Template:Kyoto/Notebook/Sep 25
From 2013.igem.org
Sep 25
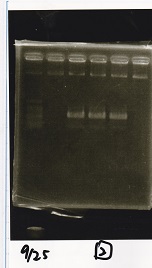
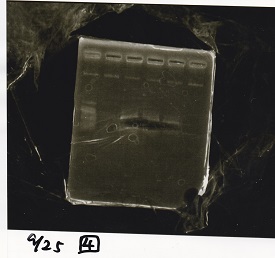
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 100bp ladder | -- | -- |
| 2 | Pcon-antisense (1C3-66E-fwd A2-662 rev) | -- | -- |
| 3 | DT (1C3-661-fwd 1C3 662 rev) | -- | -- |
| 4 | spinach-DT(1C3-661-fwd 1C3 662 rev) | -- | -- |
| 5 | Pcon-antisense-spinach-DT(1C3-66E fwd A2-661 rev) | -- | -- |
| 6 | 100bp ladder | -- | -- |
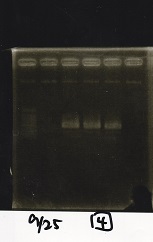
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 2 | 100bp ladder | -- | -- |
| 3 | Ptet (1C3 66E fwd-1C3 660 rev) | -- | -- |
| 4 | Pcon-attenuator(1C3-660 fwd A2-661 rev) | -- | -- |
| 5 | DT(1C3 662 fwd-1C3 661 rev) | -- | -- |
| 6 | Pcon-attenuator-aptamer-DT(1C3 66E fwd-A2-661-rev) | -- | -- |
| 7 | 100bp ladder | -- | -- |
Colony PCR
| Sample | base pair |
|---|---|
| 9/23 Pcon-PT181 attenuator-aptamer12-1R-DT-(5~12) | 859 |
| 9/23 Pcon-PT181 attenuator-RBS-GFP-DT-(5~8) | 1527 |
| 9/23 Pcon-PT181 attenuator-pSB1C3-(5~8) | 601 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94 °C | 94 °C | 55 °C | 68 °C | -- |
| 2 min | 30 s | 30 s | 1min5s | 30 cycles |
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | Pcon-PT181 attenuator-aptamer12-1R-DT-5 | -- | -- |
| 2 | Pcon-PT181 attenuator-aptamer12-1R-DT-6 | -- | -- |
| 3 | Pcon-PT181 attenuator-aptamer12-1R-DT-7 | -- | -- |
| 4 | Pcon-PT181 attenuator-aptamer12-1R-DT-8 | -- | -- |
| 5 | Pcon-PT181 attenuator-aptamer12-1R-DT-9 | -- | -- |
| 6 | Pcon-PT181 attenuator-aptamer12-1R-DT-10 | -- | -- |
| 7 | Pcon-PT181 attenuator-aptamer12-1R-DT-11 | -- | -- |
| 8 | Pcon-PT181 attenuator-aptamer12-1R-DT-12 | -- | -- |
| 9 | 1kbp ladder | -- | -- |
| 10 | Pcon-PT181 attenuator-RBS-GFP-DT-5 | -- | -- |
| 11 | Pcon-PT181 attenuator-RBS-GFP-DT-6 | -- | -- |
| 12 | Pcon-PT181 attenuator-RBS-GFP-DT-7 | -- | -- |
| 13 | Pcon-PT181 attenuator-RBS-GFP-DT-8 | -- | -- |
| 14 | Pcon-PT181 attenuator-pSB1C3-5 | -- | -- |
| 15 | Pcon-PT181 attenuator-pSB1C3-6 | -- | -- |
| 16 | Pcon-PT181 attenuator-pSB1C3-7 | -- | -- |
| 17 | Pcon-PT181 attenuator-pSB1C3-8 | -- | -- |
Transformation
