Team:BYU Provo/Notebook/CholeraDetection/Fallexp/Period1/Dailylog
From 2013.igem.org
| Line 130: | Line 130: | ||
<font size="4"> '''9/14/13''' </font> | <font size="4"> '''9/14/13''' </font> | ||
[[File:9.14.13_56.jpg|250px|center|link=]] | [[File:9.14.13_56.jpg|250px|center|link=]] | ||
| - | |||
{{TeamBYUProvoFooter}} | {{TeamBYUProvoFooter}} | ||
Latest revision as of 06:38, 26 September 2013
| ||
|
|
9/4/2013 BYU iGEM can count on every member, now that summer is over and all are again enrolled in classes. We took time at the beginning of class to list our main projects (aside from research itself) from here on out: they include cloning our parts into the iGEM backbone plasmid, filming a synthetic biology techniques video to collaborate with another team, finishing the website, and publishing our synthetic biology Children's Book and collecting data to demonstrate that children who read it and the included parental guide improve their understanding of science generally and synthetic biology in particular. Kelton and I are responsible to see the book published, and the data collected. We set ourselves the following deadlines: Sep. 8th: Have in our possession the edited, shaded version of the book. Redge (the illustrator) confirmed that he would finish by Sunday the 8th. Sep. 11th: Submit the book for publishing. Sep. 20th: Distribute paper copies/pdf copies to elementary school teachers and parents. Children will be given a pre test before reading the book, and later a post test. The test results will allow us to quantifiably show that our book improves understanding of synthetic biology among young children. Sep. 25th: Compile and analyze the data. Several weeks ago, I read a paper indicating that lysogenic lambda is induced to lysis by host cell recognition of AHL's, the molecules used in bacterial quorum sensing systems. I thought that our bacterial strain with the Lambda lysogen, TT9907, might recognize cholera's autoinducer, and perhaps that might trigger Lambda. I streaked a line of TT9907 next to cholera and then away from cholera. No bacteria grew next to cholera, but the strain did grow away from cholera! Now, we need to show that it is indeed lambda that is killing the cell and not some toxin from cholera. We designed an experiment with 6 bacterial strains: 2 without the lambda lysogen, 1 with the lambda lysogen, and three that may have the lambda lysogen. The latter three strains are each from single colonies that grew following an electroporation procedure to get TT9907 to take up the plasmid pIG87. Amazingly, top agar lawn assays with a drop of hydrogen peroxide added to stress lambda into lysis show that after electroporation, there are no plaques. Either Lambda gets out during electroporation or TT9907 loses its susceptibility to lambda. Anyway, we took these six strains and streaked a line of them next to a patch of cholera and away from the patch. Depending on how the bacteria grow we'll be able to say if cholera (perhaps some toxin) kills TT9907 or lambda kills TT9907. KK, KP Set up transformations on the third for pIG91 (pSB1C3)into DH5alpha. I used a purified plasmid from the 2012 iGEM plate and from the tube provided this year. 2012 sample didn't work. 50ul