Header
This section provides a short summary of our main experimental achievements.
Contents |
Nanoparticles:
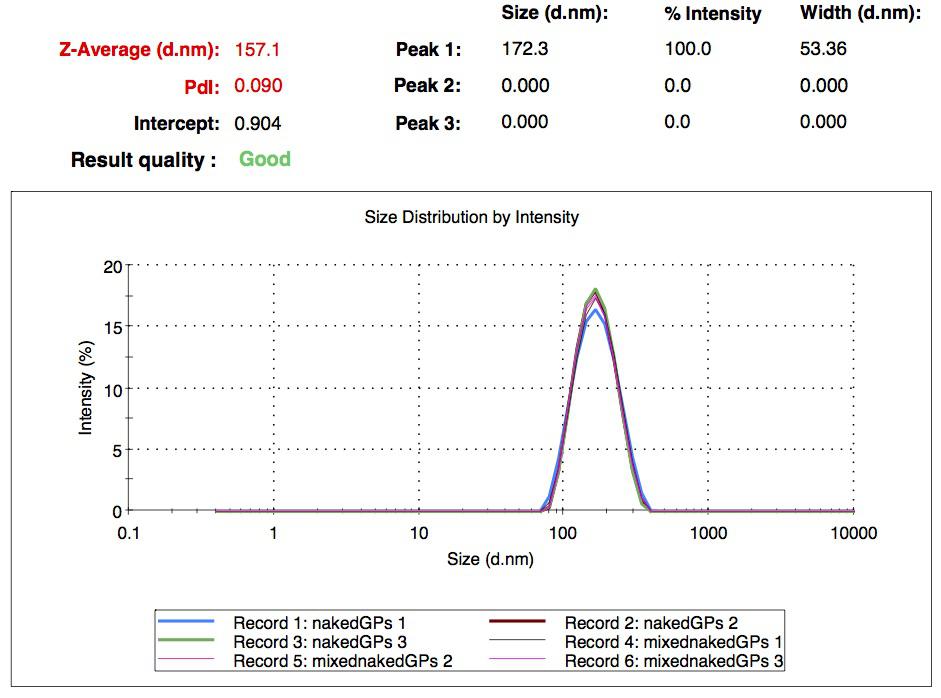
We successfully synthesized different batches of nanoparticles, whose mean diameter is in a range between 200 nm and 300 nm. We ended up with seven different collections of samples: simple naked gelatin nanoparticles, simple biotinylated nanoparticles, CY5-labeled biotinylated nanoparticles, rGFP loaded nanoparticles (naked and biotinylated) and FITC-Dextran loaded nanoparticles (naked and biotinylated). All of them were characterized by DLS. These experiments show that two cargo transport strategies are possible: an external labeling (the one we used with CY5) or an internal loading (with FITC-Dextran and rGFP).
