Team:KU Leuven/Project/Glucosemodel/EBF
From 2013.igem.org
| Line 260: | Line 260: | ||
<div class="row-fluid"> | <div class="row-fluid"> | ||
<div class="span12 white"> | <div class="span12 white"> | ||
| - | <div class=" | + | <div class="span6"> |
<p align="justify"> | <p align="justify"> | ||
As another approach to prove our constructs (<a href="http://parts.igem.org/Part:BBa_K1060009">BBa_K1060009</a>, <a href="http://parts.igem.org/Part:BBa_K1060011">BBa_K1060011</a> and <a href="http://parts.igem.org/Part:BBa_K1060014">BBa_K1060014</a>), we performed the protein extraction experimet.<br/> | As another approach to prove our constructs (<a href="http://parts.igem.org/Part:BBa_K1060009">BBa_K1060009</a>, <a href="http://parts.igem.org/Part:BBa_K1060011">BBa_K1060011</a> and <a href="http://parts.igem.org/Part:BBa_K1060014">BBa_K1060014</a>), we performed the protein extraction experimet.<br/> | ||
| Line 266: | Line 266: | ||
</p> | </p> | ||
</div> | </div> | ||
| - | <div class=" | + | <div class="span6"> |
<img src="https://static.igem.org/mediawiki/2013/7/76/EBFS_protein.JPG"> | <img src="https://static.igem.org/mediawiki/2013/7/76/EBFS_protein.JPG"> | ||
<p align="justify" class="greytext"> | <p align="justify" class="greytext"> | ||
Revision as of 23:27, 4 October 2013
Secret garden
Congratulations! You've found our secret garden! Follow the instructions below and win a great prize at the World jamboree!
- A video shows that two of our team members are having great fun at our favourite company. Do you know the name of the second member that appears in the video?
- For one of our models we had to do very extensive computations. To prevent our own computers from overheating and to keep the temperature in our iGEM room at a normal level, we used a supercomputer. Which centre maintains this supercomputer? (Dutch abbreviation)
- We organised a symposium with a debate, some seminars and 2 iGEM project presentations. An iGEM team came all the way from the Netherlands to present their project. What is the name of their city?
Now put all of these in this URL:https://2013.igem.org/Team:KU_Leuven/(firstname)(abbreviation)(city), (loose the brackets and put everything in lowercase) and follow the very last instruction to get your special jamboree prize!


E-β-Farnesene
In this part, we will give some more information about the E-β-farnesene (aka EBF) part of the project. EBF is an alarm pheromone, released by almost all of the 4000 aphid species known thus far in response to the presence of predators (eg the ladybug) or other disturbances. In response to the produced EBF, aphids change their metabolism and turn into a winged form, allowing them to "flee the scene" and thus increase their survival rate. Apart from the short term repelling effect, EBF can also cause long term effects : changes in aphid’s development, fecundity, survival when introduced to different growth stages, etc. Moreover, natural aphid predators such as the ladybugs are attracted by EBF.
Hence, having our BanAphids produce EBF should help to repel aphids from our plant of choice. In the following sections, we will give you a general background of EBF synthase followed by an overview of the model and the genes, the wetlab work and the biobricks we built for the EBF part. Finally we have made some suggestion how to optimise the production of EBF in the future. For this we will take you on a tour through the pathways that result in EBF and the problems that arise with this. Of course we have added possible solutions to these problems.
We cloned and expressed the EBF synthase gene in E. coli, to break up (2E,6E)-farnesyl diphosphate into (E)-β-farnesene (EBF) and diphosphate (see reaction scheme below).
The enzyme prefers bivalent cations as cofactors; eg a Mg2+ concentration of 5 mM should be beneficial for EBF function. The ideal pH for EBF synthase will be between 5.5-7.

The EBF construct we designed consists of constitutive promoter with a lac operator, the EBF synthase itself and a double terminator. EBF is not only made by aphids but also by plants and other organisms in a form of bio-mimicry. We obtained two different sources of the EBF gene. One gene originates from the soil bacterium Streptomyces coelicolor (Centre of Microbial and Plant Genetics of KU Leuven). We chose this plant-residing bacterium because it would be a perfect chassis for the ultimate expression of EBF in our E. coligy system. The other EBF gene is from the plant Artemisia annua (sweet wormwood) and was a kind gift from Professor Peter Brodelius (Kalmar University, Sweden). Here we were inspired with the plant origin. The KM for the Artemisia annua protein is calculated at 0.0021 mM, with a Kcat/KM=4.5 and a turnover number of 0.0095 s-1. For the Streptomyces coelicolor protein the KM is 0.0168 mM and the turnover number 0.019 s-1.
Unfortunately, the EBF synthase from Streptomyces coelicolor is a bifunctional enzyme, not only processing β-farnesene but also containing albaflavenone synthase activity. For this reason, we chose to follow up on the Artemisia annua gene and product.