phusion PCRs- This is to obtain biobrick parts of Cro and Qrr4/RFP for the database. Cro: template pIG87 primers BI264/265. Qrr4/RFP: template pIG89 primers BI266/267. PCR was successful. The samples were cleaned up. On the fourth mini preped pIG91. Set up 30ul digests with pIG91, Cro PCR, and Qrr4/RFP PCR. Restriction enzymes used were XbaI and SpeI. pIG91 didn't show up at all on the low melt. Cro was a smear but I cut out what I could. Qrr4 looked good. Set up 50ul PCRs for SdiA (723bps). Two reactions were set up both had the template E.coli (didn't matter what kind) and primers BI282/283. For one reaction I used phusion polymerase and the other I used taq. The taq reaction didn't turn out well but the other was fine. Cleaned it. CH
9/6/13 Our results from the side-by-side plate comparison of cholera with several strains of bacteria are encouraging! We grew a patch of V. cholerae for one week on LB plates. Then, we streaked a line of a bacteria adjacent to the patch and another away from the patch. We did this for six different strains on six different plates. None of the bacterial strains plated adjacent to cholera grew. All strains grew away from cholera. However, TT9907, the strain with lambda lysogen, showed plaques! There were distinct plaques on the line of bacteria leading away from Cholera. Through what we believe is a quorum sensing pathway, the lambda lysogen recognizes that E.Coli has detected cholera. It mobilizes excision from the genome, replicates, and lyses it's host. We prepared top agar lawns of our 6 strains, with cholera plated in the middle, to show more clearly the result we saw today. Clarice transformed the iGEM backbone into E.Coli and grew it up in an overnight. From those cells we isolated the plasmid, pIG91, in two Eppendorfs, at concentrations of 52 ng/uL and 82 ng/uL. Monday we will digest the plasmid and the SdiA gene to splice it in and submit to the registry. KK, KP
9/9/13 The children's book is shaded! We read it in class today and confirmed that everyone enjoyed it. Redge needs only to make a few grammatical edits. Hopefully tomorrow or Wednesday we'll get an appointment with Dr. Jaime Jensen, a member of the biology department here at BYU. Dr. Jensen earned her doctorate in Biological Science Education, and we hope that she will give us some good ideas on how to maximize the educational value of our book and its appending parental guide. And, we have wonderful news from the top agar lawns we plated on Wednesday. Our results confirm that cholera does NOT kill bacteria that grow next to it, but DOES indeed stress bacteriophage lambda such that it lyses its host cell. See our photos to the right! KK, KP Assessed all the cloning that we need to do soon. Constructs that need to be finished: Qrr4/RFP in pIG91, Cro in pIG91, SdiA in pIG91, FtsQ/GFP in pIG91, and Ydiv/GFP in pIG91. FtsQ and Ydiv are both reporter proteins for SdiA. Set up 50ul phusion PCRs for cloning 1.FtsQ promoter, 2.ydiV promoter, and 3.HapA/GFP soeing PCR. 1. template E.coli primers BI287/286. 2. template E.coli primers BI288/284. 3. template pIG09 (pGLO) primers HapA/GFP first half and BI285. Everything had bright bands except #3. Need to do next soeing step in PCR for #1 and 2. CH
