Team:BYU Provo/Notebook/CholeraDetection/Fallexp/Period3/Dailylog
From 2013.igem.org
| Line 87: | Line 87: | ||
<font size="4"> '''10/23/13''' </font> | <font size="4"> '''10/23/13''' </font> | ||
| - | Today we set up top agar tests to check our SdiA- knockout strain. We are hoping to see plaques tomorrow | + | Today we set up top agar tests to check our SdiA- knockout strain. We are hoping to see plaques tomorrow. We plated on four different plates. We plated on a kanemycin plate with cholera added, a kanemycin plate with just the cells, a LB plate with cholera, and a LB plate with H202. |
<br> | <br> | ||
Revision as of 21:37, 23 October 2013
| ||
|
|
10/9/2013 Set up overnights of SdiA+/- strains for later infection with bacteriophage Lambda. Lysogens will plated next to Cholera to prove that SdiA is necessary for lytic induction of bacteriophage Lambda. Begin growing a stockpile of Cholera patches (x10) Begin looking for 3D modeling software. Set up overnights of CRO + DH5alpha for later infection with bacteriophage Lambda. 10/10 Plate top agar assays of SdiA+/-, infect with bacteriophage Lambda, grow at 37 C.
10/11/13 Streak singles and start overnights with LOTS of SdiA+/- lysogens, PLATE NEXT TO CHOLERA. PCR SdiA+ from boiled K12 template and from not-boiled K12. 10/12 Plate SdiA+/- Top Agar Assays with Cholera in the middle (6 SdiA+ lysogens and 6 SdiA- lysogens)
10/14/13 Failed (contaminated) top agar tests!! Remake overnight cultures, and plate more lysogens (Amber and Arick); PCR clean-up of SdiA+, digest with PST1 and Xba1 10/15 Top agar SdiA+/- assays, run SdiA on low-melt gel and ligate
10/16/13 Re-plate top agar assays for SdiA+ and SdiA- lysogens, with cholera or with H202 as a positive control.
10/18/13 This time, our top agar lawns were not contaminated, however, we did not see the results we expected! Not one plate had any plaques, not our lawns of SdiA+ lysogens, nor SdiA- lysogens, not by H202 induction nor induction by v. cholerae's AHLs. So, either our lysogens aren't extremely lysogenic, or the do not have prophage Lambda and are not lysogens at all. There is evidence that they are indeed lysogens, because Wednesday I streaked lines of each lysogen across a place and attempted to infect with a microliter of bacteriophage Lambda. None of the lysogens showed any plaques, meaning that none could be superinfected. We started a PCR reaction for each plate with primers targeting the CRO gene in bacteriophage Lambda. If we see bands on the gel for CRO, then we know that they are lysogens.
10/21/13 We visualized the PCR reactions from Friday on a gel. The only plate that appears to indeed have bacteriophage Lambda is one SdiA- lysogenic lawn (see gel). We began an overnight culture of this lysogen, and plated it on LB/CAM just to ensure that it isn't contaminated, because our SdiA- knockout strain has kanemycin resistance integrated into the genome. 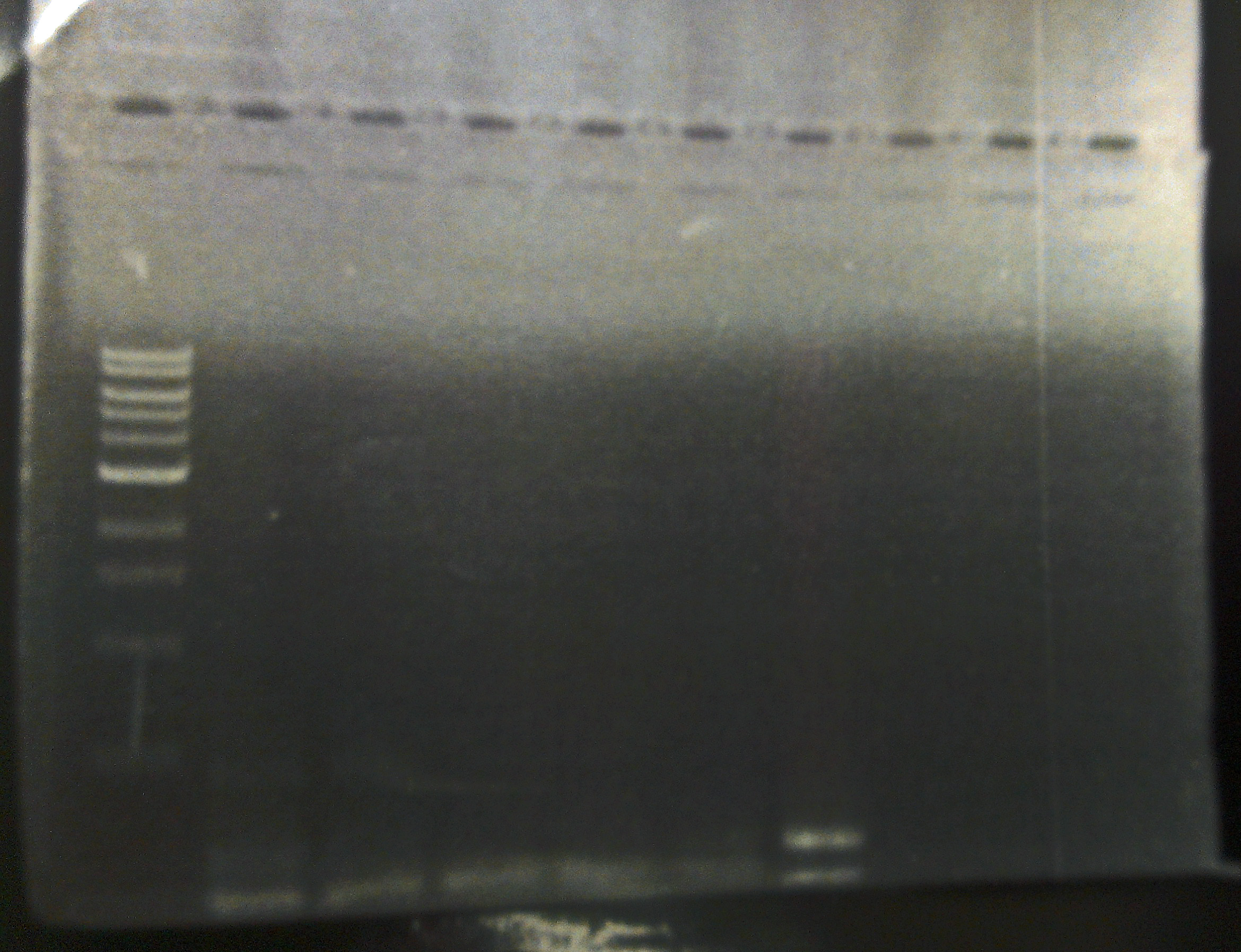
10/23/13 Today we set up top agar tests to check our SdiA- knockout strain. We are hoping to see plaques tomorrow. We plated on four different plates. We plated on a kanemycin plate with cholera added, a kanemycin plate with just the cells, a LB plate with cholera, and a LB plate with H202.
10/25/13
|
|
 "
"