Cell surface expression:
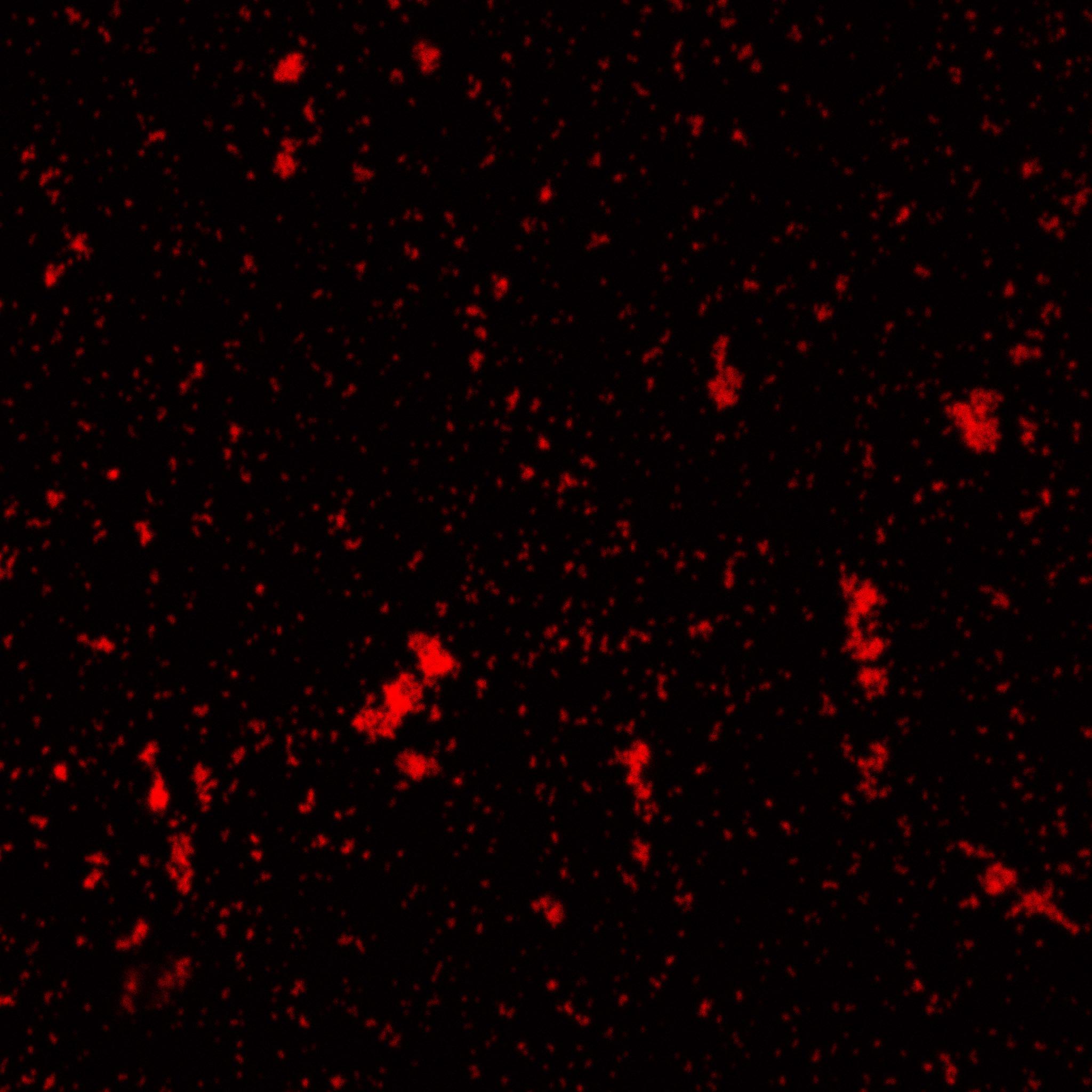
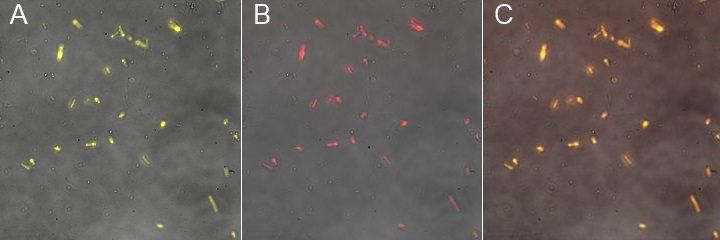
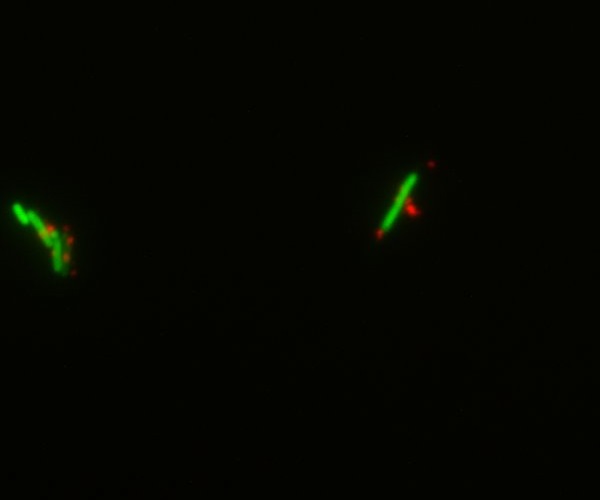
Characterization of an existing part: The part BBa_K523013 (INP-YFP construct to export YFP at the membrane) had been characterized only by a comparison of pellet and supernatant fluorescence after centrifugation. We wanted to characterize it better to be sure that it was expressed on the outer membrane. We successfully showed that the YFP-INP was expressed at the membrane.