For our construct, our first choice was a medium strength promoter with medium RBS (BBa_K608006); we nonetheless also made the construct with a strong promoter and RBS. We used BBa_B0015 for the double terminator. The lac operator in front of the EBF synthase gene will allow us to switch on/off the transcription of EBF synthase gene.
Gettin' the gene
In the case of EBF synthase gene from Streptomyces coelicolor, we amplified this gene with the a colony PCR. The EBF synthase gene from Artemisia annua was received in the pET28 vector from professor Brodelius (Kalmar University, Sweden). In this gene an additional EcoRI restriction site is present, which will conflict with the standard iGEM cloning work. Therefore we removed this site via site directed mutagenesis after transferring the gene into the iGEM pSB1C3 backbone.
Cutting and pasting
Once we obtained the target gene (EBF) in the standard pSB1C3 backbone, we started our cloning work. We used a pSB1C3 with promoter or terminator as chassis, cut this open and inserted the gene of interest. The 3A assembly method was too difficult most likely because the size of promoter and terminator are so different compared to the gene insert. When ligating the insert in front of the double terminator, we cut the vector with EcoRI and XbaI, and the insert with EcoRI and SpeI. The promotor vector on the other hand is cut with SpeI and PstI restriction sites, and the insert is cut with XbaI and PstI restriction sites. This works because SpeI and XbaI are isoschizomers.
Ligations were performed in parallel in two different ways. In one setup we ligated for 20 minutes at 16 ℃, and in comparison, the second ligation of the same products was conducted at 16 ℃ overnight.
For transformation, we used both chemically competent cells and electrocompetent cells. Electroporation had a higher efficiency when compared to heat shock transformation.
Confirmation
After we observed observed colonies the next day, we needed to confirm the products. The first step we did was usually a colony PCR to check if the insert was in the vector, this was followed up by digestion confirmation after the plasmid extraction. Only the plasmids which succeeded in both controls were send for sequencing, the final confirmation.
gBlocks
Meanwhile, we also built the EBF construct with a lac operator between the promoter and gene, using the gBlock principle. We designed the gBlocks, assembled them and ligated the insert into pSB1C3 backbone. The colonies obtained also went through the three confirmation steps mentioned above before we were satisfied.
For more details of the labwork and the wetlab difficulties as well as how we overcame them, please consult our wetlab journal.
After we overcame a lot of difficulties, we finally made the following bricks at the end of the summer.





With our constructs being ready, we tested them with several aphid experiments.
Our pilot experiment tested brick BBa_K1060009. We placed aphids on a leaf in the middle of a huge petri dish, an EBF-producing bacterium plate on the left, a control on the right, the results video seemed positive since we observed that the general trend of aphid movement was towards the direction away from the EBF-producing bacterium.
Moreover, we tried another set-up with brick BBa_K1060011. This time we connected the leaves that were on EBF-producing bacterium plate and control plate with the leaf in the middle where the aphids resided, therefore they have the opportunity to move onto the other leaves. However there was no significant difference between the amount of aphids on the two leaves. This may due to the fact that in this construct, there is a lac operator after the promoter, the lacI in the bacteria may interfere with the production of EBF.
In addition, we also examined the aphid's behavior without a leaf as a starting point. We put 30 aphids in the middle of a huge petri dish, on the left side sat a leaf with 10µl of EBF-producing bacteria and on the right side placed a normal leaf as control. Hence the aphids were offered the chance to go searching for food. After 2 hours we counted the number of aphids on the leaves, there were 4 aphids on the leaf where the EBF was produced and 6 aphids on the control leaf, the rest of aphids just walked randomly in the big petri dish. Due to the small amount of aphids used, this did not yield significant results.
Our pilot experiments indicated a trend in the right direction, however, to finally confirm this with sufficient scientific confidence level, more experiments with larger amounts of aphids have to be performed. Several aspects of the setup will be optimized in the future. For example the amount of bacteria, the strength of the promoter, the ventilation of the setup, the incubation time and the temperature, etc. The reason for this is that the concentration of EBF is essential to trigger the desired response in the aphids. Both too high and too low concentrations will lead to aphid insensitivity.
EBF synthase expression experiment
As another approach to prove our constructs (BBa_K1060009, BBa_K1060011 and BBa_K1060014), we performed the protein extraction experimet.
The size of EBF synthase is around 66kDa, even though we cannot see a clear band of desired size in the SDS-PAGE gel, it could be due to the amount of the protein is not sufficient to show up in the gel but sufficient to be active in our aphids experiments. Since the protein is not tagged, we cannot extract them with western blott. It's highly recommendable to perform a gas chromatography to confirm the presence of EBF synthase.