9/11/13 At 2:00 we joined Dr. Jaime Jensen for an insightful, productive and encouraging meeting about our book. Dr. Jensen has her Ph.D in biology education, and was able to offer some very good insights on our book, its parental guide, and the pre/post test questions we will administer. Unfortunately, we have a significant roadblock: our project has not been IRB approved. We weren't aware that in order to ask children to take a test on their understanding of synthetic biology and collect data there must be IRB approval. So, we've either got to luck out with an exempt status, or rethink collecting data from school classrooms. It would be less paperwork for us if we cleared a parental consent form with the IRB and visited a library, where we could ask parents to sign the form right there. I found a protocol for purifying bacteriophage lambda! We hope to be able to do that tomorrow. KK, KP Set up 50ul Digestions as follows: 1. Cro PCR clean(redo), XbaI/SpeI, NEB4. 2. pIG91 mini, XbaI/SpeI, NEB4 (added CIP with NEB3 for 30mins at end of digestion). 3. SdiA PCR clean, PstI/XbaI, NEB4. 4. pIG91 mini, PstI/XbaI, NEB4. 5. Hap/GFP soeing PCR clean, PstI/EcoRI-HF, NEB4. 6. pJG80 mini, PstI/EcoRI-HF, NEB4. 7. pIG91 mini, EcoRI-HF/PstI, NEB4. All of them appeared to work fine. Ligations: 1. Cro into pIG91 (Xb/Sp). 2. Qrr4/RFP into pIG91 (Xb/Sp). 3. pIG91 (Xb/Sp) vector only control. 4. SdiA into pIG91 (Pst/Xb). 5. vector only control pIG91(Pst/Xb). 6. HapA/GFP (Pst/Eco) into pIG80 (Pst/Eco). 7. pIG80 (Pst/Eco) vector only control. After letting them ligate for about half an hour transformed into dH5alpha. 1-5 were plated onto LB-chloramphenicol and 6,7 onto LB-Amp. Results: 1. many colonies. 2. 5 colonies. 3. 2 colonies. 4. many colonies. 5. many colonies. 6. none. 7. none. Collected colonies that were the transformants we wanted for further verification. 50ul soeing PCR to attach GFP to FtsQ or ydiV (SdiA reporters). 1. Template pIG09 primers FtsQ PCR clean and BI285. 2. Template pIG09 primers ydiV PCR clean and BI285. #1 didn't work need to redo. #2 worked fine cleaned up. CH
9/12/13    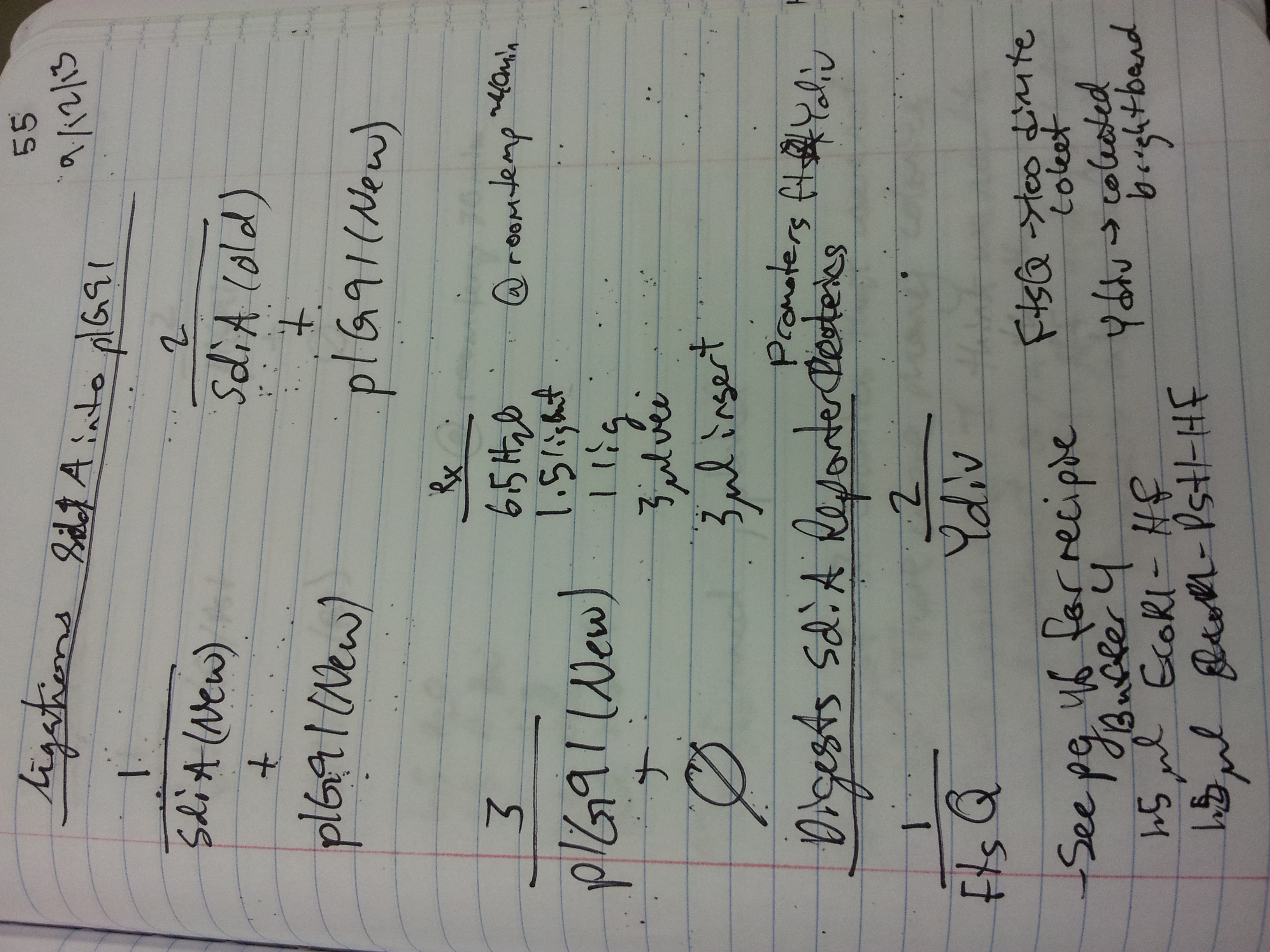
9/13/13 Yesterday, Thursday, we purified bacteriophage lambda, and today we set up spot tests to verify the presence of the phage. Our purification protocol was as follows: we grew overnight cultures of TT9907 with lambda lysogen in a 30 degree Celsius incubator for 18 hours. After incubation, we heat shocked the culture in 2 mL aliquots at 43 degrees Celsius for 10 minutes. We recombined the aliquots and incubated them at 37 degrees Celsius for 6.5 hours to allow lambda to replicate. We again aliquoted the cultures into 1.5 mL aliquots, added 400 uL chloroform, vortexed, and allowed to sit for 10 minutes. After sitting, we centrifuged at 8000 rpm for 10 minutes to separate cellular debris from bacteriophage in the supernatant. We added finishing touches to our book, including a dust jacket and a few blurbs on the back, and placed the order with an online publishing company! That will give us one hard copy; Monday at noon we also have a meeting with Giovanni Tata, the director of BYU Creative Works, to discuss publication of the book!! KK, KP
9/14/13 
|
|
 "
"