Team:Heidelberg/Templates/DelH week16
From 2013.igem.org
m |
|||
| Line 47: | Line 47: | ||
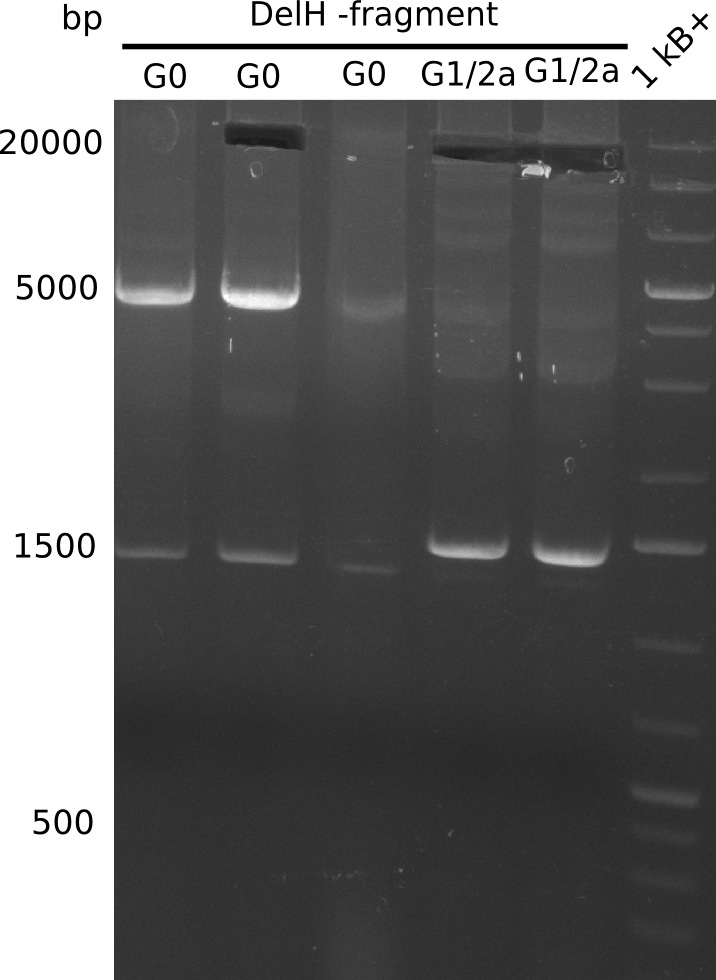
Expected bands: 18 Kb (G0), 13 Kb (G1/2a) | Expected bands: 18 Kb (G0), 13 Kb (G1/2a) | ||
[[File:Heidelberg_20130812 3xG0 2xG1-2a.png|200px|thumb|right|'''Fig.16.1''' gel of amplified DelH-fragment(loaded 20 µL of PCR) <br> ''l1-3:'' G0 (complete DelH-fragment with DN11 & HM08), ''l4-5:'' G1/2b (3/4 of the DelH fragment amplified with DN11 & HM06), ''l6:'' 1kb+ ladder <br> ''l2'' shows the expected band at 18 Kb. ''l4-5'' showed expected band at 13 Kb. All three bands were cut out.]] | [[File:Heidelberg_20130812 3xG0 2xG1-2a.png|200px|thumb|right|'''Fig.16.1''' gel of amplified DelH-fragment(loaded 20 µL of PCR) <br> ''l1-3:'' G0 (complete DelH-fragment with DN11 & HM08), ''l4-5:'' G1/2b (3/4 of the DelH fragment amplified with DN11 & HM06), ''l6:'' 1kb+ ladder <br> ''l2'' shows the expected band at 18 Kb. ''l4-5'' showed expected band at 13 Kb. All three bands were cut out.]] | ||
| - | + | ||
Gel shows expected bands. | Gel shows expected bands. | ||
:=> They were cut and gel isolated. | :=> They were cut and gel isolated. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
===Generation of Plasmid DelH 11-08=== | ===Generation of Plasmid DelH 11-08=== | ||
====Colony-PCR CP.W16.A==== | ====Colony-PCR CP.W16.A==== | ||
| Line 94: | Line 95: | ||
<br/> | <br/> | ||
[[File:Heidelberg_20130812 colonyPCR.png|200px|thumb|right|'''Fig.16.2''' gel of screening PCRs with DN07 & VF2 primer (loaded 20 µL of PCR) <br> ''l1:'' 2log ladder, ''l2:'' colony PCR of 1 µl Gibson transformed,''l3:'' colony PCR of 14 µl Gibson transformed,''l4-5:'' 2 colony PCRs of BB - 14 µl Gibson transformed, ''l6:'' 2log ladder <br> none of the colonies is positive, of course the BB shows not the expected band of 663 bp (for screening). The first colony shows a band at ~1.7 kb.]] | [[File:Heidelberg_20130812 colonyPCR.png|200px|thumb|right|'''Fig.16.2''' gel of screening PCRs with DN07 & VF2 primer (loaded 20 µL of PCR) <br> ''l1:'' 2log ladder, ''l2:'' colony PCR of 1 µl Gibson transformed,''l3:'' colony PCR of 14 µl Gibson transformed,''l4-5:'' 2 colony PCRs of BB - 14 µl Gibson transformed, ''l6:'' 2log ladder <br> none of the colonies is positive, of course the BB shows not the expected band of 663 bp (for screening). The first colony shows a band at ~1.7 kb.]] | ||
| - | + | ||
None of the colonies is positive, of course the BB shows not the expected band of 663 bp (for screening). The first colony shows a band at ~1.7 kb. | None of the colonies is positive, of course the BB shows not the expected band of 663 bp (for screening). The first colony shows a band at ~1.7 kb. | ||
