Team:TU-Munich/Notebook/Labjournal
From 2013.igem.org
LouiseFunke (Talk | contribs) (→Week 4) |
LouiseFunke (Talk | contribs) (→Wednesday, May 15th`) |
||
| Line 3,170: | Line 3,170: | ||
</div> | </div> | ||
| - | == '''Wednesday, May 15th | + | == '''Wednesday, May 15th''' == |
| + | |||
| + | |||
| + | |||
</div> | </div> | ||
Revision as of 17:27, 15 May 2013

Labjournal
Week 1
Monday, April 22nd
Transformation of E. coli XL1 blue with Phytochrome B (2-908 N-terminal amino acids) (BBa_K801031, RFC25, pSB1C3)
Investigator: Jeff, Leonie, Rosario
Aim of the experiment: Transformation of Phytochrome B for protein fusion.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 2 µl of DNA was added to 100 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Miniprep of pTUM100 with pGAL, pTEF1, pTEF2, pADH and RFC25 compatible RFP generator
Investigator: Jeff, Leonie, Rosario
Aim of the experiment: Miniprep of pTUM100 with pGAL, pTEF1, pTEF2, pADH and RFC25 compatible RFP generator
Procedure:
- Miniprep was performed after manufacturer's protocol (QIAprep Miniprep, QIAGEN)
Sequencing of RFP-Generator (RFC25, pSB1C3)
Investigator: Jeff, Leonie, Rosario
Aim of the experiment: Sequencing of RFP-Generator (RFC25, pSB1C3)
Procedure:
Sequencing batch were prepared after manufacturer's protocol. (15 µl of plasmid DNA (50 - 100 ng) and 2 µl sequencing primer)
Tuesday, April 23rd
Picking of of E. coli XL1 blue with Phytochrome B (2-908 N-terminal amino acids) (BBa_K801031, RFC25, pSB1C3)
Investigator: Jeff, Leonie, Rosario, Florian
Aim of the experiment: Picking of of E. coli XL1 blue with Phytochrome B (2-908 N-terminal amino acids) (BBa_K801031, RFC25, pSB1C3)
Procedure:
- pSB1C3 plasmid with BBa_K801031 (PhyB 2 - 908 aa, RFC25): Colonies were picked from chloramphenicol plates.
- Picked pipette tips was transferred into cell-culture tubes with air-permeable, sterile cover. Each tube contain 4 mL of LB-medium + 4 µL chloramphenicol(1000x).
- 4 colonies were picked.
- These tubes were transferred in a cell culture shaker at 37 °C and were incubated overnight
Analytical digestion and gelelectrophoresis of RFP-generator (RFC25, pSB1C3, P4 & P5)
Investigator: Jeff, Leonie, Rosario, Florian
Aim of the experiment: Analytical digestion and gelelectrophoresis of RFP-generator (RFC25, pSB1C3, P4 & P5).
Procedure:
- Batch for analytical digestion for P4 with NgoMIV+AgeI-HF
| volume | reagent |
| 2.5 µl | Plasmid DNA P4 |
| 2 µl | NEBuffer 4 (10x) |
| 0.25 µl | NgoMIV (10 U/µl) |
| 0.25 µl | AgeI-HF (20 U/µl) |
| 15 µl | ddH2O |
| =20 µl | TOTAL |
- Batch for analytical digestion for P5 with NgoMIV+AgeI-HF
| volume | reagent |
| 2.5 µl | Plasmid DNA P5 |
| 2 µl | NEBuffer 4 (10x) |
| 0.25 µl | NgoMIV (10 U/µl) |
| 0.25 µl | AgeI-HF (20 U/µl) |
| 15 µl | ddH2O |
| =20 µl | TOTAL |
- Incubation for 90 min at 37 °C.
- Analytical gelelectrophoresis was performed at 90 V for 60 min.
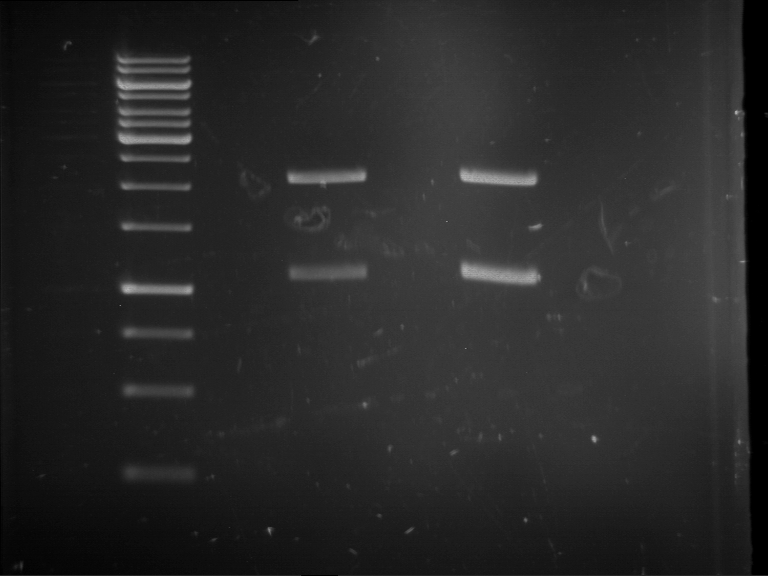
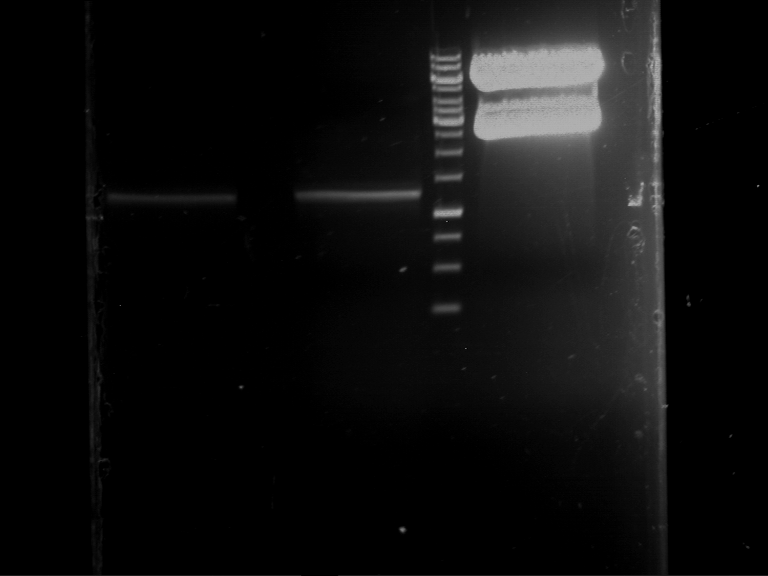
Results:
| 1 kbp ladder DNA ladder | P4 | P5 |
| Mutation successful | Mutation successful! |
- Parts are compliant and do not contain RFC25 forbidden restriction sites.
Sequencing of pTUM vectors with pGAL, pADH, pTEF1, pTEF2
Investigator: Jeff, Leonie, Rosario, Florian
Aim of the experiment: Sequencing of pTUM vectors with pGAL, pADH, pTEF1, pTEF2
Procedure:
Sequencing batch were prepared after manufacturer's protocol. (15 µl of plasmid DNA (50 - 100 ng) and 2 µl sequencing primer).
The different vectors we sequenced received the following barcodes:
- ADH in pTUM100: FR01002265
- TEF1 in pTUM100: FR01002266
- TEF2 in pTUM100: FR01002266
- GAL in pTUM100: FR01002268
Sequencing of TEF2 in pTUM100 was not interpretable. The other sequences were consistent with the sequences in the parts registry.
Wednesday, April 24th
Miniprep of Phytochrome B (2-908 N-terminal amino acids) (BBa_K801031, RFC25, pSB1C3)
Investigator: Jeff, Leonie, Florian
Aim of the experiment: Miniprep of Phytochrome B (2-908 N-terminal amino acids) (BBa_K801031, RFC25, pSB1C3).
Procedure:
- Miniprep was performed after manufacturer's protocol (QIAprep Miniprep, QIAGEN)
Analytical digestion and gelelectrophoresis of Phytochrome B (2-908 N-terminal amino acids) (BBa_K801031, RFC25, pSB1C3), P7 - P10
Investigator: Jeff, Leonie, Florian
Aim of the experiment: Analytical digestion and gelelectrophoresis of Phytochrome B (2-908 N-terminal amino acids) (BBa_K801031, RFC25, pSB1C3), P7 - P10.