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| Plac+tet aptamer12+Pcon tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181attenuator-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-Spinach-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181antisense+DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181attenuator+tet aptamer12-DT+Pcon-tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon tetR-DT+Ptet+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon tetR-DT+Pcon-GFP-DT+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-pT181attenuator-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Ptet-pT181antisense+Spinach-DT+Pcon-RBS-GFP-DT+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Pcon-pT181antisense+Spinach-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Pcon-tet aptamer12-DT+Ptet-pT181antisense+Spinach-DT+Pcon-pT181attenuator+RBS-GFP-DT | 1µL | 10µL | 11µL | -- |
| Name | Sample | Competent Cells | Total | Plate |
|---|---|---|---|---|
| aptamer12-DT(EcoRI+XbaI)+Pcon-pT181attenuator(EcoRI+SpeI) | 2µL | 20µL | 22µL | -- |
| Ptet(SpeI+PstI)+RBS-GFP-DT(XbaI+PstI)) | 2µL | 20µL | 22µL | -- |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/25 aptamer12_1R-DT(EcoRI&XbaI) | 4.5 µL | Pcon-pT181 attenuator(EcoRI&SpeI) | 7.7 µL | 6.1 µL |
| experiment | 9/17 pSB1C3(EcoRI&SpeI) | 2.0 µL | 9/3 pT181 attenuator(EcoRI&SpeI) | 14 µL | 8 µL |
| experiment | 9/25 Pcon-pT181 attenuator(EcoRI&XbaI) | 4.5 µL | 9/25 RBS-GFP-DT(EcoRI&SpeI) | 19.3 µL | 11.9 µL |
| experiment | 9/25 Pcon-RBS-GFP-DT(EcoRI&XbaI) | 4.2 µL | Pcon-RBS-tetR-DT(EcoRI&SpeI) | 20 µL | 12.1 µL |
Restriction Enzyme Digestion
| 9/24 Pcon-attenuator-aptamer-DT(µL) | EcoRI(µL) | SpeI(µL) | Buffer(µL) | MilliQ(µL) | total(µL) | |
|---|---|---|---|---|---|---|
| 2 cuts | 8.6 | 1 | 1 | 3 | 16.4 | 30 |
| NC | 0.4 | 0 | 0 | 1 | 8.6 | 10 |
| 9/11 DT(µL) | EcoRI(µL) | XbaI(µL) | Buffer(µL) | BSA(µL) | MilliQ(µL) | total(µL) | |
|---|---|---|---|---|---|---|---|
| 2 cuts | 10.4 | 1 | 1 | 3 | 3 | 11.6 | 30 |
| NC | 0.5 | 0 | 0 | 1 | 1 | 7.5 | 10 |
Liquid Culture
| Sample | medium |
|---|---|
| DT | -- |
| pT181attenuator | -- |
| Pcon-RBS-tetR-DT | -- |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 2 | Pcon-pT181antisense | -- |
| 3 | ||
| 4 | ||
| 5 | -- | -- |
| 6 | DT | -- |
| 7 | ||
| 8 | ||
| 9 | -- | -- |
| 10 | Spinach-DT | -- |
| 11 | ||
| 12 |
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | 100bp ladder | -- |
| 3 | Pcon-antisense-spinach-DT(E-1A) | -- |
| 4 | Pcon-antisense-spinach-DT(E-1A) | -- |
| 5 | Pcon-antisense-spinach-DT(E-1A) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 7 | 100bp ladder | -- |
| 9 | Ptet(E-0) | -- |
| 10 | Ptet(E-0) | -- |
| 11 | Ptet(E-0) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 13 | 100bp ladder | -- |
| 15 | Pcon-attenuator(0-1A) | -- |
| 16 | Pcon-attenuator(0-1A) | -- |
| 17 | Pcon-attenuator(0-1A) | -- |