:=> None of the colonies harbours DelH plasmid. | :=> None of the colonies harbours DelH plasmid. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
====Colony-PCR CP.W16.B==== | ====Colony-PCR CP.W16.B==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 186: | Line 188: | ||
<br/> | <br/> | ||
[[File:Heidelberg_20130816 2log ScreenG0 ScreenG1-2a.jpg|200px|thumb|right|'''Fig.16.4''' Gel of amplified fragments (loaded 20 µL) <br> ''lane 1'' 2log ladder, ''lane 2'' screening G0, ''lane 3'' screening G1/2a ]] | [[File:Heidelberg_20130816 2log ScreenG0 ScreenG1-2a.jpg|200px|thumb|right|'''Fig.16.4''' Gel of amplified fragments (loaded 20 µL) <br> ''lane 1'' 2log ladder, ''lane 2'' screening G0, ''lane 3'' screening G1/2a ]] | ||
| - | + | ||
Gel shows expected bands. | Gel shows expected bands. | ||
:=> Both PCRs were positive. Proof of functionality of DN07 screening primer and that both fragments are the expected DelH fragments. | :=> Both PCRs were positive. Proof of functionality of DN07 screening primer and that both fragments are the expected DelH fragments. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
===Amplification of DelH G0=== | ===Amplification of DelH G0=== | ||
====Gradient PCR Conditions G0.W16.B==== | ====Gradient PCR Conditions G0.W16.B==== | ||
| Line 237: | Line 240: | ||
<br/> | <br/> | ||
[[File:Heidelberg_20130817 2kb+ 4xG0 4xG1 2kb+.png|200px|thumb|right|'''Fig.16.5''' amplified fragment of DelH (loaded 20 µL of PCR) <br>''l1:''1kb+ ladder, ''l2-5:'' G0 = complete DelH, ''l6-9:'' G1/2a, ''l10:''1kb+ ladder <br> ''l2-5:'' show not the expected band at 18Kb. ''l6-9'' show the expected band at 13.5 Kb and was cut out. But it shows also other unspecific bands.]] | [[File:Heidelberg_20130817 2kb+ 4xG0 4xG1 2kb+.png|200px|thumb|right|'''Fig.16.5''' amplified fragment of DelH (loaded 20 µL of PCR) <br>''l1:''1kb+ ladder, ''l2-5:'' G0 = complete DelH, ''l6-9:'' G1/2a, ''l10:''1kb+ ladder <br> ''l2-5:'' show not the expected band at 18Kb. ''l6-9'' show the expected band at 13.5 Kb and was cut out. But it shows also other unspecific bands.]] | ||
| - | + | ||
Gel does not show expected band. | Gel does not show expected band. | ||
:=> Further optimize PCR conditions. | :=> Further optimize PCR conditions. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
====PCR Conditions G0.W16.C==== | ====PCR Conditions G0.W16.C==== | ||
Because the amplification of DelH GO didn't work yesterday, we try it again with one picked colony and 1 µl of a glycerol stock at a constant annealing temperature of 65°C. 65°C was the temperature on which it worked the last time. | Because the amplification of DelH GO didn't work yesterday, we try it again with one picked colony and 1 µl of a glycerol stock at a constant annealing temperature of 65°C. 65°C was the temperature on which it worked the last time. | ||