Procedure:
- Batch for analytical digestion for P7 with NgoMIV+AgeI-HF
| volume | reagent |
| 2.5 µl | Plasmid DNA P7 |
| 2 µl | NEBuffer 4 (10x) |
| 0.25 µl | NgoMIV (10 U/µl) |
| 0.25 µl | AgeI-HF (20 U/µl) |
| 15 µl | ddH2O |
| =20 µl | TOTAL |
- Batch for analytical digestion for P8 with NgoMIV+AgeI-HF
| volume | reagent |
| 2.5 µl | Plasmid DNA P8 |
| 2 µl | NEBuffer 4 (10x) |
| 0.25 µl | NgoMIV (10 U/µl) |
| 0.25 µl | AgeI-HF (20 U/µl) |
| 15 µl | ddH2O |
| =20 µl | TOTAL |
- Batch for analytical digestion for P9 with NgoMIV+AgeI-HF
| volume | reagent |
| 2.5 µl | Plasmid DNA P9 |
| 2 µl | NEBuffer 4 (10x) |
| 0.25 µl | NgoMIV (10 U/µl) |
| 0.25 µl | AgeI-HF (20 U/µl) |
| 15 µl | ddH2O |
| =20 µl | TOTAL |
- Batch for analytical digestion for P10 with NgoMIV+AgeI-HF
| volume | reagent |
| 2.5 µl | Plasmid DNA P10 |
| 2 µl | NEBuffer 4 (10x) |
| 0.25 µl | NgoMIV (10 U/µl) |
| 0.25 µl | AgeI-HF (20 U/µl) |
| 15 µl | ddH2O |
| =20 µl | TOTAL |
- Incubation for 90 min at 37 °C.
- Analytical gelelectrophoresis was performed at 90 V for 60 min.
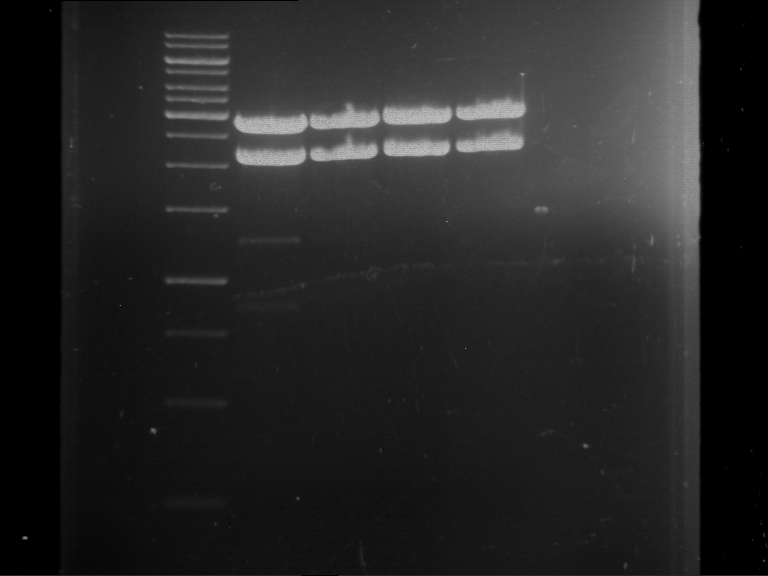
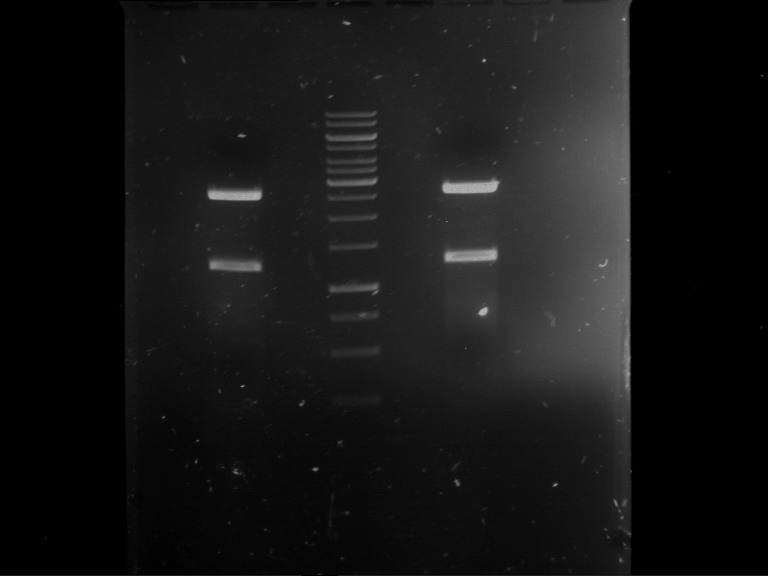
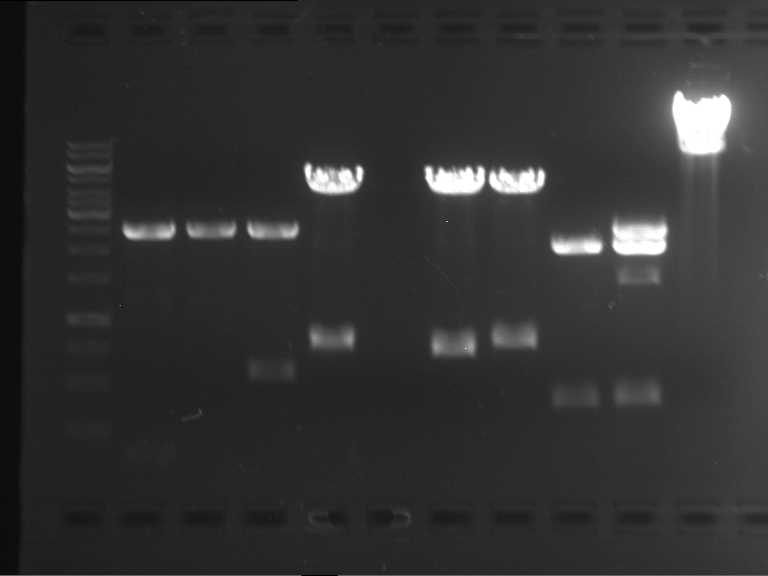
Results:
| 1 kbp ladder DNA ladder | P7 | P8 | P9 | P10 |
| Part is correct | Part is correct | Part is correct | Part is correct |
Transformation of E. coli XL1 blue with EreA and EreB (pDEST14)
Investigator: Jeff, Leonie, Florian
Aim of the experiment: Transformation of E. coli XL1 blue with EreA and EreB (pDEST14).
Procedure:
- Plasmid DNA was received dried in paper from McMaster University.
- DNA was resuspended in ddH2O
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 2 µl of DNA was added to 100 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- The transformated cells were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated on chlorampenicol and ampicillin plates.
Week 2
Monday, April 29nd
Miniprep of pRK792, pRK793, pRIL, ereA, ereB
Investigator: Leonie
Aim of the experiment: Miniprep of pRK792, pRK793, pRIL, ereA, ereB
Procedure:
- Miniprep was performed after manufacturer's protocol (QIAprep Miniprep, QIAGEN)
- Concentrations were determined by measuring the absorption:
| Plasmid | c [ng/µl] |
| pRK792 | 214,5 |
| pRK793 | 154,3 |
| pRIL | 6,1 |
| ereA | 183,4 |
| ereB | 98,3 |
- The procedure will have to be repeated for pRIL
Midiprep of RFC25 compatible RFP generator in (pSB1C3)
Investigator: Andreas, Florian
Aim of the experiment: Midiprep of RFC25 compatible RFP generator in (pSB1C3)
Procedure:
- Midiprep was performed after manufacturer's protocol (QIAprep Midiprep, QIAGEN)
Tuesday, April 30th
Preparative digestion and gelelectrophoresis of mINF1 and Leptin with HindIII and EheI
Investigator: Louise, Johanna
Aim of the experiment: Testing the enzymes, since the earlier digestion with probably older enzymes did not work (no bands on gel) .
Procedure:
- Batch for preparative digestion for mINF1 with HindIII and EheI
| volume | reagent |
| 10 µl | Plasmid DNA mINF11 |
| 5 µl | Tango Buffer (10x) |
| 2 µl | HindIII (10 U/µl) |
| 3 µl | EheI (10 U/µl) |
| 30 µl | ddH2O |
| =50 µl | TOTAL |
- Batch for preparative digestion for Leptin with HindIII and EheI
| volume | reagent |
| 20 µl | Plasmid DNA Leptin |
| 5 µl | Tango Buffer (10x) |
| 2 µl | HindIII (10 U/µl) |
| 3 µl | EheI (10 U/µl) |
| 20 µl | ddH2O |
| =50 µl | TOTAL |
- Incubation for 3 h at 37 °C.
Transformation of E. coli XL1 blue with PSH21 (P17)
Investigator: Johanna, Louise
Aim of the experiment: Transformation of E. coli XL1 blue with PSH21.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice for 10 minutes.
- 5 µl of DNA was added to 250 µl of competent cells and gently mixed.
