Team:NCTU Formosa/results
From 2013.igem.org
Mike821005 (Talk | contribs) (→Expected sRNA regulation efficiency) |
|||
| Line 16: | Line 16: | ||
</p> | </p> | ||
| - | <p> | + | <p>We measured the OD value along with the florescence expression of the reporter gene for each biobrick mentioned above. Fig 3. is the result obtained. We calculated the normalized expression of the y-axis by dividing fluorescence expression with the OD value measured, since the higher the OD value is, the larger the amount of bacteria that can express florescence would be. As shown in Fig 3., the the normalized expression of the biobrick with B0034 is the highest and the one with target 1, K1017202, is second highest, while the other two of B0032 and B0030 show weak expressions. This result implies that target 1 can, in fact, serve as a functional RBS. In comparison to other RBS, target 1 can provide moderate translational efficiency that is just lower than that of highly efficient B0034 and higher than those of commonly used RBS such as B0030 and B0032. </p> |
<span class="undetermined">(undetermined)</span> | <span class="undetermined">(undetermined)</span> | ||
Revision as of 10:22, 24 September 2013
The current progress of our project, including detailed information of the experimental data and the overall evaluation of the practicability of this project.
Contents |
RBS efficiency
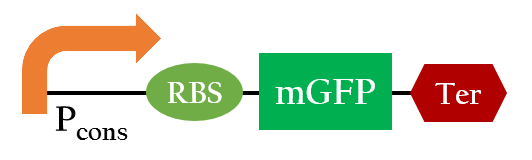
We used the following biobrick to test the efficiency of different ribosome binding sites:
- Pcons+BBa_B0034+mRFP+Ter
- Pcons+[http://parts.igem.org/wiki/index.php?title=Part:BBa_K1017202 BBa_K1017202]+mRFP+Ter
- Pcons+BBa_B0030+mRFP+Ter
- Pcons+BBa_B0032+mRFP+Ter
- control:pet40
We measured the OD value along with the florescence expression of the reporter gene for each biobrick mentioned above. Fig 3. is the result obtained. We calculated the normalized expression of the y-axis by dividing fluorescence expression with the OD value measured, since the higher the OD value is, the larger the amount of bacteria that can express florescence would be. As shown in Fig 3., the the normalized expression of the biobrick with B0034 is the highest and the one with target 1, K1017202, is second highest, while the other two of B0032 and B0030 show weak expressions. This result implies that target 1 can, in fact, serve as a functional RBS. In comparison to other RBS, target 1 can provide moderate translational efficiency that is just lower than that of highly efficient B0034 and higher than those of commonly used RBS such as B0030 and B0032.
(undetermined)
37 degrees Celsius RBS
Using biobrick, Pcons+37°C RBS+mGFP+J61048, we tested the function of the 37°C RBS under different temperatures.
As shown on Fig 3. the expression of mGFP under 37°C and 42°C reached a higher OD value in less time than the expression below 37°C. This demonstrates the fact that 37°C RBS functions more efficiently above or at 37°C. On the other hand, the fact that 37°C RBS leads to moderate expression even under 37°C degree might be a problem, since then we cannot make sure that only single gene is expressed under 37°C.
