Team:DTU-Denmark/Protein Models
From 2013.igem.org
(→NOR) |
|||
| Line 1: | Line 1: | ||
{{:Team:DTU-Denmark/Templates/StartPage|Protein Models}} | {{:Team:DTU-Denmark/Templates/StartPage|Protein Models}} | ||
<div class="overviewPage"> | <div class="overviewPage"> | ||
| - | It turns out that the structures of some of the proteins we want our mutants to express have been determined. | + | It turns out that the structures of some of the proteins we want our mutants to express have been determined. While others could be homology modelled due to existing 3D models that had high enough similarity to the ones that we are working with. |
==Mutant 1== | ==Mutant 1== | ||
| - | |||
===AMO=== | ===AMO=== | ||
Ammonia monooxygenase | Ammonia monooxygenase | ||
Revision as of 22:23, 1 October 2013
Protein Models
It turns out that the structures of some of the proteins we want our mutants to express have been determined. While others could be homology modelled due to existing 3D models that had high enough similarity to the ones that we are working with.
Contents |
Mutant 1
AMO
Ammonia monooxygenase
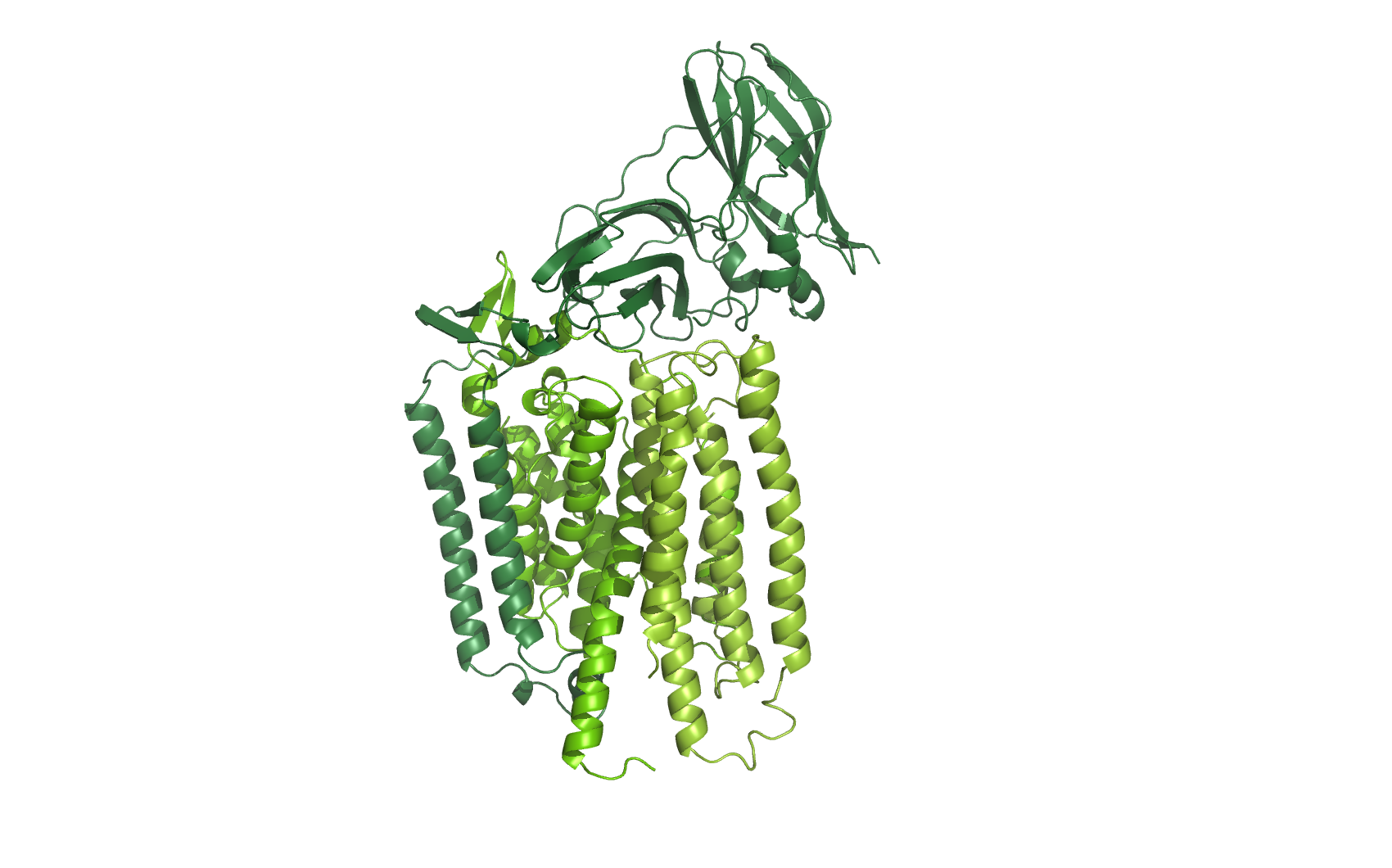
homology models of all three subunits of AMO