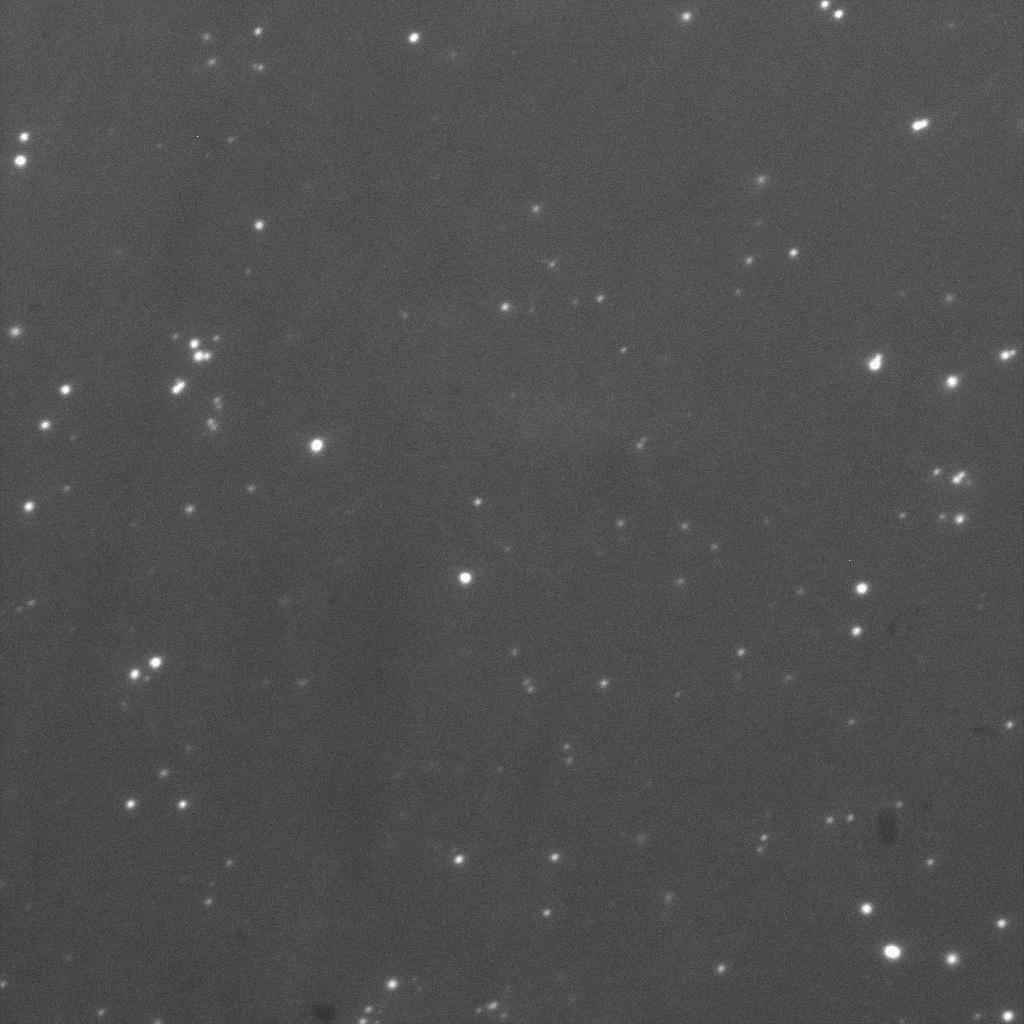
This fluorescence microscopy image shows that YFP direct excitation signal colocalizes with YFP-antibodysignal, meaning that the protein is not only at the membrane, but at the outer one.