- 60 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding the DNA to 2 ml LB-medium.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- The transformated cells were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated on chlorampenicol and ampicillin plates.
Thursday, May 2nd
Miniprep of pSH21 (pAutoRex8) containing AlcR/AlcA promotor system
Investigator: Katrin
Aim of the experiment: Miniprep of pSH21 (pAutoRex8) containing AlcR/AlcA promotor system
Procedure:
- Miniprep was performed after manufacturer's protocol (QIAprep Miniprep, QIAGEN)
- The concentration was measured (900 ng/µl)
Friday, May 3rd
Transformation of E. coli XL1 blue with pSB1C3-GFP, pSB1C3-mkate2, pSB1C3-mCherry
Investigator: Andreas
Aim of the experiment: Transformation of E. coli XL1 blue with pSB1C3-GFP, pSB1C3-mkate2, pSB1C3-mCherry.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice for 10 minutes.
- 5 µl of DNA was added to 250 µl of competent cells and gently mixed.
- 60 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding the DNA to 2 ml LB-medium.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- The transformated cells were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated on chlorampenicol and ampicillin plates.
Plating of received E. coli containing biobricks
Investigator: Jeff, Andreas
Aim of the experiment: The received biobricks were already transformed in E. coli and were in an agar stabs. These E. coli cells were transferred with an inoculation loop on antibiotic selection plates and were incubated over night.
Operational sequence:
- Bacterias containing plasmids with biobricks were transferred with a sterile inoculation loop on antibiotic plates and were incubated at 37 °C overnight.
- The biobricks were:
| Name | Function |
|---|---|
| [[http://partsregistry.org/Part:BBa_K157004 BBa_K157004]] | Fluoresceine A -binding derivative of a Lipocalin |
| [[http://partsregistry.org/Part:BBa_K863000 BBa_K863000]] | bpul (laccase from Bacillus pumilus) with T7 promoter, RBS and HIS tag |
| [[http://partsregistry.org/Part:BBa_K909009 BBa_K909009]] | cDNA of UV-B sensing protein UVR8 from Arabidopsis thaliana |
| [[http://partsregistry.org/Part:BBa_K337013 BBa_K337013]] | constitutive promoter SV40 |
| [[http://partsregistry.org/Part:BBa_K747096 BBa_K747096]] | constitutive promoter CMV |
| [[http://partsregistry.org/Part:BBa_K414002 BBa_K414002]] | constitutive promoter CaMV35S |
| [[http://partsregistry.org/Part:BBa_K147002 BBa_K147002]] | xylE gene coding for catechol-1,2-dioxygenase |
| [[http://partsregistry.org/Part:BBa_E2020 BBa_E2020]] | Cerulean |
| [[http://partsregistry.org/Part:BBa_K404319 BBa_K404319]] | mCFP |
| [[http://partsregistry.org/Part:BBa_E0040 BBa_E0040]] | GFPmut3b |
| [[http://partsregistry.org/Part:BBa_K592010 BBa_K592010]] | amilGFP |
| [[http://partsregistry.org/Part:BBa_E2050 BBa_E2050]] | mOrange |
| [[http://partsregistry.org/Part:BBa_K399001 BBa_K399001]] | mStrawberry |
| [[http://partsregistry.org/Part:BBa_E1010 BBa_E1010]] | mRFP1 |
| [[http://partsregistry.org/Part:BBa_E2060 BBa_E2060]] | mCherry |
| [[http://partsregistry.org/Part:BBa_K592012 BBa_K592012]] | eforRed |
| [[http://partsregistry.org/Part:BBa_K592011 BBa_K592011]] | cjBlue |
| [[http://partsregistry.org/Part:BBa_K864401 BBa_K864401]] | aeBlue |
| [[http://partsregistry.org/Part:BBa_K592009 BBa_K592009]] | amilCP, blue chromoprotein |
| [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K414000 BBa_K414000]] | RD29A Promoter + Strong Plant Kozak (RBS) + RFP |
| [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K414006 BBa_K414006]] | 35S Promoter + (Strong Plant RBS + GFP) From K414001 |
| [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K414007 BBa_K414007]] | DREB1C promoter |
| [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K678001 BBa_K678001]] | alcA promoter |
| [[http://partsregistry.org/Part:BBa_K243004 BBa_K243004]] | Short linker |
| [[http://partsregistry.org/Part:BBa_K243005 BBa_K243005]] | Middle linker |
| [[http://partsregistry.org/Part:BBa_K243006 BBa_K243006]] | Long Linker |
| [[http://partsregistry.org/Part:BBa_K243029 BBa_K243029]] | GSAT Linker |
| [[http://partsregistry.org/Part:BBa_K243030 BBa_K243030]] | SEG Linker |
Sunday, May 5th
Picking of the plated E. coli containing biobricks and the parts received from LMU
Investigator: Florian
Aim of the experiment: The colonies from the plates were transferred into liquid LB medium, so that minipreps can be performed the next day.
Operational sequence:
- Tubes with 5 ml LB medium and the corresponding antibiotic were prepared.
- A single colony from each plate was transferred into a tube with a pipet tip.
- The biobricks were:
| Label | Name | Function |
|---|---|---|
| p19 | mCherry in pSB1C3 | |
| p20 | [[http://partsregistry.org/Part:BBa_K823039 BBa_K823039]] | GFP in pSB1C3 |
| p21 | [[http://partsregistry.org/Part:BBa_K823029 BBa_K823029]] | mKate2 in pSB1C3 |
| p22 | [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K414006 BBa_K414006]] | 35S Promoter + (Strong Plant RBS + GFP) From K414001 |
| p23 | [[http://partsregistry.org/Part:BBa_K414002 BBa_K414002]] | constitutive promoter CaMV35S |
| p24 | [[http://partsregistry.org/Part:BBa_K909009 BBa_K909009]] | cDNA of UV-B sensing protein UVR8 from Arabidopsis thaliana |
| p25 | [[http://partsregistry.org/Part:BBa_K399001 BBa_K399001]] | mStrawberry |
| p26 | [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K414007 BBa_K414007]] | DREB1C promoter |
| p27 | [[http://partsregistry.org/Part:BBa_K863000 BBa_K863000]] | bpul (laccase from Bacillus pumilus) with T7 promoter, RBS and HIS tag |
| p28 | [[http://partsregistry.org/Part:BBa_K337013 BBa_K337013]] | constitutive promoter SV40 |
| p29 | [[http://partsregistry.org/Part:BBa_K592011 BBa_K592011]] | cjBlue |
| p30 | [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K678001 BBa_K678001]] | alcA promoter |
| p31 | [[http://partsregistry.org/Part:BBa_K592012 BBa_K592012]] | eforRed |
| p32 | [[http://partsregistry.org/Part:BBa_K592010 BBa_K592010]] | amilGFP |
| p33 | [[http://partsregistry.org/Part:BBa_K864401 BBa_K864401]] | aeBlue |
| p34 | [[http://partsregistry.org/Part:BBa_K404319 BBa_K404319]] | mCFP |
| p35 | [[http://partsregistry.org/wiki/index.php?title=Part:BBa_K414000 BBa_K414000]] | RD29A Promoter + Strong Plant Kozak (RBS) + RFP |
| p36 | [[http://partsregistry.org/Part:BBa_K592009 BBa_K592009]] | amilCP, blue chromoprotein |
| p37 | [[http://partsregistry.org/Part:BBa_K747096 BBa_K747096]] | constitutive promoter CMV |
| p38 | [[http://partsregistry.org/Part:BBa_K147002 BBa_K147002]] | XylE dioxygenase (catechol dioxygenase) |
| p39 | [[http://partsregistry.org/Part:BBa_K243005 BBa_K243005]] | Middle linker |
| p40 | [[http://partsregistry.org/Part:BBa_K243029 BBa_K243029]] | GSAT Linker |
| p41 | [[http://partsregistry.org/Part:BBa_E0040 BBa_E0040]] | GFPmut3b |
| p42 | [[http://partsregistry.org/Part:BBa_K243006 BBa_K243006]] | Long Linker |
| p43 | [[http://partsregistry.org/Part:BBa_K243030 BBa_K243030]] | SEG Linker |
| p44 | [[http://partsregistry.org/Part:BBa_K243004 BBa_K243004]] | Short linker |
| p45 | [[http://partsregistry.org/Part:BBa_K157004 BBa_K157004]] | Fluoresceine A -binding derivative of a Lipocalin |
| p46 | [[http://partsregistry.org/Part:BBa_E2060 BBa_E2060]] | mCherry |
| p47 | [[http://partsregistry.org/Part:BBa_E2020 BBa_E2020]] | Cerulean |
| p48 | [[http://partsregistry.org/Part:BBa_E1010 BBa_E1010]] | mRFP1 |
| p49 | [[http://partsregistry.org/Part:BBa_E2050 BBa_E2050]] | mOrange |
- There were no colonies on the plate with K864401 (p33), therefore the LB medium could not be infected
- The tubes were placed into the incubator at 37 °C
Week 3
Monday, May 6th
Miniprep of the prepared E. coli containing biobricks and the parts from LMU
Investigators: Florian, Johanna, Jefferey
Aim of the experiment: A miniprep of the overnight culture of our E. coli containig our biobricks and LMU parts was performed using the Qiagen miniprep kit.
