Team:Tsinghua-E/Part1
From 2013.igem.org
| (20 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
<html xmlns="http://www.w3.org/1999/xhtml"> | <html xmlns="http://www.w3.org/1999/xhtml"> | ||
<style type="text/css"> | <style type="text/css"> | ||
| - | #content{height: | + | #content{height:1800px;} |
| - | p{font-size: | + | p{font-size:130%} |
.memberx {position:absolute;top:60px;left:5px;height:215px;width;300px;} | .memberx {position:absolute;top:60px;left:5px;height:215px;width;300px;} | ||
.memberx span{z-index:50;position:relative;width:453px;display:block;top:-215px;left:305px;color:#000000} | .memberx span{z-index:50;position:relative;width:453px;display:block;top:-215px;left:305px;color:#000000} | ||
| Line 17: | Line 17: | ||
</div> | </div> | ||
| - | <div class="tmc" style="height: | + | <div class="tmc" style="height:1685px;"> |
</div> | </div> | ||
| - | <div class="neirong" style="height: | + | <div class="neirong" style="height:1665px;"> |
| - | + | ||
</html> | </html> | ||
| - | [[File: | + | |
| - | + | <h2>[[File:4_1s.png|68px|left]] '''Part 1: THU-E Mutation Part''' </h2> | |
| + | [[File:Plasmid1.png|446px|right]] | ||
| + | |||
| + | A plasmid used for the construction of high-diversity library in vivo ingenome level. In this vector, highly error-prone <em>dnaQ</em> mutant, <em>mutD</em><html><a href="#_ENREF_1" title="Lou, 2012 #499">[1]</a></html>was cloned downstream of <em>araBAD</em> promoter to control the mutation rate of the target genome by the concentration of <em>araBAD</em> promoter’s inducer, L-arabinose, in a strict manner.<em>E. Coli</em> JM109 carrying different vectors of pBAD_B0030-<em>mutD</em>-<em>sfGFP</em>, pBAD_B0032-<em>mutD</em>-<em>sfGFP</em> and pBAD_SDA_RBS-<em>mutD-sfGFP</em>(this RBS sequence was derived from the RBS sequence upstream of sfGFP in original AraC_pBAD_CI_OR222-sfGFP vector<html><a href="#_ENREF_2" title="Lou, 2012 #499">[2]</a></html>)were constructed. By detecting the induced fluorescence intensity, we found that pBAD_B0030-<em>mutD</em>-<em>sfGFP</em>, andpBAD_SDA_RBS-<em>mutD- sfGFP</em>have relatively higher <em>mutD</em> expression. The increaseof mutation rate induced by our mutation part was measured by quantifying the reversion of rifampinresistance caused by mutation in genome.pBAD_SDA_RBS-<em>mutD- sfGFP</em>could increase the genome mutation rate up to 10 times compared with negative control with 1g/L induction concentration of L-arabinose. <br /> | ||
| + | |||
<html> | <html> | ||
<p align="center"><br /> | <p align="center"><br /> | ||
| + | <img border="0" width="446" src="/wiki/images/a/a3/Part1II.png" /> <br /> | ||
| + | Figure.1 Conception illustration of the working mechanism of <em>mutD</em> | ||
<img border="0" width="446" src="/wiki/images/7/70/Part1I.png" /> <br /> | <img border="0" width="446" src="/wiki/images/7/70/Part1I.png" /> <br /> | ||
Figure.2 rifampicin reversion mutants caused by <em>mutD</em> expression and the counts by agar plate<br /> | Figure.2 rifampicin reversion mutants caused by <em>mutD</em> expression and the counts by agar plate<br /> | ||
| - | + | </p> | |
| - | + | ||
<br/> | <br/> | ||
| - | + | <br/> | |
| - | + | <p><font style="font-size: 12px">[1] Schaaper, R. M. Mechanisms of mutagenesis in the Escherichia-Coli mutator mutd5 - role of DNA mismatch repair. <em>Proc. Natl. Acad. Sci. U. S. A.</em> 85, 8126-8130,doi:10.1073/pnas.85.21.8126 (1988).</font></p> | |
| - | + | <p><font style="font-size: 12px">[2] Lou, C. B., Stanton, B., Chen, Y. J., Munsky, B. & Voigt, C. A. Ribozyme-based insulator parts buffer synthetic circuits from genetic context. <em>Nature Biotechnology</em> 30, 1137-+, doi:10.1038/nbt.2401 (2012).</font></p> | |
</div> | </div> | ||
</html> | </html> | ||
Latest revision as of 03:07, 28 September 2013