| Line 286: | Line 290: | ||
<br/> | <br/> | ||
[[File:Heidelberg_20130818 G0 1kb+.png|200px|thumb|right|'''Fig.16.6''' gel of amplified DelH G0 fragment (loaded 20 µL of PCR) <br> ''l1:'' G0 amplified from picked colony, ''l2:'' G0 amplified from glycerol stock, ''l3:'' 1 kb+ ladder <br> ''l3'' shows expected band at 18 kb and was cut]] | [[File:Heidelberg_20130818 G0 1kb+.png|200px|thumb|right|'''Fig.16.6''' gel of amplified DelH G0 fragment (loaded 20 µL of PCR) <br> ''l1:'' G0 amplified from picked colony, ''l2:'' G0 amplified from glycerol stock, ''l3:'' 1 kb+ ladder <br> ''l3'' shows expected band at 18 kb and was cut]] | ||
| - | + | ||
Gel shows expected band. | Gel shows expected band. | ||
:=> It was cut and gel extracted. Also, we will run again 3 samples with the same conditions. | :=> It was cut and gel extracted. Also, we will run again 3 samples with the same conditions. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
====PCR Conditions G0.W16.D==== | ====PCR Conditions G0.W16.D==== | ||
{| class="wikitable" style="float:left; margin-right:1em" | {| class="wikitable" style="float:left; margin-right:1em" | ||
| Line 335: | Line 340: | ||
<br/> | <br/> | ||
[[File:Heidelberg_20130818 1kb+ 3xG0.png|200px|thumb|right|'''Fig.16.7''' gel of amplified DelH G0 in same conditions (loaded 20 µL of PCR) <br> ''l1:'' 1kb+ ladder, ''l2:'' G0 amplified with DN11 & HM08, ''l3:'' G0, ''l4:'' G0 <br> ''l2-4''show the expected band at 18 Kb and was cut]] | [[File:Heidelberg_20130818 1kb+ 3xG0.png|200px|thumb|right|'''Fig.16.7''' gel of amplified DelH G0 in same conditions (loaded 20 µL of PCR) <br> ''l1:'' 1kb+ ladder, ''l2:'' G0 amplified with DN11 & HM08, ''l3:'' G0, ''l4:'' G0 <br> ''l2-4''show the expected band at 18 Kb and was cut]] | ||
| - | + | ||
Gel shows expected fragments. | Gel shows expected fragments. | ||
:=> They were cut and gel extracted. | :=> They were cut and gel extracted. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
===Amplification of DelH G1/2a=== | ===Amplification of DelH G1/2a=== | ||
====Gradient PCR Conditions G1/2a.W16.B==== | ====Gradient PCR Conditions G1/2a.W16.B==== | ||
| Line 386: | Line 392: | ||
<br/> | <br/> | ||
[[File:Heidelberg_20130817 2kb+ 4xG0 4xG1 2kb+.png|200px|thumb|right|'''Fig.16.5''' amplified fragment of DelH (loaded 20 µL of PCR) <br>''l1:''1kb+ ladder, ''l2-5:'' G0 = complete DelH, ''l6-9:'' G1/2a, ''l10:''1kb+ ladder <br> ''l2-5:'' show not the expected band at 18Kb. ''l6-9'' show the expected band at 13.5 Kb and was cut out. But it shows also other unspecific bands.]] | [[File:Heidelberg_20130817 2kb+ 4xG0 4xG1 2kb+.png|200px|thumb|right|'''Fig.16.5''' amplified fragment of DelH (loaded 20 µL of PCR) <br>''l1:''1kb+ ladder, ''l2-5:'' G0 = complete DelH, ''l6-9:'' G1/2a, ''l10:''1kb+ ladder <br> ''l2-5:'' show not the expected band at 18Kb. ''l6-9'' show the expected band at 13.5 Kb and was cut out. But it shows also other unspecific bands.]] | ||