Procedure:
- Miniprep was performed after manufacturer's protocol (QIAprep Miniprep, QIAGEN)
- Resulting concentrations were determined by measuring the absorption:
| Label | Name | Concentration [ng/µl] |
|---|---|---|
| p19 | mCherry in pSB1C3 | 40,2 |
| p20 | GFP in pSB1C3 | 58,5 |
| p21 | mKate2 in pSB1C3 | 131,3 |
| p22 | 35S Promoter + (Strong Plant RBS + GFP) From K414001 | 42,8 |
| p23 | constitutive promoter CaMV35S | 39,3 |
| p24 | cDNA of UV-B sensing protein UVR8 from Arabidopsis thaliana | 30,0 |
| p25 | mStrawberry | 46,6 |
| p26 | DREB1C promoter | 34,8 |
| p27 | bpul (laccase from Bacillus pumilus) with T7 promoter, RBS and HIS tag | 52,5 |
| p28 | constitutive promoter SV40 | 58,7 |
| p29 | cjBlue | 44,4 |
| p30 | alcA promoter | 26,9 |
| p31 | eforRed | 64,8 |
| p32 | amilGFP | 59,3 |
| p34 | mCFP | 78,4 |
| p35 | RD29A Promoter + Strong Plant Kozak (RBS) + RFP | 61,2 |
| p36 | amilCP, blue chromoprotein | 89,2 |
| p37 | constitutive promoter CMV | 83,6 |
| p38 | XylE dioxygenase (catechol dioxygenase) | 57,4 |
| p39 | Middle linker | 64,3 |
| p40 | GSAT Linker | 109,6 |
| p41 | GFPmut3b | 118,1 |
| p42 | Long Linker | 133,0 |
| p43 | SEG Linker | 107,5 |
| p44 | Short linker | 60,2 |
| p45 | Fluoresceine A -binding derivative of a Lipocalin | 82,7 |
| p46 | mCherry | 230,2 |
| p47 | Cerulean | 25,7 |
| p48 | mRFP1 | 299,3 |
| p49 | mOrange | 181,2 |
Sequencing of FluA, SV40, CaMV35S, xylE, LMU mKate2, LMU GFP, LMU mCherry, DREB1C, RD29A
Investigators: Andreas, Florian, Johanna, Jefferey, Rosario
Aim of the experiment: Sequencing of genes for which no prior sequencing results were available.
Procedure: The DNA was prepared for sequencing according to the Eurofins SmartSeq Kit protocol (15 µl of plasmid DNA (50 - 100 ng) and 2 µl sequencing primer).
The different genes we sequenced received the following barcodes:
| Label | Name | Barcode |
|---|---|---|
| p45 | FluA | FR01002355 |
| p28 | SV40 | FR01002356 |
| p23 | CaMV35S | FR01002357 |
| p38 | xylE | FR01002358 |
| p21 | LMU mKate2 | FR01002359 |
| p20 | LMU GFP | FR01002360 |
| p19 | LMU mCherry | FR01002345 |
| p46 | mCherry | FR01002346 |
| p26 | DREB1C | FR01002347 |
| p35 | RD29A | FR01002348 |
Preparation of ordered primers
Investigators: Andreas, Jefferey, Rosario
Aim of the experiment: Dissolving of primer pallets to provide a concentration of 100 pmol/µl
Procedure: We added an amount of water which gave us a final primer concentration of 100 pmol/µl.
Primers were labeled as follows:
| Label | Oligonr. |
|---|---|
| 6 | EreA_for |
| 7 | EreA_rev |
| 8 | EreA_SDM1_for |
| 9 | EreA_SDM1_rev |
| 10 | EreA_SDM2_for |
| 11 | EreA_SDM2_rev |
| 12 | EreA_SDM3_for |
| 13 | EreA_SDM3_rev |
| 14 | EreB_for |
| 15 | EreB_rev |
| 16 | EreB_SDM1_for |
| 17 | EreB_SDM1_rev |
| 18 | EreB_SDM2_for |
| 19 | EreB_SDM2_rev |
| 20 | EreB_SDM3_for |
| 21 | EreB_SDM3_rev |
Tuesday, May 7th
PCR, purification and analytical geleletrophoresis of EreA (P15) and EreB (P16)
Investigator: Jeff, Louise, Florian, Rosario, Andi, Johanna
Aim of the experiment: PCR of EreA (P15) and EreB (P16).
Procedure:
Operational sequence:
- PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer (P15: O46 (EreA_for); P16: O31 (EreB_for)) |
| 1 µl | 10 µM Reverse Primer (P15: O47 (EreA_rev); P16: O32 (EreB_rev)) |
| 0.25 µL | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µL) |
| 1 µl | Plasmid DNA (P15; P16) |
| 35.75 µL | ddH2O Water |
| =50 µL | TOTAL |
- Mix with pipette
- The gradient PCR program was performed after following scheme with following conditions (Tm=54 °C; ΔG=2 °C; P15 in row 11(=52 °C); P16 in row 1 (=54.0 °C)):
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| Tm=54 °C; ΔG=2 °C | 60 s | |
| 68 °C | 80 s | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | infinite |
- After PCR, the product was purified with QIAquick PCR Purification Kit, Qiagen.
- 9 µl of the purified PCR products were mixed with 1 µl DNA loading buffer (10x) for analytical gelelectrophoresis
- Analytical gelelectrophoresis was performed at 90 V for 60 min in 1% agarose gel.
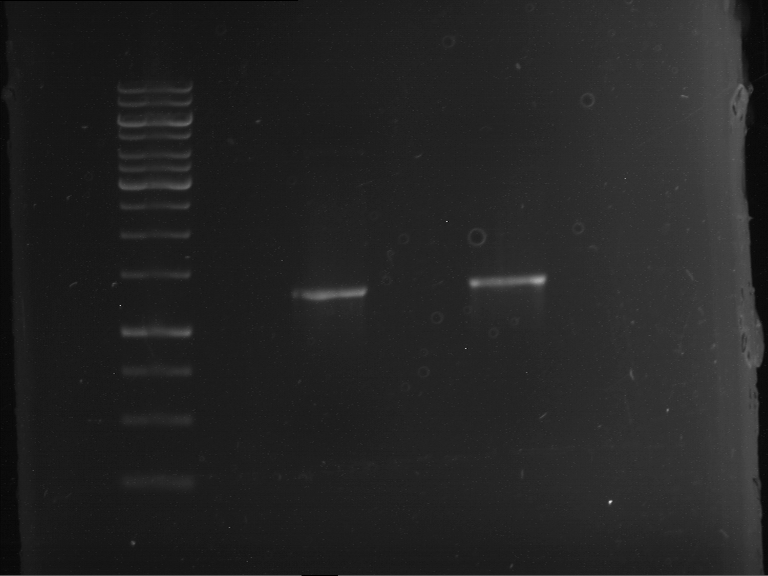
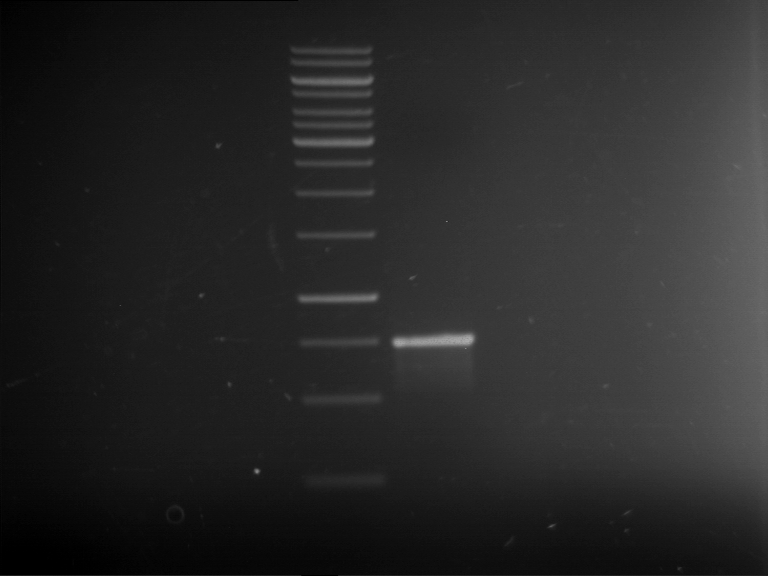
| 1 kbp ladder | F1 | F2 |
| Fragment-size like expected | Fragment-size like expected |
Preparative digestion and gelelectrophoresis of PhyB (P9) and PIF3-20aa linker (P50) with AgeI and PstI as well as 20aalinker-PIF3 (P51) with EcoRI and NgoMIV
Investigator: Jeff, Andi, Florian, Rosario, Louise, Johanna
Aim of the experiment: Preparative digestion and gelelectrophoresis of PhyB (P9) and PIF3-20aa linker (P50) with AgeI and PstI as well as 20aalinker-PIF3 (P51) with EcoRI and NgoMIV .