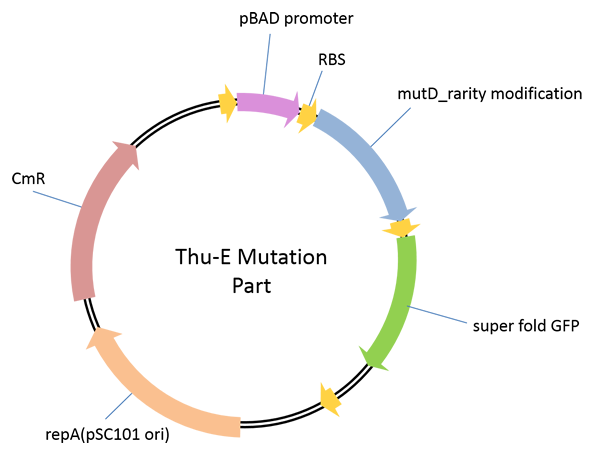
Part 1: THU-E Mutation Part
A plasmid used for the construction of high-diversity library in vivo ingenome level. In this vector, highly error-prone dnaQ mutant, mutD[1]was cloned downstream of araBAD promoter to control the mutation rate of the target genome by the concentration of araBAD promoter’s inducer, L-arabinose, in a strict manner.E. Coli JM109 carrying different vectors of pBAD_B0030-mutD-sfGFP, pBAD_B0032-mutD-sfGFP and pBAD_SDA_RBS-mutD-sfGFP(this RBS sequence was derived from the RBS sequence upstream of sfGFP in original AraC_pBAD_CI_OR222-sfGFP vector[2])were constructed. By detecting the induced fluorescence intensity, we found that pBAD_B0030-mutD-sfGFP, andpBAD_SDA_RBS-mutD- sfGFPhave relatively higher mutD expression. The increaseof mutation rate induced by our mutation part was measured by quantifying the reversion of rifampinresistance caused by mutation in genome.pBAD_SDA_RBS-mutD- sfGFPcould increase the genome mutation rate up to 10 times compared with negative control with 1g/L induction concentration of L-arabinose.

Figure.1 Conception illustration of the working mechanism of mutD
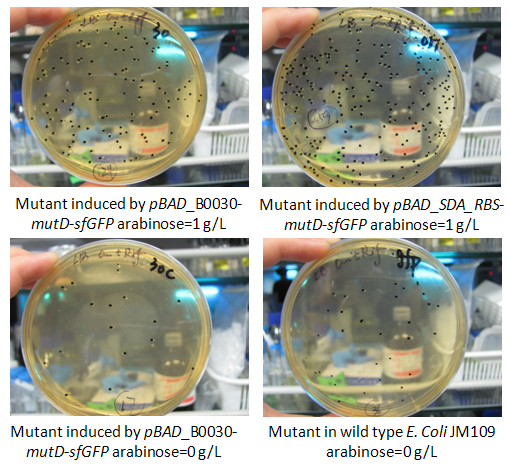

Figure.2 rifampicin reversion mutants caused by mutD expression and the counts by agar plate
[1] Schaaper, R. M. Mechanisms of mutagenesis in the Escherichia-Coli mutator mutd5 - role of DNA mismatch repair. Proc. Natl. Acad. Sci. U. S. A. 85, 8126-8130,doi:10.1073/pnas.85.21.8126 (1988).
[2] Lou, C. B., Stanton, B., Chen, Y. J., Munsky, B. & Voigt, C. A. Ribozyme-based insulator parts buffer synthetic circuits from genetic context. Nature Biotechnology 30, 1137-+, doi:10.1038/nbt.2401 (2012).
 "
"