| Lane | DNA | Enzyme |
|---|---|---|
| 19 | 100bp ladder | -- |
| 21 | Pcon-attenuator-aptamer-DT(E-1A) | -- |
| 22 | Pcon-attenuator-aptamer-DT(E-1A) | -- |
| 23 | Pcon-attenuator-aptamer-DT(E-1A) | -- |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| DT(2-1) | 24.9 | 1.73 | 0.37 |
| Ptet(E-0) | 12.8 | 1.59 | 0.90 |
| Pcon-antisense(E-2A) | 16.5 | 1.72 | 1.07 |
| Pcon-attenuator-aptamer-DT(E-1A) | 22.1 | 1.75 | 0.94 |
| Pcon-antisense-spinach-DT(E-1A) | 31.0 | 1.78 | 1.15 |
| spinach-DT(1-2) | 23.4 | 1.85 | 1.11 |
| Pcon-attenuator(0-1A) | 22.5 | 1.76 | 0.46 |
| DT(1-2) | 25.1 | 1.78 | 1.21 |
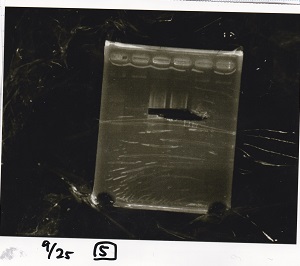
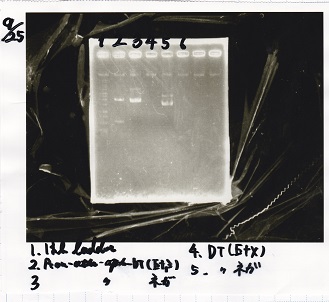
Electrophoresis
| Lane | Sample | Enzyme1 | Enzyme2 |
|---|---|---|---|
| 1 | 1kbp ladder | -- | -- |
| 2 | Pcon-pT181 attenuator-aptamer 12_1R-DT | EcoRI | SpeI |
| 3 | Pcon-pT181 attenuator-aptamer 12_1R-DT | -- | -- |
| 4 | DT | EcoRI | XbaI |
| 5 | DT | -- | -- |
Gel Extraction
| Lane | DNA | Enzyme |
|---|---|---|
| 1 | DT | EcoRI+XbaI |
| 2 | DT | EcoRI+XbaI |
| 3 | Pcon-pT181 attenuator-aptamer12_1R-DT | EcoRI+SpeI |
| 4 | Pcon-pT181 attenuator-aptamer12_1R-DT | EcoRI+SpeI |
| 5 | Pcon-pT181 attenuator-aptamer12_1R-DT | EcoRI+SpeI |
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-pT181 attenuator-aptamer12_1R-DT(EcoRI&SpeI) | 11.8 | 1.80 | 0.46 |
| DT(EcoRI+XbaI) | 42.6 | 1.84 | 0.98 |
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-pT181anntisense-spinach-DT(4k5) | - |
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/24 pSB1C3(EcoRI+Spel) | 10µL | 9/25 PconpT181attenuator-aptamer12-1R-DT(EcoRI+Spel) | 5.9µL | 8µL |
PCR
| 800pg/μL DT | KOD plus | 10x buffer | dNTP | MgSO4 | 1C3_GG2_fwd | iC3_GG1_rev | MilliQ | total |
|---|---|---|---|---|---|---|---|---|
| 1 | 0.5 | 2.5 | 2.5 | 1.5 | 0.75 | 0.75 | 15.5 | 25 |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 98°C | 57°C | 68°C | -- |
| 2min | 10s | 30s | 24s | 30cycles |
Transformation
| Name | Plate |
|---|---|
| aptamer12-1R-DT | CP |
| pSB1C3(EcoRI+SpeI)+pT181attenuator(EcoRI+Spal) | CP |
| Pcon-pT181attenuator(EcoRI+XbaI)+RBS-GFP-DT(EcoRI+SpeI) | Amp |
| Pcon-RBS-GFP-DT(EcoRI+XbaI)+Pcon-RBS-tetR-DT(EcoRI+SpeI) | Amp |
| pSB1C3(EcoRI+SpeI)+Pcon-pT181attenuator-aptamer12-1R-DT(EcoRI+SpeI) | CP |
Electrophoresis
Column Refining
| Name | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| 8/25 DT(2-1) | -- | -- | -- |
Restriction Enzyme Digestion
Concentrate PCR product by evaporator until volume become less than 3µL, add all sulution and pipetting.