β-farnesene is a terpenoid that is converted from farnesyl pyrophosphate (FPP) by the enzyme β-farnesene synthase (EC 4.2.3.47).
FPP is the precursor of β-farnesene, that is produced by the building blocks, the molecules isopentenyl pyrophosphate (IPP) and its isomer dimethylallylpyrophosphate (DMAPP).
These precursors of farnesyl pyrophosphate can be produced by several metabolic pathways. Most prokaryotes use the non-mevalonate or DXP pathway, producing IPP starting from glyceraldehyde-3-phosphate and pyruvate. Eukaryotes, except for plants, exclusively use the mevalonate pathway, producing IPP starting from acetyl-CoA. Plants use both pathways.
On the left you can see the non-mevalonate pathway or DXP pathway, showing the conversion of pyruvate and glyceraldehyde-3-phosphate to the terpenoid precursor IPP and its isomer DMAPP.
Pyr = pyruvate, G3P = glyceraldehyde-3-phosphate, DXP = 1-deoxy-D-xylulose 5-phosphate, MEP = 2-C-methylerythritol 4-phosphate, CDP-ME = 4-phosphocytidyl-2-C-methylerythritol, CDP-MEP = 4-phosphocytidyl-2-C-methyl-D-erythritol 2-phosphate, MEcPP = 2-C-methyl-D-erythritol 2,4-cyclopyrophosphate, HMB-PP = (E)-4-Hydroxy-3-methyl-but-2-enyl pyrophosphate, DXS = DXP synthase, DXR = DXP reductase, CMS = CDP-ME synthase, CMK = CDP-ME kinase, MCS = MEcPP synthase, HDS = HMB-PP synthase, HDR = HMB-PP reductase
Since FPP is such an important precursor, used for the biosynthesis of lots and lots of compounds, it might turn out that once we insert a plasmid containing the β-farnesene synthase gene, we obtain only a very small amount of β-farnesene, since the precursor amount wasn't increased and there simply isn’t enough FPP available to produce the amount of β-farnesene to satisfy the EBF synthase enzyme we brought in.
A solution may be to implement plasmids which engineer a mevalonate pathway in E. coli, thereby upregulating the production of FPP. This larger amount of FPP may then be converted to β-farnesene, creating a large enough amount of this volatile for our uses. This was demonstrated many times by J.D. Keasling in S. cerervisiae, while Martin et al., (2003) implemented this mevalonate pathway in E. coli. In the article, they described their successful efforts to create a high level production of amorphadiene by introducing the mevalonate pathway in E. coli. However, expression of this heterologous pathway led to such an abundance of isoprenoid precursors that cells ceased to grow or mutated to overcome the toxicity.
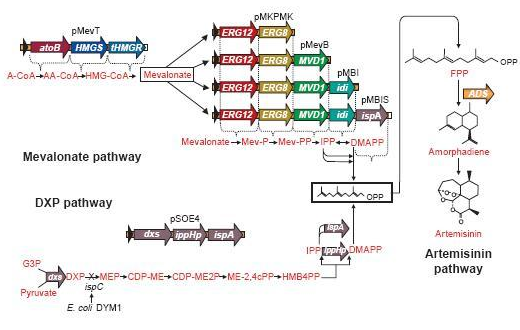
Since there are eight genes responsible for the mevalonate pathway, Martin et al. decided to split them up into two parts. A first plasmid named pMevT, responsible for the conversion of acetyl-CoA to mevalonate, harboring the atoB, HMGS and tHMGR genes into a pBAD33 vector, and a second one named pMBIS, harboring the ERG12, ERG8, MVD1, idi and ispA genes into a pBBR1MCS-3 plasmid. Coexpression of these two operons in an ispC deficient E. coli strain produced the terpenes, even in the absence of mevalonate, indicating that the mevalonate pathway works. A combined expression of their recombinant mevalonate pathway and the synthetic gene product (ADS in their case) resulted in greatly improved yields.
Even though we do not need a very high production of EBF it would be definitely better to optimise the pathway, by using the plasmids pMevT and pMBIS. Implementing them into our BanAphids along with the synthetic β-farnesene synthase gene could result into high yields of β-farnesene. This way the amount of EBF can be easily changed by changing the amount of bacteria or the concentration of Mg2+.
Due to the short amount of time iGEM offered we did not yet started doing this, but this is definitely something future teams might look into.
Kajiwara S., Fraser P., Kondo K., Misawa N., Expression of an exogenous isopentenyl diphosphate isomerase gene enhances isoprenoid synthesis in Escherichia coli, Biochem J. 324, 421-426 (1997).
Martin V., Pitera D., Withers S., Newman J., Keasling J., Engineering a mevalonate pathway in Escherichia coli for production of terpenoids, Nature Biotechnology 21(7), 796-802 (2003).
 "
"