Team:NCTU Formosa/biobricks
From 2013.igem.org
(Difference between revisions)
(→BBa_K1017301) |
(→BBa_K1017781) |
||
| (66 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
{{:Team:NCTU Formosa/source/header}} | {{:Team:NCTU Formosa/source/header}} | ||
{{:Team:NCTU Formosa/source/cover-biobricks}} | {{:Team:NCTU Formosa/source/cover-biobricks}} | ||
| - | + | __NOTOC__ | |
| + | <div class="li" style="clear:both"><div class="card"> | ||
===Parts submitted to the Registry=== | ===Parts submitted to the Registry=== | ||
| - | <div class="li"><div class="card"> | + | <groupparts>iGEM013 NCTU_Formosa</groupparts> |
| + | </div></div> | ||
| + | |||
| + | <div class="li" style="clear:both"><div class="card"> | ||
| + | ===Brief Information=== | ||
| + | Please click on the name of the parts for detailed information that is hosted in the Registry website. | ||
====Light-regulated system==== | ====Light-regulated system==== | ||
| - | ====== | + | ======BBa_K1017726====== |
| - | * | + | *pcya and ho1 are the enzymes needed to convert heme into chromophore phycocyanobiline(PCB), the red light sensor. Since pcya and ho1 are not naturally produced in E.coli, we use P<sub>cons</sub> upstream in order to make E.coli continuously expressed them to synthesize PCB. |
| - | <br>[[File: | + | <br>[[File:Nctu_pcya_ho1.jpg|450px]] |
======BBa_K1017301====== | ======BBa_K1017301====== | ||
| - | + | *Cph8 is a chimeric light receptor. It is a fusion of the photoreceptor cph1 and the envZ histidine kinase. cph1 is only active when it binds the chromophore phycocyanobiline (PCB). | |
| - | * | + | <br>[[File:Nctu_cph8.jpg|150px]] |
| - | + | ||
| - | + | ||
| - | + | ||
======BBa_K1017781====== | ======BBa_K1017781====== | ||
| - | *[[File:Nctu_lacI_plac.jpg| | + | *This is the key part to change Ompc promoter intro red promotor. The lacI will inhibit the function of P<sub>lac</sub>, thus, the original outcomes will be converted into our expected result. |
| + | <br>[[File:Nctu_lacI_plac.jpg|300px]] | ||
| - | ====== | + | ======BBa_K1017101====== |
| - | *[[File: | + | *Ompc promtor, which can sense red light with the presence of cph8. It will be turned on in the dark ,and be turned off in the bright. For the convenient use, we add lacI and lac promotor downstream the biobrick. This part, so called red promotor, can be activated under red light, and inactive in the dark. |
| - | + | <br>[[File:Nctu_Pred.jpg|450px]] | |
| - | + | ||
| - | + | ||
====Temperature-regulated system==== | ====Temperature-regulated system==== | ||
======BBa_K1017602====== | ======BBa_K1017602====== | ||
| - | *[[File:Nctu_37rbs_mGFP.jpg| | + | *By using mGFP as a reporter gene, we can test whether the 37 °C RBS works. |
| + | <br>[[File:Nctu_37rbs_mGFP.jpg|200px]] | ||
======BBa_K1017603====== | ======BBa_K1017603====== | ||
| - | *[[File:Nctu_37rbs_luxr.jpg|200px]] | + | *In our circuit, this biobrick is the part of P<sub>lux</sub>'s activation when the temperature reaches to 37<sup>o</sup>C. |
| - | + | <br> | |
| + | [[File:Nctu_37rbs_luxr.jpg|200px]] | ||
| - | + | ====Small RNA-regulated system==== | |
| - | ==== | + | ======BBa_K1017403====== |
| - | + | *The sRNA is the complement of its rRBS. It can regulate the downstream of rRBS in RNA level by binding onto the rRBS when it is transcribed in order to interrupt ribosomes' work. In addition, adding Plux upstream makes the sequence be controlled by luxR/AHL complex.<br> | |
| - | *The sRNA | + | [[File:Nctu_plux_srna1.jpg|250px]] |
| - | <br>[[File: | + | |
======BBa_K1017404====== | ======BBa_K1017404====== | ||
| - | *[[File:Nctu_srna2.jpg| | + | *The sRNA is the complement of its rRBS. It can regulate the downstream of rRBS in RNA level by binding onto the rRBS when it is transcribed in order to interrupt ribosomes' work.<br>[[File:Nctu_srna2.jpg|200px]] |