| 9/25 DT(2-1) | 10*cut smart Buffer | BsaI-HF | MilliQ | total |
|---|---|---|---|---|
| 3.0 µL | 2.0 µL | 0.3 µL | 14.7 µL | 20.0 µL |
Incubate 37°C 12hour
Liquid Culture
| Sample | medium |
|---|---|
| Pcon-GFP-DT | LB(Amp) |
| RBS-GFP-DT | LB(CP) |
| Pcon-spinach-DT | LB(Amp) |
| Pcon-tetRaptamer-DT | LB(Amp) |
| Pcon-attenuator-DT | LB(Amp) |
| spinach-DT | LB(CP) |
| Pcon-antisense-spinach-DT | LB(Amp) |
Plating
Mix Liquid Culture 1mL and LB(+Amp), incubate 37°C 30min
| Sample | Use plate | The number of plate |
|---|---|---|
| Pcon-GFP-DT | M9(+Amp) | 3 |
| Pcon-GFP-DT | LB(+Amp) | 3 |
</div> incubate 37°C
Colony PCR
| Sample | base pair |
|---|---|
| 9/24 GGA-1(1~8) | 3177 |
| 9/24 GGA-2(1~8) | 3189 |
| 9/24 GGA-3(1~8) | 2989 |
| 9/24 GGA-4(1~8) | 2939 |
| 9/24 GGA-7(1) | 2489 |
| GGA-16(1) | 2717 |
| RBS-GFP-DT | -- |
| PreDenature | Denature | Annealing | Extension | cycle |
|---|---|---|---|---|
| 94°C | 94°C | 55°C | 68°C | -- |
| 5min | 30s | 30s | 3min15s | 30cycles |
RNA Extraction
sumple 0.25mL
Add 1mLISOGEN-LS
Lysis or homogenization
Store for 5 min, at room temperature
Add 0.2mL Chloroform
Shake vigorously for 15sec.
Store for 2~3min.at room temperature
Centrifuge 12K*g for 15min. at 4°C
Extract aqueous phase
Add 0.5mL isopropanol
Store for 5~10min. at room temperature
Centrifuge 12K*g for 10min. at 4°C
Remove aqueous phase
Add at least 1mL 70% ethanol
Vortex
Centrifuge 7.5K*g for 5min. at 4°C
Remove aqueous phase
Dry briefly
Add ddH2O, TE(pH8.0) or 0.5% SDS
Dissolve
Total RNA solution
| DNA | concentration[µg/mL] | 260/280 | 260/230 |
|---|---|---|---|
| Pcon-RBS-GFP-DT | 2783 | 1.95 | 1.48 |
Add 91.2µL, Store at freezer
Total 250µL</div>
cDNA Synthesis
| volume | 7 µL |
|---|---|
| 7*cDNA Wipcout Buffer | 1 µL |
| Template RNA(9/25 Pcon-RBS-GFP-DT) | 3 µL |
| RNase-frec water | 4 µL |
Incubate 42°C 2min, then on ice immediately
Add
| 5*Quantiscript Reverse Transcriptase | 0.5 µL |
|---|---|
| 5*Quantiscript RT Buffer | 2 µL |
| RT Primer Mix | 0.5µL |
Incubate 42°C 15min, then incubate 95°C 3min
Electrophoresis
Ligation
| state | Vector | Inserter | Ligation High ver.2 | ||
|---|---|---|---|---|---|
| experiment | 9/13 Pcon-pT181-attenuator(SpeI&PstI) | 3.5µL | 9/25 RBS-GFP-DT(XbaI&PstI) | 13.5µL | 8.5 µL |
 "
"