Team:Heidelberg/Templates/Indigoidine Introduction
From 2013.igem.org
Most modules of non-ribosomal peptide synthetase (NRPS) pathways consist of three domain types: condensation, adenylation and thiolation domain (see Figure 1a), also called peptidyl-carrier-protein domain (PCP)-domain (reviewed in <bib id="pmid16895337"/>)). Remarkably, precisely this order of domains is highly conserved among different NRPS pathways except for the very first and last module of each NRPS. In the initial module, the A-domain is always followed by a T-domain. The last module of an NRPS usually ends with a TE-domain. During the process of non-ribisomal peptide (NRP) synthesis, a new amino acid is first adenylated by the A-domain and then bound to the T-domain via a thioester bond. The C-domain catalyzes the condensation of the substrate - which is bound to the T-domain of the previous module - and the amino acid of the next module. The T-domain itself shows no substrate specificity but acts as a carrier domain, which keeps the peptide attached to the NRPS module complex. The core element of every T-domain is a conserved 4’-phosphopanthetheinylated (4’-PPT) serine. The 4’-PPT residue is added by a 4’-Phosphopanthetheinyl-transferase (PPTase), which converts the NRPS apo-enzyme to its active holo-form (see Figure 1b).
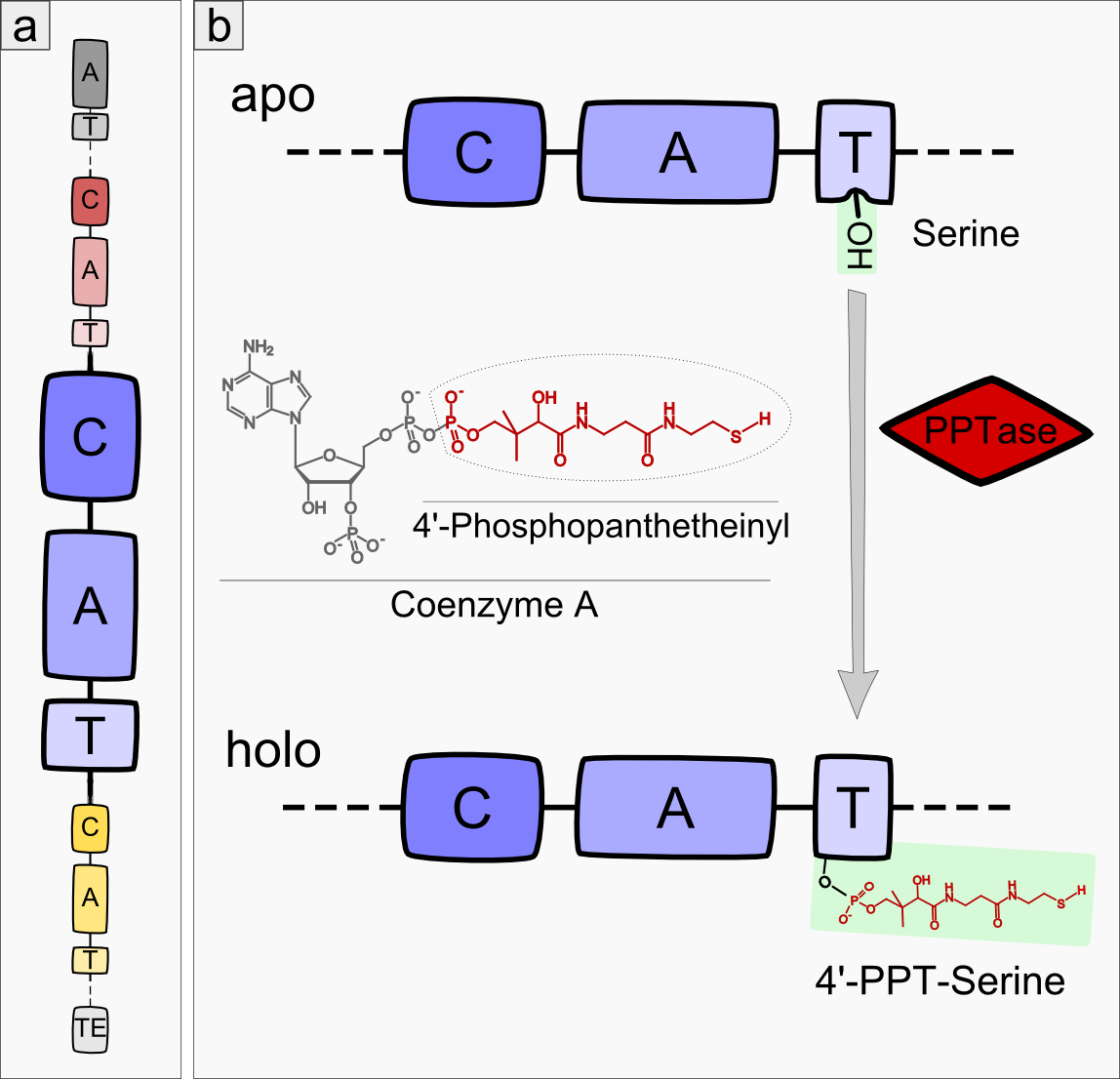 Besides these fundamental domains (C-domain, A-domain and T-domain), some NRPS modules incorporate additional domains enlarging the amount of potential catalytic reactions, such as cyclization, epimerization or oxidation of the amino acid [Reference].
Besides these fundamental domains (C-domain, A-domain and T-domain), some NRPS modules incorporate additional domains enlarging the amount of potential catalytic reactions, such as cyclization, epimerization or oxidation of the amino acid [Reference].
For example, a single module of P. luminescens laumondii TT01 (DSM15139) contains an internal oxidation domain (Ox-domain) in its A-domain and a special TE-domain (Figure 2a). This enzyme first adenylates L-glutamine (A-domain), which is then attached to the T-domain. The TE-domain cleaves and catalyzes the cyclization of the substrate, which is further oxidized by the Ox-domain. The oxidation of two cyclic glutamines results in the formation of an insoluble small molecule (Figure 2b)[Reference].
The small molecule produced by the pathway described above is a blue-colored pigment called indigoidine. Accordingly, the catalytic NRPS is referred to as indigoidine synthetase or blue pigment synthetase encoded by various bacterial strains such as S. lavendulae subsp. lavendulae (ATCC11924) or P. luminescens (<bib id="Takahashi2007"/><bib id="Brachmann2012"/>). Previous publications showed that replacing the T-domain of the blue pigment synthetase bpsA from S. lavendulae with T-domains of other NRPS modules results in a loss of function, i.e. the engineered indigoidine synthetase does not produce the blue pigment (<bib id="Owen2012"/). So far the only T-domain exchange that resulted in a functional bpsA was achieved using the T-domain of entF, - a gene involved in the enterobactin biosynthesis of E. coli. For this approach, random mutagenesis was performed to yield various mutated entF T-domains, some of which were capable to preserve the enzyme function when being introduced into bpsA (<bib id="Owen2012"/>). Other studies revealed that it is possible to exchange the A-domains of NRPS modules in B. subtilis resulting in modified non-ribosomal peptide products (<bib id="Doekel2000"/> ). Additionally, the selectivity of an NRPS module for specific amino acids could be modified by altering the conserved motif in the active site of the A-domain (<bib id="Thirlway2012"/>). Furthermore, since the endogenous 4'-Phoshopanthetheinyl-transferase (PPTase) entD from E. coli has been reported to exhibit low efficiency in activating heterologous NRPS pathways, most research in the field of NRPS involves co-expression of another PPTase (<bib id="Pfeifer2001"/>), bib id="Takahashi2007"/>).
 "
"