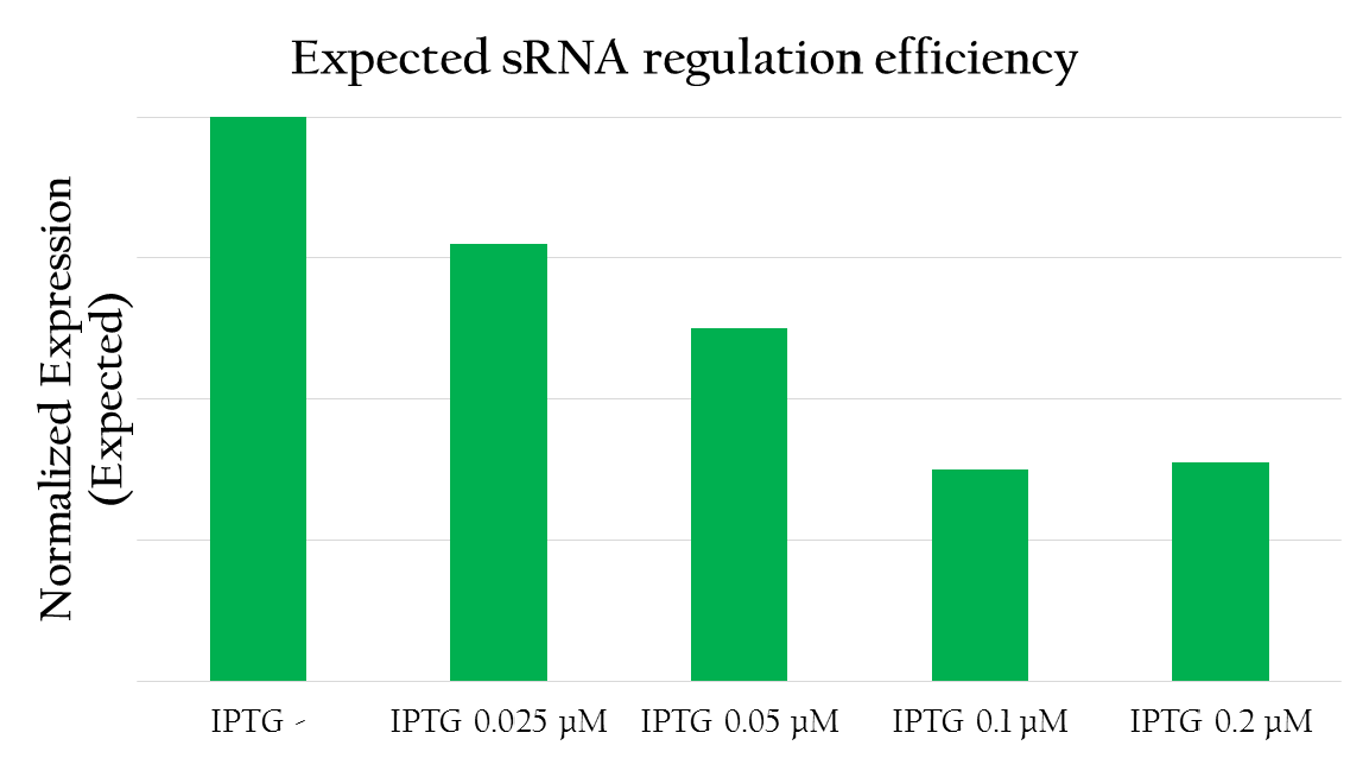
Expected sRNA regulation efficiency
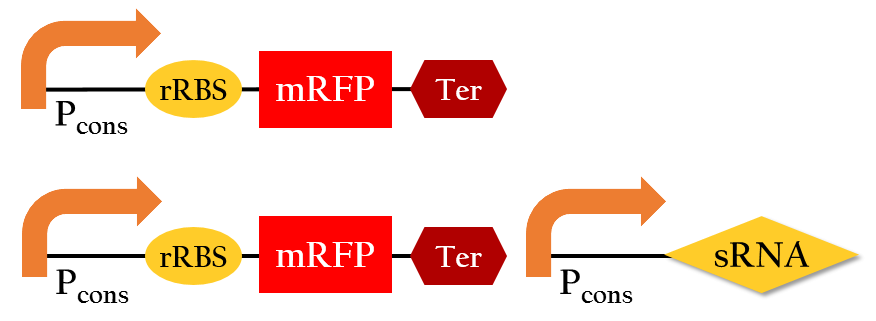
To test the regulation efficiency of sRNA, we employed the following biobricks: Pcons + rRBS + mGFP + J61048 and Pcons + B0030 + lacI + J61048 + Plac + sRNA.
The graph above shows the relationship between the concentration of IPTG added and the red florescent measured. The more IPTG added, the more sRNA can be produced by Plac as IPTG serves to activate the promoter. In other words, the amount of sRNA and the concentration of IPTG forms a liner relationship. With more IPTG, and therefore, more sRNA the amount of red fluorescent measured decreases. This implies that sRNA can effectively regulate the expression of RFP by decreasing the translation efficiency and the stability of rRBS.
 "
"