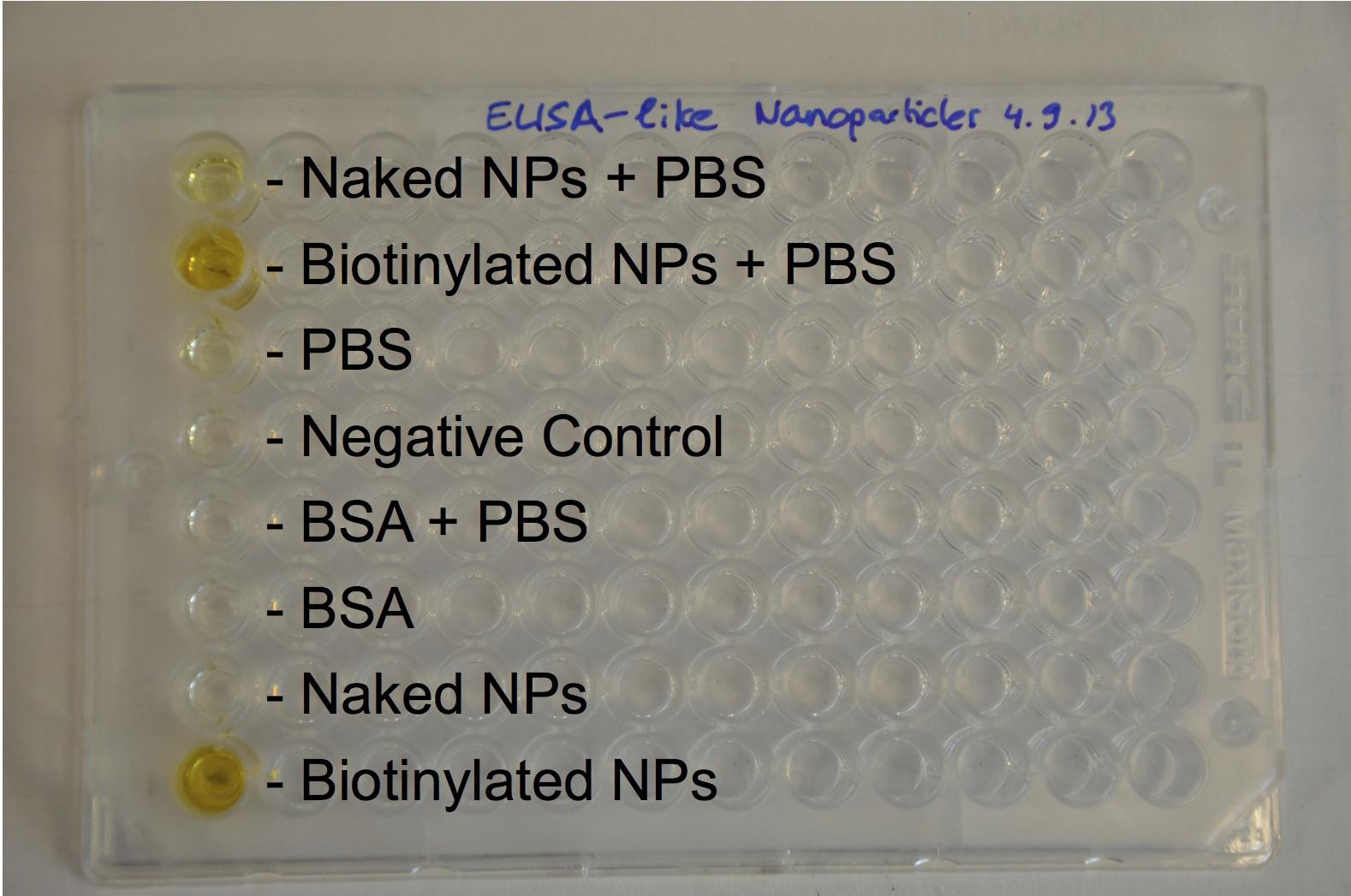
Expression of streptavidin at the cell surface: Gibson assemblies of the three different streptavidin constructs worked. and sequencing results matched with what expected. The growth curve of transformed E.Coli showed delayed growth, but bacteria still divide with an acceptable rate. The assay with a fluorescent biotin supposed to bind streptavidin gave some positive results (some bacteria appeared flurescent when excited at the corresponding wavelenght) but since the negative control also showed fluorescence, nothing could be proved. However, a Western blot against streptavidin showed bands at the expected size (53 kDa) of streptavidin, proving that it was expressed.