At the time of writing neither of the the three subunits from Nitrosomonas Europae is available as 3D structures, so each of the subunits was identified at UNIPROT, and their sequence was homologymodelled using [http://www.cbs.dtu.dk/services/CPHmodels/ CPHmodels-3.2]
| Resource | AmoA | AmoB | AmoC |
|---|---|---|---|
| Uniprot | [http://www.uniprot.org/uniprot/Q04507 Q04507] | [http://www.uniprot.org/uniprot/Q04508 Q04508] | [http://www.uniprot.org/uniprot/Q82T63 Q82T63] |
| Template pdb code and chain | [http://www.pdb.org/pdb/explore/explore.do?structureId=1yew 1YEW.A] | [http://www.pdb.org/pdb/explore/explore.do?structureId=1yew 1YEW.B] | [http://www.pdb.org/pdb/explore/explore.do?structureId=3rgb 3RGB.C] |
| Coverage | 90.5 | 86.2 | 66.4 |
| E-value | 1e-87 | 3e-67 | 8e-43 |
The three subunits were combined and aligned to the subunits of the [http://www.pdb.org/pdb/explore/explore.do?structureId=1yew Crystal structure] of particulate methane monooxygenase, hence an overlap of two alpha helices.
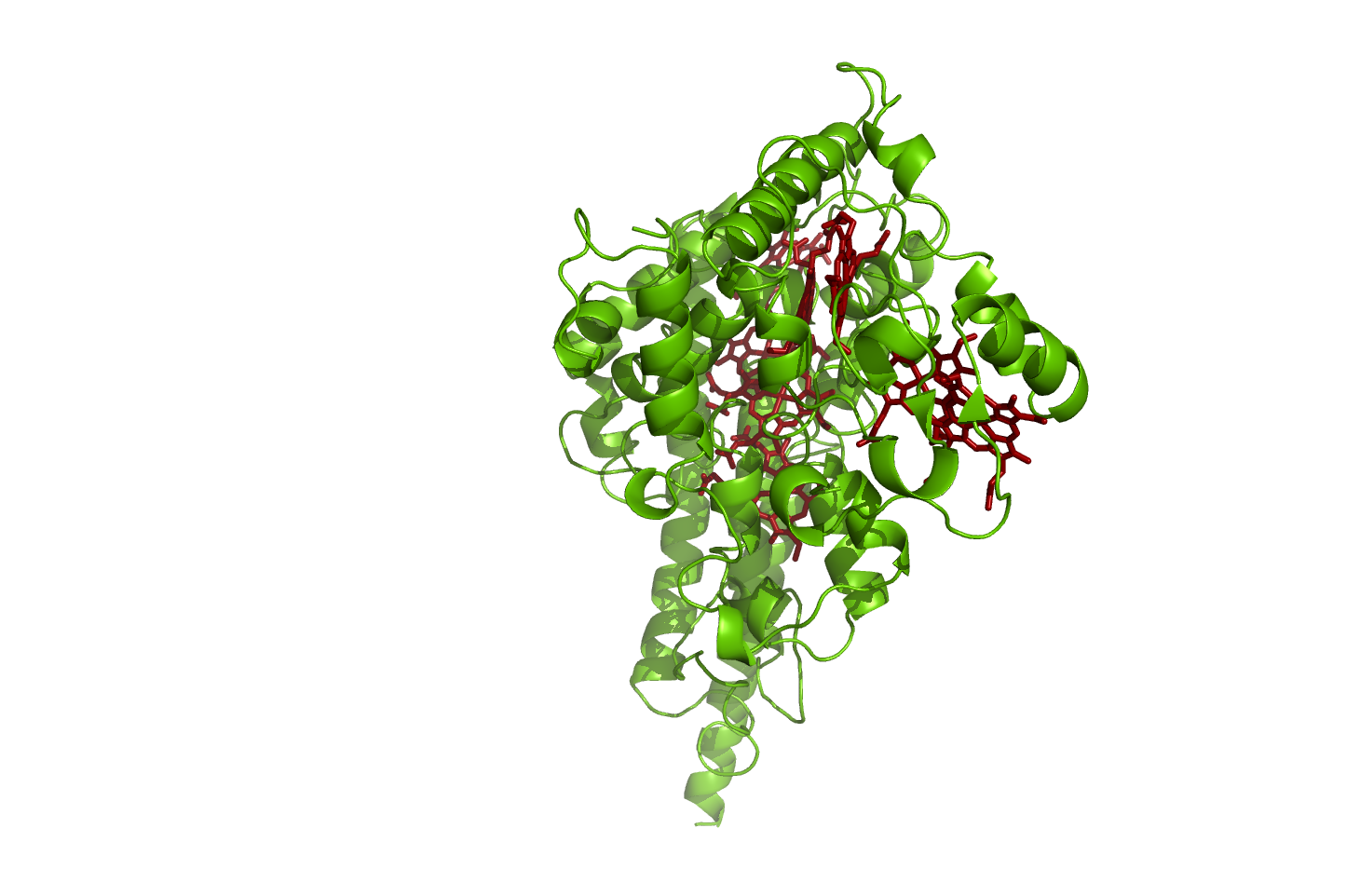
Hao
Hydroxylamine oxidoreductase
source [http://www.rcsb.org/pdb/explore.do?structureId=1fgj 1FGJ]