Procedure:
- Batch for preparative digestion of PhyB (P9) and PIF3-20aa linker (P50) with AgeI and PstI
| volume | reagent |
| 20 µl | Plasmid DNA P9/P50 |
| 4 µl | NEBuffer 4 |
| 0.4 µl | BSA (100x) |
| 1 µl | AgeI-HF |
| 1 µl | PstI-HF |
| 13.6 µl | ddH2O |
| =40 µl | TOTAL |
- Batch for preparative digestion of 20aalinker-PIF3 (P51) with EcoRI and NgoMIV
| volume | reagent |
| 20 µl | Plasmid DNA P51 |
| 4 µl | NEBuffer 4 |
| 1 µl | EcoRI-HF |
| 1 µl | NgoMIV |
| 14 µl | ddH2O |
| =40 µl | TOTAL |
- Incubation for 3 h at 37 °C.
- 4 µl of DNA loading buffer (10x) were added to the reaction batches after digestion and were loaded on an 1% low-melting agarose gel for preparative gelelectrophoresis.
- Preparative gelelectrophoresis was performed at 70 V for 90 min.
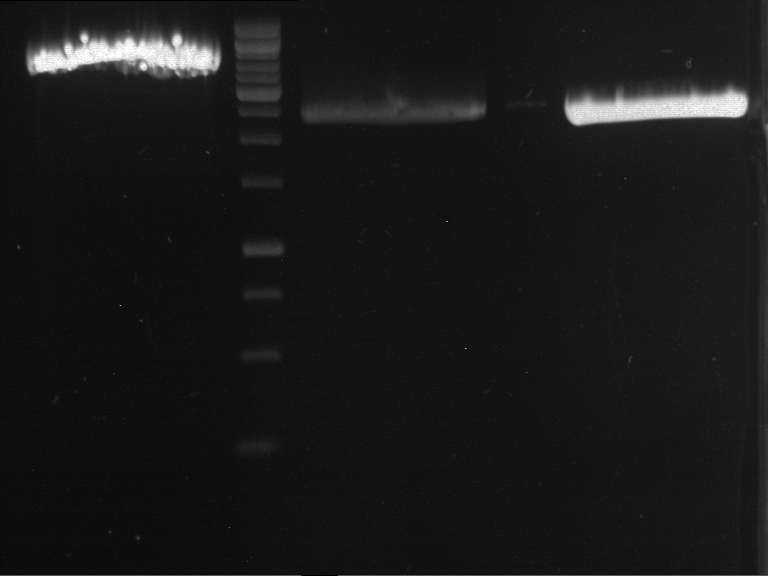
| P9 digestion = F3 | 1 kbp ladder | P50 digestion = F4 | P51 digestion = F5 |
| Fragment-size like expected; band was cut out | Fragment-size like expected; band was cut out | Fragment-size like expected; band was cut out |
- Bands were extracted by QIAquick Gel Extraction Kit, QIAGEN.
Wednesday, May 8th
Preparative digestion and gelelectrophoresis of PCR product of EreA (F1) and EreB (F2) with XbaI & AgeI
Investigator: Jeff, Andi, Florian, Rosario, Louise
Aim of the experiment: Preparative digestion and gelelectrophoresis of PCR product of EreA (F1) and EreB (F2) with XbaI & AgeI.
Procedure:
- Batch for preparative digestion of PCR product of EreA (F1) and EreB (F2) with XbaI & AgeI.
| volume | reagent |
| 25 µl | PCR product F1/F2 |
| 5 µl | NEBuffer 4 |
| 0.5 µl | BSA (100x) |
| 1 µl | AgeI-HF |
| 1 µl | XbaI |
| 17.5 µl | ddH2O |
| =50 µl | TOTAL |
- Batch for preparative digestion of P52 for preperation of pSB1C3 vector:
| volume | reagent |
| 20 µl | Plasmid DNA P52 |
| 4 µl | NEBuffer 4 |
| 0.4 µl | BSA (100x) |
| 1 µl | AgeI-HF |
| 1 µl | XbaI |
| 13.6 µl | ddH2O |
| =40 µl | TOTAL |
- Incubation for 3 h at 37 °C.
- 5 µl of DNA loading buffer (10x) were added to the reaction batches for F1 & F2 and 4 µl of DNA loading buffer (10x) were added to the reaction batches for Pxx after digestion and were loaded on an 1% agarose gel for preparative gelelectrophoresis.
- Preparative gelelectrophoresis was performed at 90 V for 60 min.
| F1 digestion = F6 | F2 digestion = F7 | 1 kbp ladder | P52 digestion = F8 |
| Length was like expected | Length was like expected | Length was like expected; the lower band was extracted |
- Bands were extracted by QIAquick Gel Extraction Kit, QIAGEN.
Ligation of F8+F6 (EreA in pSB1C3), F8+F7 (EreB in pSB1C3)
Investigator: Jeff, Andi, Florian, Rosario, Louise
Aim of the experiment: Ligation of F8+F6 (EreA in pSB1C3), F8+F7 (EreB in pSB1C3). Quickchanges have still to be performed.
Procedure:
Ligation batch for F8+F6 (EreA in pSB1C3), F8+F7 (EreB in pSB1C3):
- Ligation batch for F8+F6
| volume | reagent |
| 1.35 µl | F8 (74.3 ng/µl, 2086 bp) |
| 6.76 µl | F6 (25.9 ng/µl, 1218 bp) |
| 2 µl | T4 ligase buffer (10x) |
| 1 µl | T4 ligase (1 U/µl) |
| 8.89 µl | ddH2O |
| =20 µl | TOTAL |
- Ligation batch for F8+F7
| volume | reagent |
| 1.35 µl | F8 (74.3 ng/µl, 2086 bp) |
| 8.78 µl | F7 (20.6 ng/µl, 1257 bp) |
| 2 µl | T4 ligase buffer (10x) |
| 1 µl | T4 ligase (1 U/µl) |
| 6.87 µl | ddH2O |
| =20 µl | TOTAL |
- Negative control was als prepared. Instead of the insert, the same volume of ddH2O was added to the reaction batches.
- Cycled ligation has been performed after following protocol in a thermocycler:
| 100 cycles | 12 °C | 60 s |
| 22 °C | 60 s | |
| 12 °C | 60 s | |
| 22 °C | 60 s | |
| 12 °C | 60 s | |
| 22 °C | 60 s | |
| 12 °C | 60 s | |
| 22 °C | 60 s | |
| Hold | 16 °C | infinite |
Lid temperature = 37 °C
Midiprep of cloning vector RFC25 RFP generating device
Investigators: Andreas, Rosario
Aim of the experiment: Midiprep of our cloning vector RFC25 RFP generating device.
Procedure: Midiprep was performed after manufacturer's protocol (QIAprep Midiprep, QIAGEN)
The yield of our vector was 1854 ng.
Thursday, May 9th
Transformation of E. coli XL1 blue with F8(pSB1C3)+F6(EreA) and F8(pSB1C3)+F7(EreB)
Investigator: Florian, Jeff, Leonie, Louise, Katrin, Ingmar
Aim of the experiment: Transformation of E. coli XL1 blue with F8(pSB1C3)+F6(EreA) and F8(pSB1C3)+F7(EreB). Quickchanges has now to be performed to remove forbidden restriction sites.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 10 µl of ligation products F8+F6, F8+F7, F8 negative control were added to 100 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Friday, May 10th
Quick Change mutagenesis to remove mutations found by sequencing in our gene constructs
Investigator: Andi, Jeff, Rosario
Aim of the experiment: Deletion of found mutations.
Operational sequence
1. PCR general setup
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | DNA template |
| 1 µl | 1:10 dilution of appropriate Oligo (= 5 pmol) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | DNA template |
| 1 µl | 1:10 dilution of appropriate Oligo (= 5 pmol) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 12 | 95 °C | 30 s |
| 55 °C | 1 min | ||
| 68 °C | 6 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were one time more applied.