Sensing:
We inserted two pH-dependent and one constitutive promoter in front of a superfolded GFP sequence ( [http://parts.igem.org/Part:BBa_I746908 Biobrick BBa_I746916, Main page] ). All three promoters as well as the respective backbones were successfully isolated and amplified by PCR.
The Gibson assemblies also worked and the sequencing results showed a 100% match between the inserted promoters and the refererence sequence.
Fluorescence measurements with the PlateReader were not conclusive since a lot of cells died in acidic pH, supposed to activate the promoter. However, even if we were not able to prove that acidic pH triggers expression of GFP, fluorescence could be seen under the microscope. This indicated that the promoters are functional. The constitutive promoter worked as expected, inducing the expression of superfolded GFP strongly. We used the fact that those cells' fluorescence could be seen by the naked eye and put the plasmid in our human practice kit (Our Kit ).
Effector:
With the exception of MMP9, the Gibson Assemblies worked for both Gelatinase GelE and MMP2. The primers that were designed had introduced a stop codon upstream of GFP. But since we had planned to do constructs with and without GFP attached to the gelatinase, we continued our experiments. The Western Blot with an anti-His tag antibody did not work. The His-Tag it may be hidden in the protein, contain a mutaion or could simply not bind to the nickel columns as they were stored improperly. An assay to purify the two enzymes MMP2 and gelE showed the presence of the proteins by comparing the assay of arabinose induce (expressing enzymes due to the arabinose induced promoter) and non arabinose induced protein.
For triggering expression of degradating enzymes, arabinose sensitive promoter was chosed. The enzymes to insert were MMP9, MMP2 and gelE, the three degrading gelatin. The part BBa_I746908 was the backbone consisting in GFP driven by pBad promoter. The enzymes to insert would be either put instead of GFP or in addition to GFP. All final plasmids would have a His-tag to purify it easily.
Sequences of the enzymes and the backbones were correctly PCR amplified. The Gibson were successfull for the MMP2 and gelE, but cells transformed with MMP9 didn't give any colony.
Sequencing results of the other plasmids showed a stop codon between enzymes and GFP, the reason why they didn't appear green.
However, the enzymes preceding GFP could still be expressed and an His-Tag purification was achieved since His-tag is placed before the stop codon. The different fractions were collected and analyzed on a SDS page :
Overall:
We can say that except minor problems, cloning succedeed well. The Gibson assemblies globally worked out and we had no trouble growing the resulting transformed bacteria. The most delicate part was the characterization of our parts with functionnal assays. However, a lot of parts showed encouraging results but would maybe need to be studied in more detail. The naoparticles is the part that worked out well, nanoparticles could be synthetized and loaded. This project was ambitious and was almost achieved, and we are really proud of sharing our results with the iGEM community!
 "
"