Team:Tianjin/Project/Design&Construction
From 2013.igem.org
(→b. what cause the leakage (from modeling information)) |
(→b. what cause the leakage (from modeling information)) |
||
| Line 344: | Line 344: | ||
[[Image:TJU-PROJECT-D4.jpg|thumb|400px|center|<b>Figure 3.</b> The result of the influence of two factors. a:Vector without RFP gene. b:Vector with PalkM-RFP part. c:Strain with both PalkM-RFP and Pc-AlkR parts.]] | [[Image:TJU-PROJECT-D4.jpg|thumb|400px|center|<b>Figure 3.</b> The result of the influence of two factors. a:Vector without RFP gene. b:Vector with PalkM-RFP part. c:Strain with both PalkM-RFP and Pc-AlkR parts.]] | ||
| - | [[Image:TJU-Project-D4-2.jpg|thumb|500px|center|<b>Figure 4.</b> ]] | + | [[Image:TJU-Project-D4-2.jpg|thumb|500px|center|<b>Figure 4.</b> Mathematical Analysis of infering the reason of leakage.]] |
Besides the high expression of AlkR, the low activity of the terminator downstream of the AlkR is also the source of leakage. In this condition, the transcription of constitutive promoter Pc may turn on the expression of RFP. | Besides the high expression of AlkR, the low activity of the terminator downstream of the AlkR is also the source of leakage. In this condition, the transcription of constitutive promoter Pc may turn on the expression of RFP. | ||
Revision as of 01:29, 29 October 2013
Contents |
First Trail
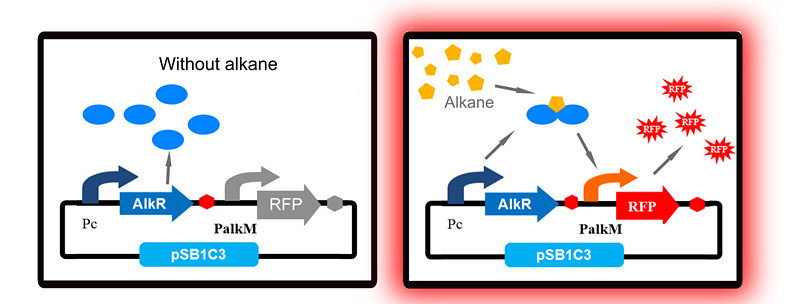
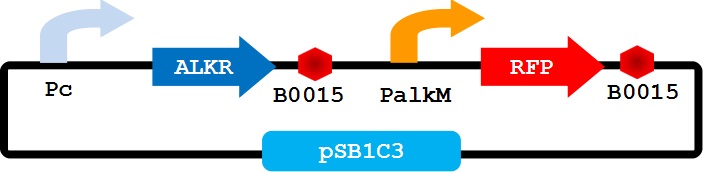
To apply the special alkane sensing system AlkR-PalkM of Acinetobacter baylyi ADP1 in E. coli, we built a gene circuit as Figure 1 showed. We used a constitutive promoter to control the expression of regulator AlkR, and RFP is controlled by PalkM. Without alkanes, the PalkM will not work, RFP will not be expressed. In the presence of alkanes, AlkR dimer will bind to the alkane molecules to form a complex, and the complex can turn on the expression of RFP which is controlled by PalkM. Then red fluorescence can be detected by naked eyes and measured by fluorescence spectrophotometer easily. Considering that another module may be put in E. coli to produce alkanes, we put Pc-AlkR-tt and PalkM-RFP-tt into one plasmid to form our Alk-Sensor.