| + | |||
| + | ======BBa_K1017202====== | ||
| + | *The sRNA, base pair with target mRNA, including the Shine-Dalgarno sequence. Thus it prevent ribosome from binding to initiate the translation. The rRBS is designed for sRNA perfect binding, and this rRBS is the RBS which can be bound only for our artificial sRNA(BBa_K1017404).<br> | ||
| + | [[File:Nctu_rbs2.jpg|150px]] | ||
| + | |||
| + | ======BBa_K1017811====== | ||
| + | *The sequence let us show the efficiency of the sRNA-rRBS binding by expressing the fluorescence with the combination of a sequence providing the transcription of the sRNA. | ||
| + | [[File:Nctu_pcons_rbs2_mrfp.jpg|350px]] | ||
======BBa_K1017401====== | ======BBa_K1017401====== | ||
| - | *This part, BBa_K1017401, includes our artificial sRNA-1 and rRBS-1. The non-coding small RNA can bind to the Shine-Dalgarno sequence on rRBS-1 by base-pairing. Once the rRBS-1 is blocked, ribosomes cannot bind to it to translate, thus, gene expressions downstream are decreased. Because of specific binding, rRBS-1 can only be bound by sRNA-1. We add P<sub>lux</sub> upstream, so this part can be regulated by luxR/AHL complex. | + | *This part, BBa_K1017401, includes our artificial sRNA-1 and rRBS-1. The non-coding small RNA can bind to the Shine-Dalgarno sequence on rRBS-1 by base-pairing. Once the rRBS-1 is blocked, ribosomes cannot bind to it to translate, thus, gene expressions downstream are decreased. Because of specific binding, rRBS-1 can only be bound by sRNA-1. We add P<sub>lux</sub> upstream, so this part can be regulated by luxR/AHL complex. There is one important thing hasn't be mentioned is that it is a temporary sequence which contains the restriction enzyme cutting sites of SpeI, EcoRI and XbaI in order for us to separate them.<br> |
| - | <br>[[File:Nctu_sRNA_rRBS1.jpg| | + | [[File:Nctu_sRNA_rRBS1.jpg|350px]] |
======BBa_K1017402====== | ======BBa_K1017402====== | ||
| - | *This part, BBa_K1017402, is similar to BBa_K1017401 mentioned above, but without P<sub>lux</sub>. rRBS-2 can only be bound by sRNA-2 due to specificity, then gene expressions downstream are decreased. | + | *This part, BBa_K1017402, is similar to BBa_K1017401 mentioned above, but without P<sub>lux</sub>. rRBS-2 can only be bound by sRNA-2 due to specificity, then gene expressions downstream are decreased. There is one important thing hasn't be mentioned is that it is a temporary sequence which contains the restriction enzyme cutting sites of SpeI, EcoRI and XbaI in order for us to separate them.<br> |
| - | <br>[[File:Nctu_srna2_rRNS.jpg| | + | [[File:Nctu_srna2_rRNS.jpg|250px]] |
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</div> | </div> | ||
</div> | </div> | ||
| + | <html> | ||
</div> | </div> | ||
</div> | </div> | ||
Latest revision as of 01:21, 29 October 2013
Biobricks
To strive for the best results, we made a number of experiment. Our team always hang together in spite of many losses.
Parts submitted to the Registry
<groupparts>iGEM013 NCTU_Formosa</groupparts>
Brief Information
Please click on the name of the parts for detailed information that is hosted in the Registry website.
Light-regulated system
BBa_K1017726
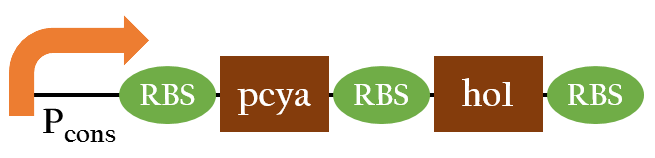
- pcya and ho1 are the enzymes needed to convert heme into chromophore phycocyanobiline(PCB), the red light sensor. Since pcya and ho1 are not naturally produced in E.coli, we use Pcons upstream in order to make E.coli continuously expressed them to synthesize PCB.
BBa_K1017301
- Cph8 is a chimeric light receptor. It is a fusion of the photoreceptor cph1 and the envZ histidine kinase. cph1 is only active when it binds the chromophore phycocyanobiline (PCB).
