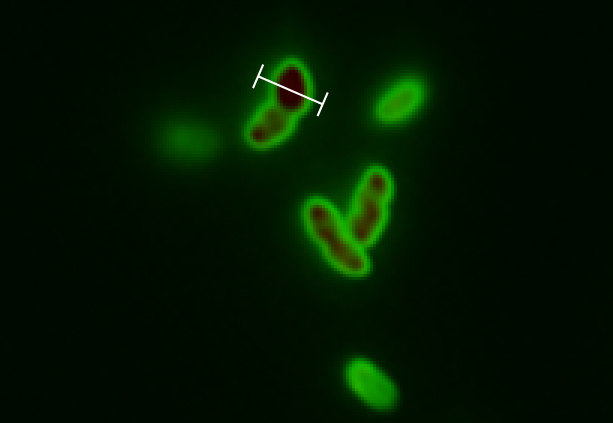
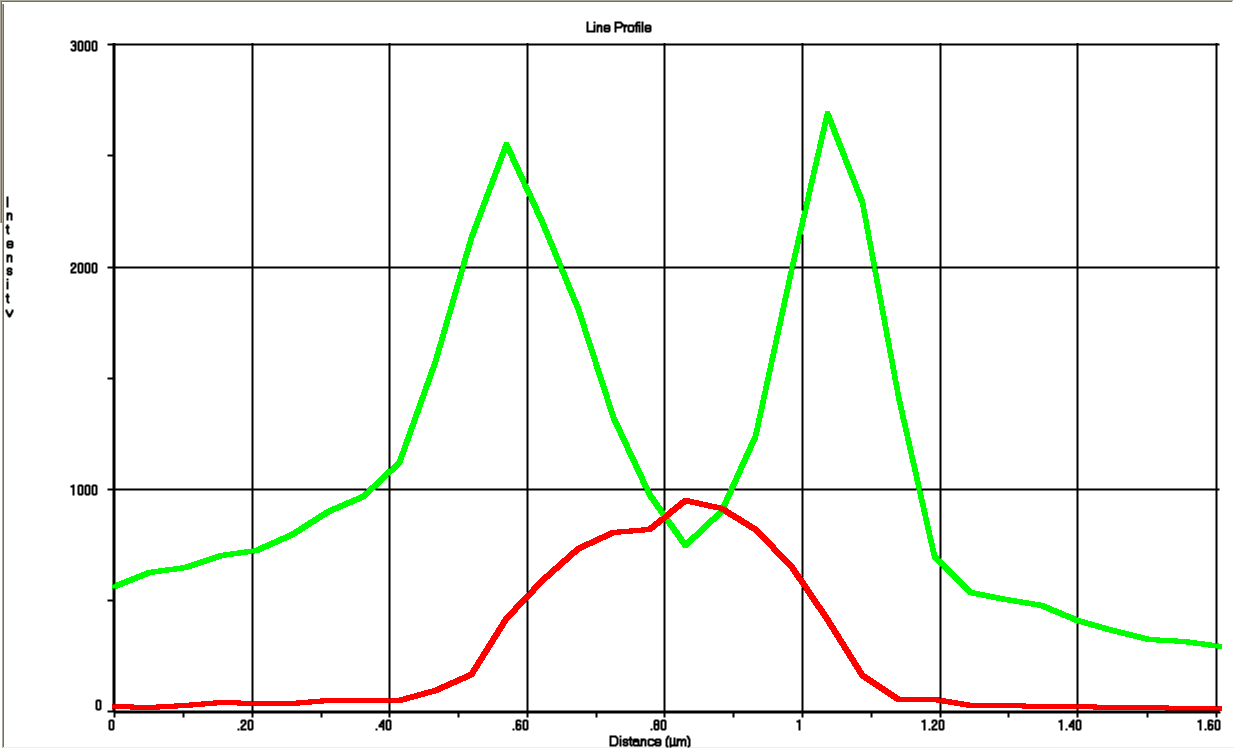
See also "Hello World project".
References
[1] Jensen, Peter Ruhdal, and Karin Hammer. "The sequence of spacers between the consensus sequences modulates the strength of prokaryotic promoters." Applied and environmental microbiology 64.1 (1998): 82-87.
 "
"