- After the PCR was finished 1 µl of the DpnI restriction enzyme (10 U/µl)was added
- Now the reaction was mixed gently and thoroughly, spinned down in a microcentrifuge for 1 minute, and immediately incubated at 37 °C for 1 hour to digest the parental supercoiled dsDNA
2. Used constructs and primers
| Quickchange number | Plasmid number | Geneconstruct | Oligo number |
| QC 1 | P27 | BPul_Laccase+ pSB1C3 | O26 & O27 |
Picking of transformed Plasmids F8 + F6
Investigator: Andi
Aim of the study: Picking from the overnight transformation of the ligated F8+F6, to see whether ligation is successful or not.
Operational sequence:
- Picking and overnight culture after standard laboratory's protocol. (CamR LB-medium)
Investigator: Andi
PCR of TEV (P14, pRK793)
Investigator: Katrin, Jeff
Aim of the experiment: PCR, purification and analytical geleletrophoresis of TEV (P14, pRK793) to produce TEV with RFC25 pre- and suffx (still contains forbidden SpeI-site).
Procedure:
Operational sequence:
- PCR reaction mixture
| volume | reagent |
| 10 µl | 5x OneTaq Standard Reaction Buffer |
| 1 µl | 10 mM dNTPs |
| 1 µl | 10 µM Forward Primer (O34 (TEV_fw)) |
| 1 µl | 10 µM Reverse Primer (O35 (TEV_rv)) |
| 0.25 µL | OneTaq Hot Start DNA Polymerase (Finally: 1.25 units/50 µL) |
| 1 µl | Plasmid DNA (P14) |
| 35.75 µL | ddH2O Water |
| =50 µL | TOTAL |
- Mix with pipette
- The PCR program was performed after following scheme with following conditions:
| Initial denaturation | 94 °C | 30 s |
| 30 cycles | 94 °C | 30 s |
| 55 °C | 60 s | |
| 68 °C | 150 s | |
| Final extension | 68 °C | 5 min |
| Hold | 4 °C | infinite |
- After PCR, the product was purified with QIAquick PCR Purification Kit, Qiagen.
- Resulting product was labeled as F12.
Digestion of P61 with S1 Nuclease
Investigator: Katrin
Oligohybridization of O22 (MinMCS_for) and O23 (MinMCS_rev)
Investigator: Leonie
Preparation of ordered primers
Investigators: Andreas, Rosario, Katrin, Ingmar
Aim of the experiment: Dissolving of primer pallets to provide a concentration of 100 pmol/µl
Procedure: We added an amount of water which gave us a final primer concentration of 100 pmol/µl.
Primers were labeled as follows:
| Label | Oligonr. |
|---|---|
| 24 | BPU_for |
| 25 | BPU_rev |
| 26 | BPU_SDM1_for |
| 27 | BPU_SDM1_rev |
| 28 | BPU_SDM2_for |
| 29 | BPU_SDM2_rev |
| 30 | BPU_SDM3_for |
| 31 | BPU_SDM3_rev |
| 32 | BPU_SDM4_for |
| 33 | BPU_SDM4_rev |
| 34 | TEV_for |
| 35 | TEV_rev |
| 36 | TEV_t387c_for |
| 37 | TEV_t387c_rev |
| 38 | N_Split_TEV_for |
| 39 | N_Split_TEV_rev |
| 40 | C_Split_TEV_for |
| 41 | C_Split_TEV_rev |
| 42 | PIF6_2-100_for |
| 43 | PIF6_2-100_rev |
| 44 | p104AgeI a71g fw |
| 45 | p104AgeI a71g rev |
| 46 | xylE_for |
| 47 | xylE_rev |
| 48 | xylE_c312a_for |
| 49 | xylE_c312a_rev |
| 50 | xylE_g483a_for |
| 51 | xylE_g483a_rev |
| 52 | xylE_a834g_for |
| 53 | xylE_a834g_rev |
Saturday, May 11th
Preparative digestion of PCR product of S219V pRK793 (F12) with XbaI & AgeI
Investigator: Louise
Aim of the experiment: Preparative digestion of PCR product of S219V pRK793 (F12) with XbaI & AgeI. Quickchange has still to be performed (forbidden SpeI restriction site).
Procedure:
- Batch for preparative digestion of PCR product S219V pRK793 (F12) with XbaI & AgeI.
| volume | reagent |
| 20 µl | PCR product F12 |
| 4 µl | NEBuffer 4 |
| 0.4 µl | BSA (100x) |
| 1 µl | AgeI-HF |
| 1 µl | XbaI |
| 13.6 µl | ddH2O |
| =40 µl | TOTAL |
- Incubation for 3 h at 37 °C
- Digested products were purified to remove nucleotides and enzymes with QIAquick PCR Purification Kit, QIAGEN.
- The digested fragment was labelled F15
Miniprep of overnight culture of transformated E.~coli XL1-Blue with F8 (pSB1C3) + F6 (EreA)
Investigator: Louise
Aim of the experiment: Miniprep of overnight culture of transformated E.~coli XL1-Blue with F8 (pSB1C3) + F6 (EreA) of 2 clones picked the day before
Procedure:
- Miniprep was performed after manufacturer's protocol (QIAprep Miniprep, QIAGEN)
- The plasmids were labelled as P63 and P64
Analytical digestion and gelelectrophoresis of P63 and P64 with AgeI-HF and XbaI
Investigator: Louise
Aim of the experiment: Analytical digestion and gelelectrophoresis of P63 and P64 with AgeI-HF and XbaI
Procedure:
- Batch for analytical digestion of P63 and P64 with AgeI-HF and XbaI
| volume | reagent |
| 2.5 µl | Plasmid DNA P63/P64 |
| 2 µl | NEBuffer 4 (10x) |
| 0.2 µl | BSA (100x) |
| 0.25 µl | XbaI(10 U/µl) |
| 0.25 µl | AgeI-HF (20 U/µl) |
| 14.8 µl | ddH2O |
| =20 µl | TOTAL |
- Incubation for 90 min at 37 °C.
- Analytical gelelectrophoresis was performed at 90 V for 60 min.
| Analytical digestion of P63 with AgeI-HF and XbaI | 1 kbp ladder | Analytical digestion of P64 with AgeI-HF and XbaI |
| Insertion successful | Insertion successful |
Preparative digestion of PCR product of EreB (F2) with XbaI & AgeI
Investigator: Louise,Leonie,Johanna
Aim of the experiment: Preparative digestion of PCR product EreB (F2) with XbaI & AgeI (second try since there did not grow any transformed bacteria in the first time).
Procedure:
- Batch for preparative digestion of PCR product and EreB (F2) with XbaI & AgeI.
| volume | reagent |
| 20 µl | PCR product F2 |
| 5 µl | NEBuffer 4 |
| 0.5 µl | BSA (100x) |
| 1 µl | AgeI-HF |
| 1 µl | XbaI |
| 22.5 µl | ddH2O |
| =50 µl | TOTAL |
- Incubation for 3 h at 37 °C
- The digested fragment was labelled F14
Purification and ligation of digested PCR product EreB (F14)
Investigator: Louise, Rosario
Aim of the experiment: PCR Purification and ligation of digested PCR product EreB (F14).
Procedure: PCR Purification of EreB PCR (F14)
- To purify the digested Fragment(F14) the QIAquick PCR Purification Kit was used. The concentration of the purified product was found to be 5 ng/µl via Nano drop.
Ligation of F14 + F8(pSB1C3)
- Ligation batch for F8+F14 (positive control)
| volume | reagent |
| 1.35 µl | F8 (74.3 ng/µl, 2086 bp) |
| 15.65 µl | F14 (5 ng/µl, 1257 bp) |
| 2 µl | T4 ligase buffer (10x) |
| 1 µl | T4 ligase (1 U/µl) |
| =20 µl | TOTAL |
- Negative control was also prepared. Instead of the insert, the same volume of ddH2O was added to the reaction batch.
- Cycled ligation has been performed after following protocol in a thermocycler:
| 99 cycles | 12 °C | 60 s |
| 22 °C | 60 s | |
| 12 °C | 60 s | |
| 22 °C | 60 s | |
| 12 °C | 60 s | |
| 22 °C | 60 s | |
| 12 °C | 60 s | |
| 22 °C | 60 s | |
| 12 °C | 60 s | |
| 22 °C | 60 s | |
| Hold | 16 °C | infinite |
Lid temperature = 37 °C
Transformation of ligation products of F10 + F11 into E.coli Xl1-Blue
Investigator: Andi
Aim of the experiment:Transformation of the ligation products of F10 + F11 and the negative control in pTUM104 in XL1 Blue.
Operation Sequence
- melting of 2x 200 µl Ca-competent E.coli XL1-Blue cells on ice
- Preparing 4 Tubes with 100 µl of Ca-competent E.coli XL1-Blue cells
- addition of 5 µl of the ligation products
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Transformation of Quickchange Product P62 into E.coli Xl1-Blue
Investigator: Andi
Aim of the experiment:Transformation of the Quickchange Product P62 and the negative control in XL1 Blue.