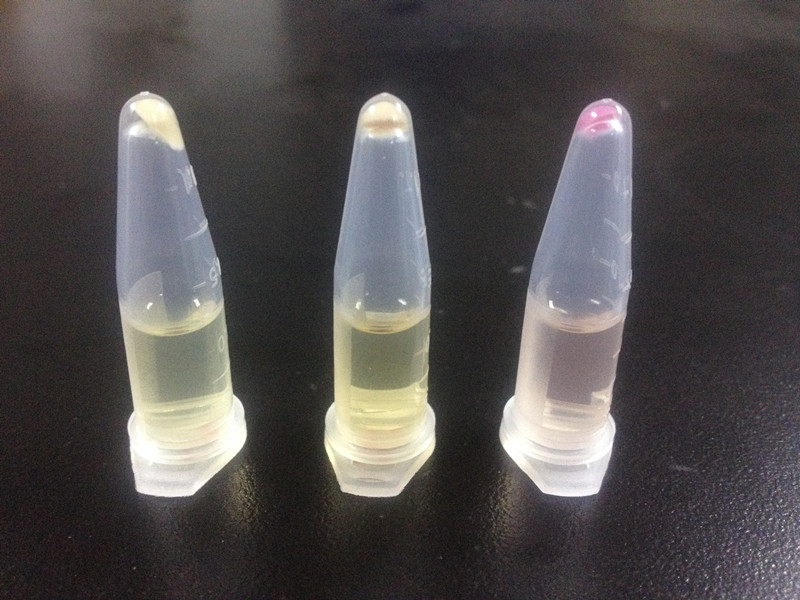
Unfortunately, our first trial has a big problem: even in the absence of alkanes, strains with our Alk-Sensor turn red, which can be detected by naked eyes, as Fig 2 shows. The leakage of our Alk-Sensor is very high. This will influence alkane detection by naked eyes, so the Alk-Sensor need to be improved.
Analysis
a. what's the influence of the high leakage
To find out the source of the problem, we built a mathematical model for the alkane sensing process of AlkR-PalkM system. (see more details for the process of modeling) The modeling result reflects the relationship between input (alkane concentration) and output (RFP concentration), showed in Figure 4. From the Figure, we find that, when leakage of the AlkR-PalkM system is very high, the dynamic range will be small. This will make it hard for us to distinguish the change of the alkane concentration, so it is crucial to improve the function of our Alk-Sensor to reduce the leakage.
b. what cause the leakage (from modeling information)
From the equation 1, which is acquired from mathematical analysis, we can find the source of leakage. In the equation, the [Pmt] and [AlkR] can be easily controlled through changing the copy number of the plasmid or strength of promoter, so we choose these two factors as our target to improve our Alk-Sensor.
To evaluate the influence of two factors, we test the part PalkM-RFP in a high copy number plasmid pSB1C3 without the expression of AlkR. The result shows in Figure 3, we can see that the strain just containing PalkM-RFP isn’t red, and the strain with both PalkM-RFP and Pc-AlkR turns red. So the leakage is mainly caused by the high expression of AlkR and PalkM is a very weak promoter without the induction of AlkR.
Besides the high expression of AlkR, the low activity of the terminator downstream of the AlkR is also the source of leakage. In this condition, the transcription of constitutive promoter Pc may turn on the expression of RFP.
Summary of the leakage of RFP:
a)Promoter alkM has some leakage, to be more specific, it binds to RNA polymerase without undergoing a conformation change and activates the transcription.
b)Protein alkR or dimerized alkR could induce promoter alkM without alkane molecules.
c)The terminator downstream of AlkR is weak,and the constitutive promoter upstream of AlkR may influence the expression of RFP.
Reconstruction
To improve the function of the Alk-Sensor and decrease the leakage, we reconstructed our device.
We used a weaker promoter to control the expression of AlkR. And then, to reduce the interference of two transcription units, we used a stronger terminator to the downstream of two transcription units.
At last, we switched the position of the two transcription unit, to guarantee that PalkM-RFP isn’t interferenced by constitutive promoter. The new reconstructed Alk-Sensor shows in Figure 5.
Through another blank test we found that the reconstruction could decrease AlkSensor’s leakage significantly. After optimization, the percentage of bacterial colonies that are red enough to be distinguished by naked eyes decreased from 95% to 15%.
At last, we used full dose octane to induce the two sensors. After a certain period of cultivation, the relative fluorescent intensity was measured. The results showed that after reconstruction the leakage decreased by 50%, and the dynamic range increased by 5 folds, which means that the optimized AlkSensor has stronger ability to distinguish different inputs. ( showed in Figure 6)
So we choose this construction as our final design for Alk-Sensor and one of our favorite biobricks.
 "
"