BBa_K1017781
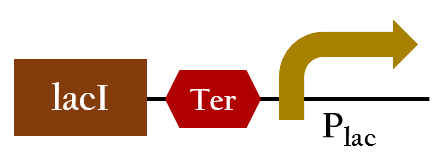
- This is the key part to change Ompc promoter intro red promotor. The lacI will inhibit the function of Plac, thus, the original outcomes will be converted into our expected result.
BBa_K1017101
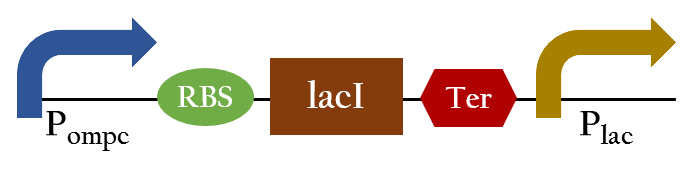
- Ompc promtor, which can sense red light with the presence of cph8. It will be turned on in the dark ,and be turned off in the bright. For the convenient use, we add lacI and lac promotor downstream the biobrick. This part, so called red promotor, can be activated under red light, and inactive in the dark.
Temperature-regulated system
BBa_K1017602
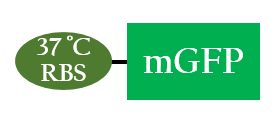
- By using mGFP as a reporter gene, we can test whether the 37 °C RBS works.
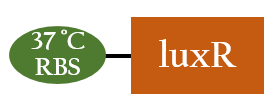
BBa_K1017603
- In our circuit, this biobrick is the part of Plux's activation when the temperature reaches to 37oC.
Small RNA-regulated system
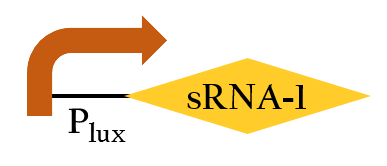
BBa_K1017403
- The sRNA is the complement of its rRBS. It can regulate the downstream of rRBS in RNA level by binding onto the rRBS when it is transcribed in order to interrupt ribosomes' work. In addition, adding Plux upstream makes the sequence be controlled by luxR/AHL complex.
BBa_K1017404
- The sRNA is the complement of its rRBS. It can regulate the downstream of rRBS in RNA level by binding onto the rRBS when it is transcribed in order to interrupt ribosomes' work.

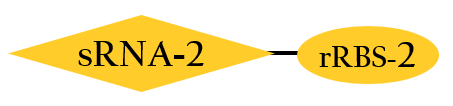
BBa_K1017202
- The sRNA, base pair with target mRNA, including the Shine-Dalgarno sequence. Thus it prevent ribosome from binding to initiate the translation. The rRBS is designed for sRNA perfect binding, and this rRBS is the RBS which can be bound only for our artificial sRNA(BBa_K1017404).
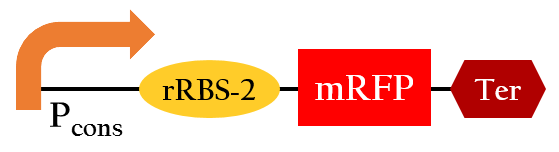
BBa_K1017811
- The sequence let us show the efficiency of the sRNA-rRBS binding by expressing the fluorescence with the combination of a sequence providing the transcription of the sRNA.
BBa_K1017401
- This part, BBa_K1017401, includes our artificial sRNA-1 and rRBS-1. The non-coding small RNA can bind to the Shine-Dalgarno sequence on rRBS-1 by base-pairing. Once the rRBS-1 is blocked, ribosomes cannot bind to it to translate, thus, gene expressions downstream are decreased. Because of specific binding, rRBS-1 can only be bound by sRNA-1. We add Plux upstream, so this part can be regulated by luxR/AHL complex. There is one important thing hasn't be mentioned is that it is a temporary sequence which contains the restriction enzyme cutting sites of SpeI, EcoRI and XbaI in order for us to separate them.
BBa_K1017402
- This part, BBa_K1017402, is similar to BBa_K1017401 mentioned above, but without Plux. rRBS-2 can only be bound by sRNA-2 due to specificity, then gene expressions downstream are decreased. There is one important thing hasn't be mentioned is that it is a temporary sequence which contains the restriction enzyme cutting sites of SpeI, EcoRI and XbaI in order for us to separate them.
 "
"