Operation Sequence
- melting of 1x 200 µl Ca-competent E.coli XL1-Blue cells on ice
- Preparing 4 Tubes with 50 µl of Ca-competent E.coli XL1-Blue cells
- addition of 1 µl of the Quickchange Product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Quickchange mutagenesis of pTUM104
Investigator: Ingmar
Aim of the experiment: Sequencing results showed a forbiden AgeI restriction site in this vector. The sdm will delete this restricition site
Procedure:
PCR
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P6 template |
| 0.5 µl | 1:10 dilution of O44 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 4 µl | Plasmid P6 template |
| 0.5 µl | 1:10 dilution of O45 (10 pmol/µL) |
| 16.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 10 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 67°C | 6 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were one time more applied.
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C. The resulting product was named fragmet 14.
Transformation into E.coli Xl1-Blue Operation Sequence
- melting of 100 µl Ca-competent E.coli XL1-Blue cells on ice
- addition of 1 µl of the PCR product
- incubation for 30 min on ice
- heat shock for 5 min at 37 °C
- transfer of cells to 1 ml LB-medium without antibiotics and incubate at 37°C and 180 rpm for 30 min
- plate 100 µl on an Amp-LB-plate
- sediment the leftover in a centrifuge (30 - 60 sec, 13 000 rpm) and resuspend the sediment in 100 µl LB-medium and plate it as well on an Amp-LB-plate
Sunday, May 12th
Analytical gelelectrophoresis of F15 (digested TEV PCR product)
Investigator: Jeff, Rosario
Aim of the experiment: Analytical gelelectrophoresis of F15 (digested TEV PCR product) to check whether PCR has worked.
Procedure:
- 5 µl of F15 was mixed with 0.55 µl of DNA loading buffer (10x) and analytical gelelectrophoresis was performed at 1% agarose gel for 60 min at 90 V.
- PCR product has expected length
Transformation of E. coli XL1 blue with ligation product F8+F15 (TEV in pSB1C3, RFC25)
Investigator: Jeff, Rosario
Aim of the experiment: Transformation of E. coli XL1 blue with ligation product F8+F15 (TEV in pSB1C3, RFC25). Further quickchanges still have to be done.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 10 µl of the ligation products were added to 200 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Transformation of E. coli XL1 blue with ligation products F8+F14 (EreB in pSB1C3, RFC25)
Investigator: Jeff, Rosario
Aim of the experiment: Transformation of E. coli XL1 blue with ligation product F8+F14 (EreB in pSB1C3, RFC25). Further quickchanges still have to be done.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 10 µl of the ligation products were added to 200 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Transformation of E. coli XL1 blue with P23
Investigator: Jeff, Rosario
Aim of the experiment: Transformation of E. coli XL1 blue with P23 for insertion of miniMCS.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 2 µl of DNA was added to 100 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Picking of transformed Plasmid F10+F11 (IRES from polio virus, blunt ligated with pSB1C3 for sequencing)
Investigator: Andi
Aim of the study: Picking of transformed Plasmid F10+F11 (AlcR blunt ligated with pSB1C3 for sequencing).
Operational sequence:
- Picking and overnight culture after standard laboratory's protocol. (CamR LB-medium)
- 10 colonies were picked.
Week 4
Monday, May 13th
Miniprep of overnight culture of transformated E.~coli XL1-Blue with F10+F11
Investigator: Rosario, Jeffery, Johanna, Flo, Leonie
Aim of the experiment: Miniprep of overnight culture of transformated E.~coli XL1-Blue F10+F11 of 10 clones picked the day before
Procedure:
- Miniprep was performed after manufacturer's protocol (QIAprep Miniprep, QIAGEN)
Analyzing sequencing results
Investigator: Leonie,Johanna
Aim of the experiment: Comparison of the sequencing results with the Biobrick (BB) sequences.
Procedure: Using Geneious to align the sequencing results with the BB sequences.
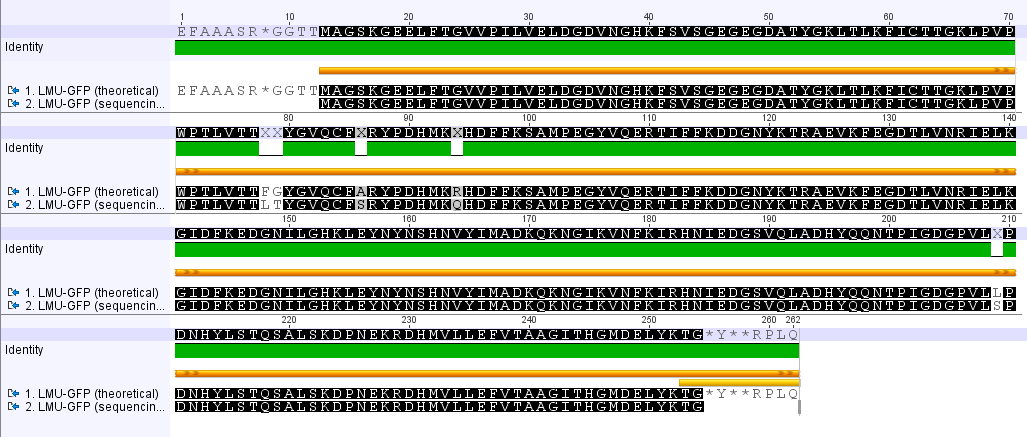
LMU GFP in RFC 25(p20)
- The provided sequence in the parts registry is totally wrong.
- The comparison of the amino acid seqeunces reveals several point mutations.
FluA in RFC 25 (p45)
- Did not yield sequencing results (does the primer bind ?)
- No presence of RCF 25 standard restriction enzymes
mCherry in RFC 25 (p46)
- Did not yield sequencing results (does the primer bind ?)
- No presence of RCF 25 standard restriction enzymes
CaMV35S in RFC 25 (p23)
- high identity
- high match with HQ699544 (Blast)
LMU mKate2in RFC 25 (p21)
- no continous ORF
- Did not yield sequencing results
DREB1C promoter in RFC 25 (p26)
- Present of fluorecent protein gene
xylE in RCF 25 (p38)
- Did not yield sequencing results
SV40 in RCF 25 (p28)
- sequencing result agrees with the laccases
- the sequencing result can't be the one of SV40
Quickchange mutagenesis SDMI of p27
Investigator: Rosario, Jeff, Flo, Andi
Aim of the experiment: Sequencing results showed a forbidden restriction site in this vector. The sdm will delete this restricition site. The resulting Plasmid was labeled 65.
Procedure:
PCR
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 0.5 µl | Plasmid P27 template |
| 0.5 µl | 1:10 dilution of O26 (10 pmol/µL) |
| 20. 5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 0.5 µl | Plasmid P27 template |
| 0.5 µl | 1:10 dilution of O27 (10 pmol/µL) |
| 20.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 10 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 67°C | 4 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were one time more applied, but the amount of cycles was increased to 20
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Quickchange mutagenesis SDMI of p38
Investigator: Jeff, Flo, Andi, Rosario
Aim of the experiment: Sequencing results showed a forbidden restriction site in this vector. The sdm will delete this restricition site. The resulting Plasmid was labeled 67.
Procedure:
PCR
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 0,5 µl | Plasmid P38 template |
| 0.5 µl | 1:10 dilution of O48 (10 pmol/µL) |
| 20.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 0.5 µl | Plasmid P38 template |
| 0.5 µl | 1:10 dilution of O49 (10 pmol/µL) |
| 20.5 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 10 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 67°C | 4 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were one time more applied, but the amount of cycles have been increased to 20
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Quickchange mutagenesis SDMI of P63
Investigator: Jeff, Rosario, Flo, Andi
Aim of the experiment: Sequencing results showed a forbidden restriction site in this vector. The sdm will delete this restricition site. The resulting Plasmid was labeled P66.
Procedure:
PCR
Reaction batch 1
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 0.25 µl | Plasmid P63 template; 1:2 dilution |
| 0.5 µl | 1:10 dilution of O8 (10 pmol/µL) |
| 20.75 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
Reaction batch 2
| volume | reagent |
| 2.5 µl | 10x Pfu Ultra II buffer |
| 0.25 µl | Plasmid P63 template; 1:2 dilution |
| 0.5 µl | 1:10 dilution of O9 (10 pmol/µL) |
| 20.75 µl | ddH2O |
| 0.5 µl | dNTP mix |
| 0.5 µl | Pfu Ultra II DNA polymerase (2.5 U / µl) |
PCR cycling parameters
| Segment | Cycles | Temperature | Time |
| 1 | 1 | 95 °C | 30 sec |
| 2 | 10 | 95°C | 30 sec |
| 55°C | 1 min | ||
| 67°C | 6 min |
- Having completed the PCR cycling parameters listed above both PCR reaction batches were mixed together and the cycling parameters listed above were one time more applied, the amount of cycles was raised to 20
- Digestion of the parental DNA with DpnI: Addition of 1 µl DpnI to the PCR batch and incubate for 1 h at 37 °C.