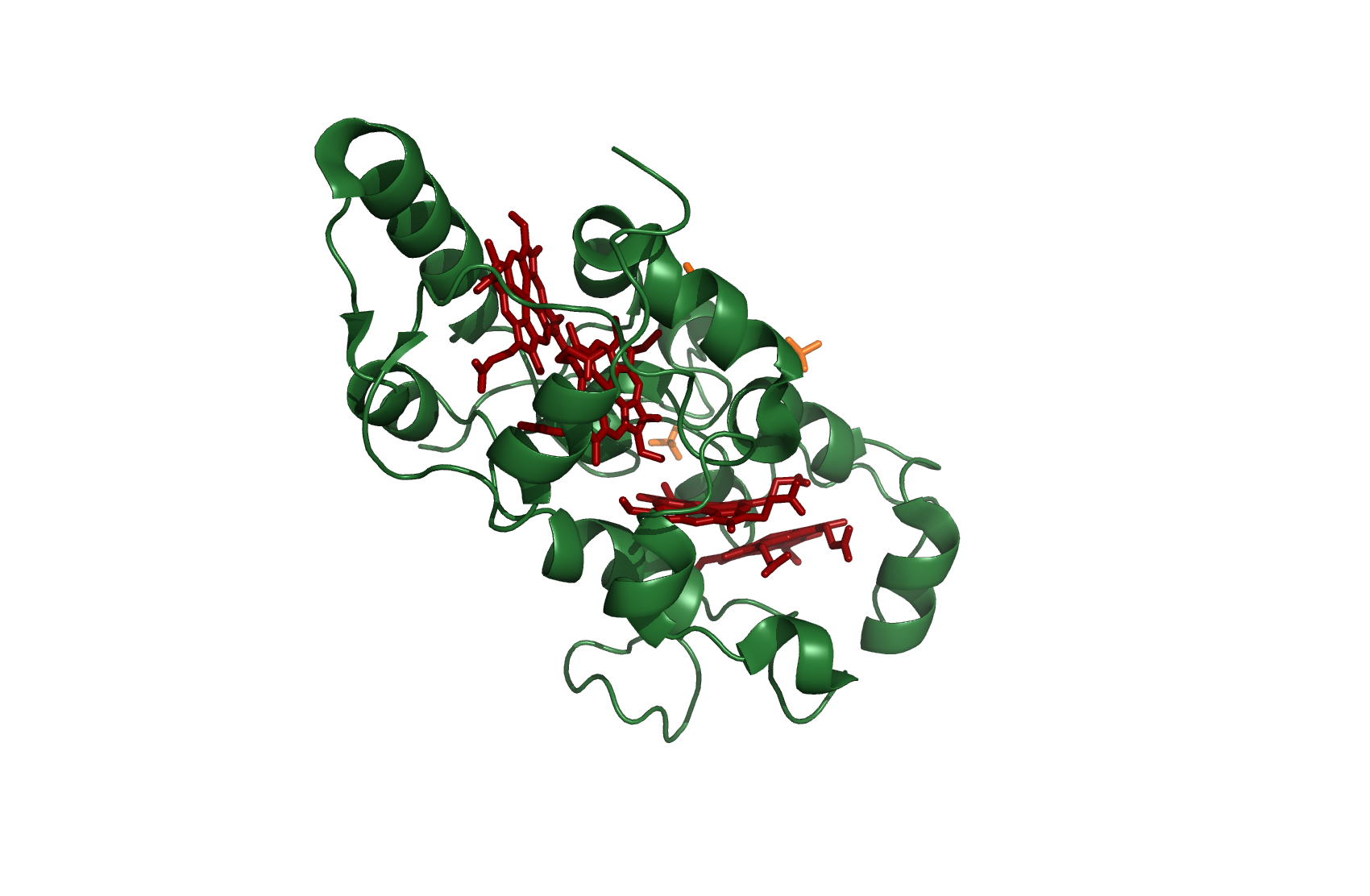
Cc554
Cytochrome c554
source [http://www.rcsb.org/pdb/explore.do?structureId=1ft5 1FT5]
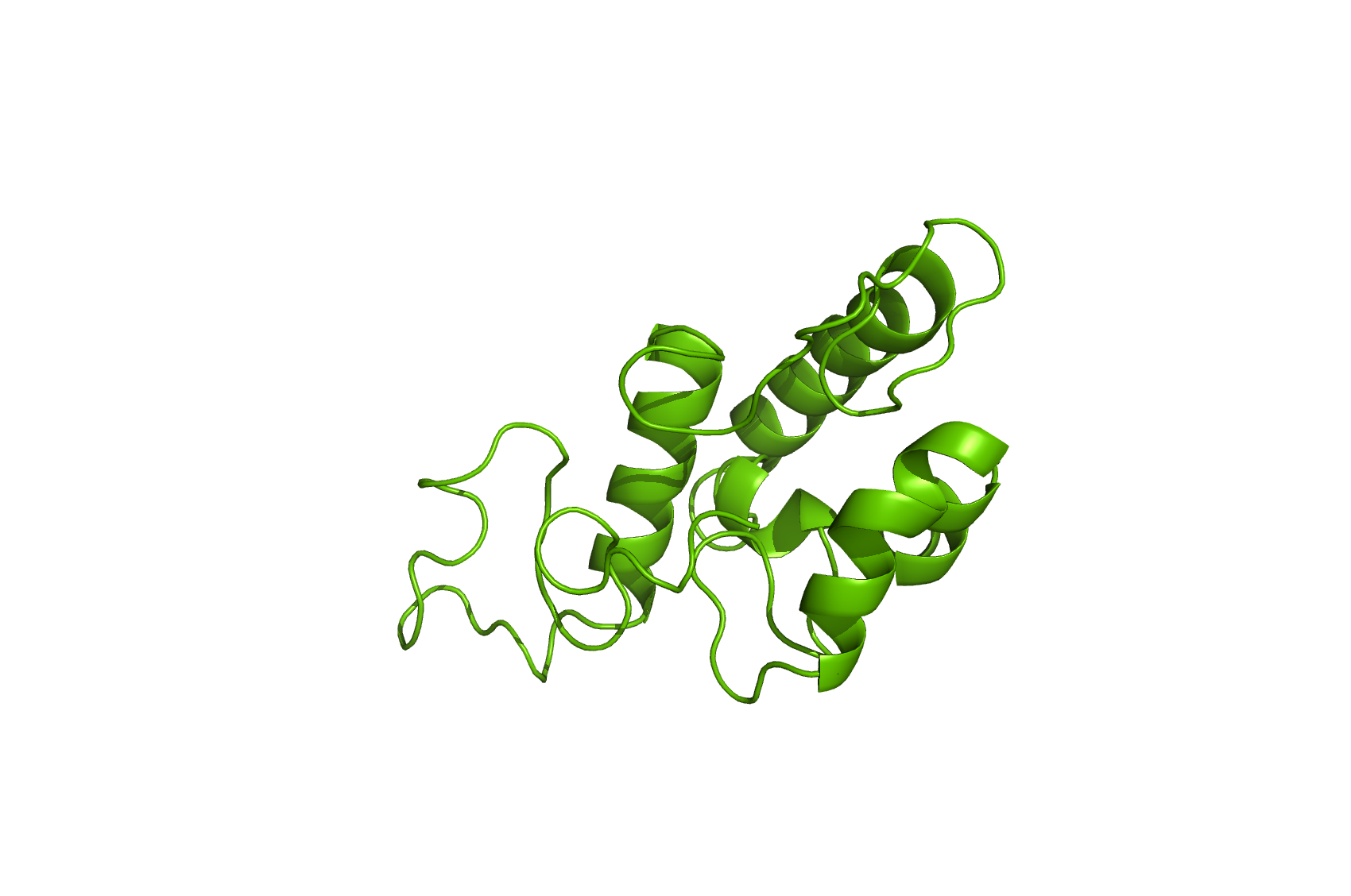
Ccm552
Homology model of Ccm552
| Resource | Ccm552 |
|---|---|
| Uniprot | [http://www.uniprot.org/uniprot/Q50926 Q50926] |
| Template pdb code and chain | [http://www.pdb.org/pdb/explore/explore.do?structureId=1j7a 2J7A.O] |
| Coverage | 68.3 |
| E-value | 6e-11 |
Mutant 2
NirS
Nitrite reductase
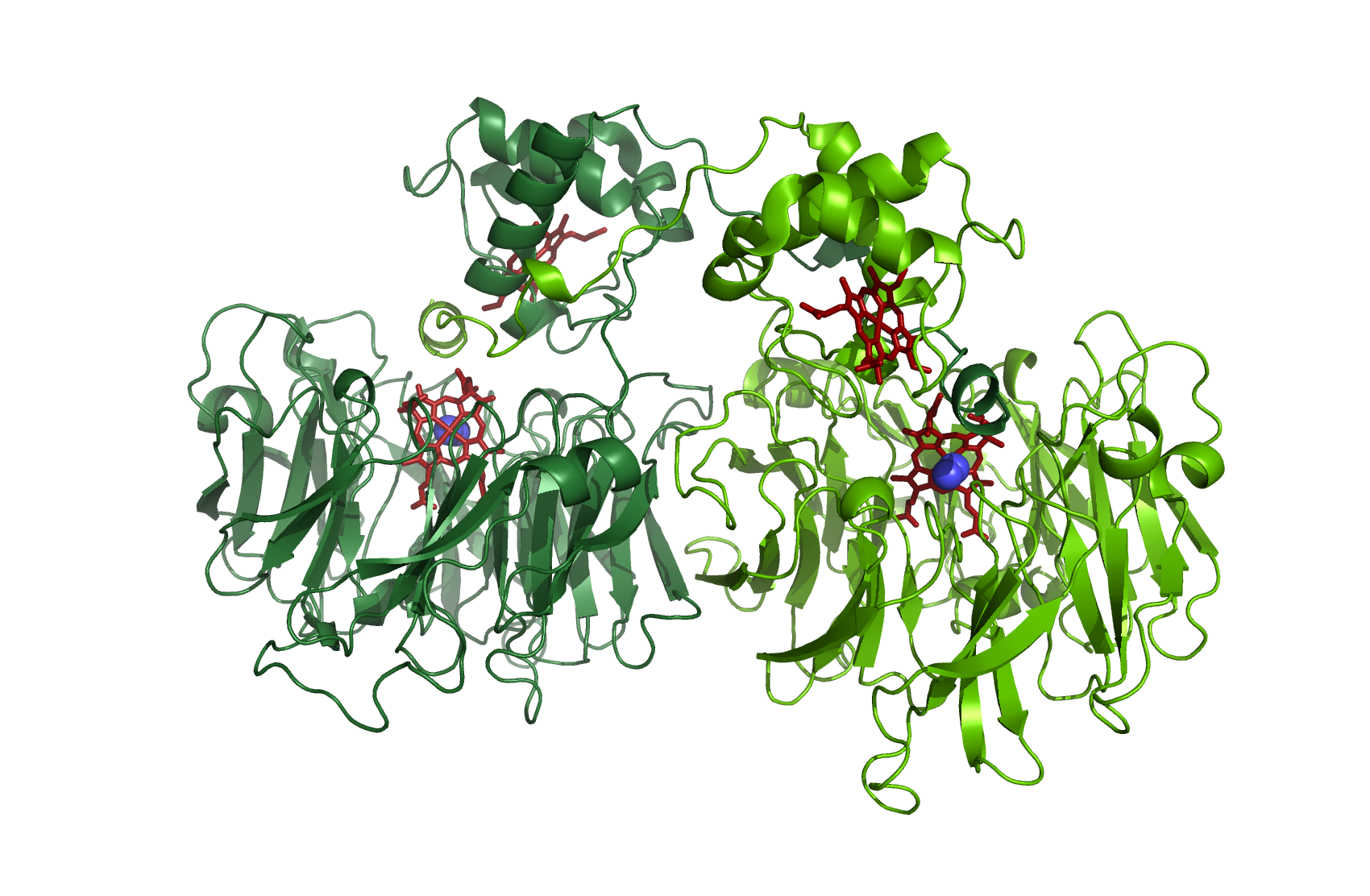
As a dimer with N2 - source [http://www.rcsb.org/pdb/explore/explore.do?structureId=1nno 1NNO]
NirM
Cytochrom c551
source [http://www.rcsb.org/pdb/explore/explore.do?structureId=2exv 2EXV]
NOR
Flavorubredoxin
Homology model of NOR
| Resource | NOR |
|---|---|
| Uniprot | [http://www.uniprot.org/uniprot/Q46877 Q46877] |
| Template pdb code and chain | [http://www.pdb.org/pdb/explore/explore.do?structureId=1ycf 1YCF.A] |
| Coverage | 82.5 |
| E-value | 3e-95 |
- All homology models were made using the free online tool [http://www.cbs.dtu.dk/services/CPHmodels/ CPHmodels v3.2]
- All models were made and visualized in [http://sourceforge.net/projects/pymol/ PyMOL v1.6]
 "
"