Transformation of E. coli XL1 blue with P65
Investigator: Jeff, Rosario, Johanna, Andi, Flo, Leonie
Aim of the experiment: Transformation of E. coli XL1 blue with P65 .
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 2 µl of DNA was added to 100 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Transformation of E. coli XL1 blue with P66
Investigator: Jeff, Rosario, Flo, Johanna, Andi, Leonie
Aim of the experiment: Transformation of E. coli XL1 blue with P66.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 2 µl of DNA was added to 100 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Transformation of E. coli XL1 blue with P67
Investigator: Jeff, Rosario, Johanna, Andi, Leonie, Flo
Aim of the experiment: Transformation of E. coli XL1 blue with P67.
Procedure:
- CaCl2 competent E. coli XL1-Blue cells were put out from the stock in -80 °C freezer and were gently thawed on ice.
- 2 µl of DNA was added to 100 µl of competent cells and gently mixed.
- 30 min incubation on ice
- 5 min. heat shock at 37 °C
- Adding of 1 ml LB-medium to each tube.
- Incubation for 45 min at 37 °C in the 180 rpm cell-culture shaker.
- 100 µl of the cell suspension was plated on one chloramphenicol plate.
- The rest were centrifuged for 1 min at 13000 rpm and the supernatant was dicarded.
- The pellet was resuspended in 100 µl of LB-medium and this concentrated cell suspension was plated again on a new chlorampenicol plate.
Tuesday, May 14th
Analytical digestion of P15 and P19 to P52
Investigator: Leonie, Florian, Johanna
Aim of the experiment: Analytical digestion and gelelectrophoresis of P15 and P19 to P52 to check the sequencing results.
Procedure:
- Batch for analytical digestion of P15 and P19 to P52
| volume | reagent |
| 5 µl | Plasmid DNA |
| 5 µl | Mastermix |
| =10 µl | TOTAL |
Batch of Mastermix
| volume | reagent |
| 40 µl | NEBuffer 4 (10x) |
| 4 µl | BSA (100x) |
| 0.2 µl | NotI(10 U/µl) |
| 148 µl | ddH2O |
| =200 µl | TOTAL |
- Incubation for 90 min at 37 °C.
- Analytical gelelectrophoresis was performed at 90 V for 60 min.
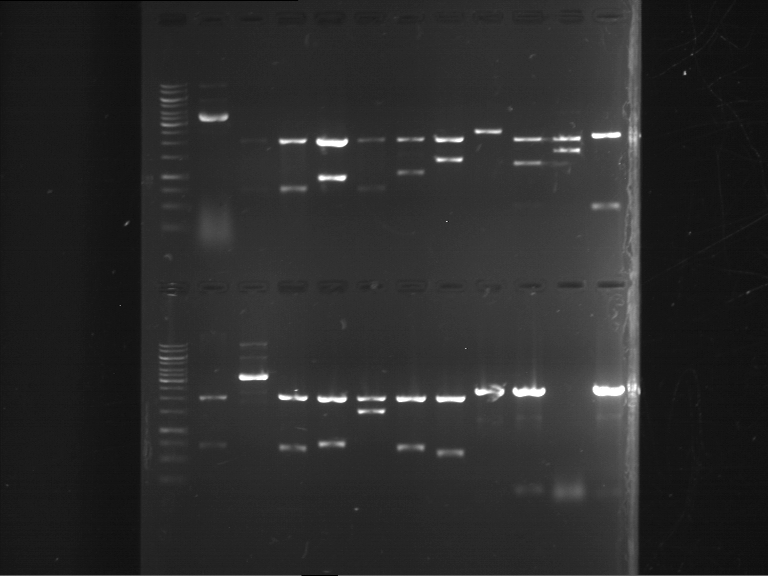
- Discussion:
- Gel 1, Row 1:
| lane | part | plasmid | band 1 | band 2 | result |
| 2 | EreA | pDEST14 | 3,25 kb | no cut, no restriction sites in pDEST14 | |
| 3 | mCherry(LMU) | pSB1C3 | 2,0 kb | 750 bp | probably right |
| 4 | GFP(LMU)(845 bp) | pSB1C3 | 2,0 kb | 750 bp | maybe switched with mKate2(LMU) |
| 5 | mKate2(LMU)(702 bp) | pSB1C3 | 2,0 kb | 900 bp | maybe switched with GFP(LMU) |
| 6 | 35S + Strong Plant + GFP(1775 bp) | pSB1C3 | 2,0 kb | 750 bp | ? |
| 7 | UVR8(1332 bp) | pSB1C3 | 2,0 kb | 1,1 kb | ? |
| 8 | mStrawberry(708 bp) | pSB1C3 | 2,0 kb | 1,4 kb | ? |
| 9 | DREB1C promoter(463 bp) | pSB1C3 | 2,5 kb | probably only one cut -> one NotI site mutated | |
| 10 | Laccase(1587 bp) | pSB1C3 | 2,0 kb | 1,3 kb | ? |
| 11 | SV40(419 bp) | pSB1C3 | 2,0 kb | 1,6 kb | this could be the laccase |
| 12 | cjBlue(696 bp) | pSB1C3 | 2,0 kb | 0,5 kb | ? |
- Gel 1, Row 2:
| lane | part | plasmid | band 1 | band 2 | result |
| 2 | alcA (843 bp) | pSB1C3 | 2,0 kb | 0,7 kb | ? |
| 3 | eforRed (681 bp) | pSB1C3 | plasmid | plasmid bands (coiled, supercoiled, etc.) | |
| 4 | amilGFP (699 bp) | pSB1C3 | 2,0 kb | 0,7 kb | right part |
| 5 | mCFP (714 bp) | pSB1C3 | 2,0 kb | 0,7 kb | right part |
| 6 | RD29A promoter (1535 bp) | pSB1C3 | 2,0 kb | 1,5 kb | right part |
| 7 | amilCP (669 bp) | pSB1C3 | 2,0 kb | 0,7 kb | right part |
| 8 | CMV promoter (580 bp) | pSB1C3 | 2,0 kb | 0,6 kb | right part |
| 9 | middle linker (24 bp) | pMA-BBFR | 2,4 kb | short insert went through the gel | |
| 10 | GSAT (108 bp) | pMA-BBFR | 2,4 kb | 0,1 kb | right part |
| 11 | GFP(720 bp) | pSB1A2 | ? | ||
| 12 | long linker (36 bp) | pMA-BBFR | 2,4 kb | short insert went through the gel |
- Gel 2:
| lane | part | plasmid | band 1 | band 2 | result |
| 2 | SEG linker (108 bp) | pMA-BBFR | 2,4 kb | short insert went through the gel | |
| 3 | short linker (12 bp) | pMA-BBFR | 2,4 kb | short insert went through the gel | |
| 4 | FluA (522 bp) | pMA-BBFR | 2,4 kb | 0,5 kb | right part |
| 5 | mCherry (769 bp) | pSB2K3 | 4,4 kb | 0,75 kb | right part |
| 6 | Cerulean (778 bp) | pSB2K3 | ? | ||
| 7 | mRFP1 (706 bp) | pSB2K3 | 4,4 kb | 0,7 kb | right part |
| 8 | mOrange (769 bp) | pSB2K3 | 4,4 kb | 0,75 kb | right part |
| 9 | PIF3-20aa (366 bp) | pSB1C3 | 2,0 kb | 0,35 kb | right part |
| 10 | 20aa-PIF3 (366 bp) | pSB1C3 | 2,0 kb | 0,35 kb | two plasmids and two insert can be seen |
| 11 | LexA (ca. 6 kb) | pSB1C3 | > 8 kb | only one cut, huge plasmid |
Picking of Plasmid P23 (CaMV35S) and Ligation Product F8 + F15 (TEV)
Investigator: Jeff, Florian
Aim of the study: Picking of Plasmid P23 (CaMV35S) and Ligation Product F8 + F15 (TEV) for Miniprep.
Operational sequence:
- Picking and overnight culture after standard laboratory's protocol. (CamR LB-medium)
- 1 colony of P23 were picked.
- 7 colonies of F8 + F15 were picked.
Sequencing of P63 & P64
Investigators: Louise
Aim of the experiment: Sequencing of genes for which no prior sequencing results were available.
Procedure: The DNA was prepared for sequencing according to the Eurofins SmartSeq Kit protocol (15 µl of plasmid DNA (50 - 100 ng) and 2 µl sequencing primer).
The different genes we sequenced received the following barcodes:
| Label | Name | Barcode1 | Barcode2 |
|---|---|---|---|
| p63 | EreA | FR01002349 | FR01002350 |
| p64 | EreA | FR01002351 | FR01002352 |
 "
"