| - | + | ||
Gel shows expected band of G1/2a, but also additional uspecific bands. | Gel shows expected band of G1/2a, but also additional uspecific bands. | ||
:=> Fragment was cut and gel extracted. | :=> Fragment was cut and gel extracted. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
===Generation of Plasmid DelH 17-08=== | ===Generation of Plasmid DelH 17-08=== | ||
====Electroporation==== | ====Electroporation==== | ||
| Line 439: | Line 446: | ||
<br/> | <br/> | ||
[[File:Heidelberg_20130818_Colony_PCR_DelH.png|200px|thumb|right|'''Fig.16.8''' gel of screening minipreped colonies 1&2 from plate with 10 µl and colonies 3-12 from plate with 100 ml Gibson assembled construct with mRFP (loaded 20 µL of PCR) <br> ''l1:'' 2log ladder, ''l2:'' 1, ''l3:'' 2, ''l4:'' 3, ''l5:'' 4, ''l6:'' 5, ''l7:'' 6, ''l8:'' 7, ''l9:'' 8, ''l10:'' 9, ''l11:'' 10, ''l12:'' 11, ''l13:'' 12 <br> ''l4-13'' show expected band = positive? ]] | [[File:Heidelberg_20130818_Colony_PCR_DelH.png|200px|thumb|right|'''Fig.16.8''' gel of screening minipreped colonies 1&2 from plate with 10 µl and colonies 3-12 from plate with 100 ml Gibson assembled construct with mRFP (loaded 20 µL of PCR) <br> ''l1:'' 2log ladder, ''l2:'' 1, ''l3:'' 2, ''l4:'' 3, ''l5:'' 4, ''l6:'' 5, ''l7:'' 6, ''l8:'' 7, ''l9:'' 8, ''l10:'' 9, ''l11:'' 10, ''l12:'' 11, ''l13:'' 12 <br> ''l4-13'' show expected band = positive? ]] | ||
| - | + | ||
Majority of colonies show expected band. | Majority of colonies show expected band. | ||
:=> Grow ON, perform mini prep and test digests as well as PCRs of fragments from isolated DNA. | :=> Grow ON, perform mini prep and test digests as well as PCRs of fragments from isolated DNA. | ||
<br/> | <br/> | ||
| + | <div style="clear:both"></div> | ||
Latest revision as of 13:17, 25 October 2013
Contents |
12-08 - 18-08-13
Increasing Yield of DelH Fragments
PCR Conditions G0.G1/2a.W16.A
| Reagent | G0 | G0 | G0 | G1/2a | G1/2a |
|---|---|---|---|---|---|
| Expected length [Kb] | 18 | 18 | 18 | 13.083 | 13.083 |
| Named | G0 | G0 | G0 | 1 | 1 |
| Template | Picked colony | Picked colony | Picked colony | Picked colony | Picked colony |
| Primer fw 10 µM | 1 µl short2 | 1 µl short2 | 1 µl short2 | 1 µl HM07 | 1 µl HM07 |
| Primer rev 10 µM | 1 µl HM08 | 1 µl HM08 | 1 µl HM08 | 1 µl HM06 | 1 µl HM06 |
| Phusion Flash Ready Mix | 10 µl | 10 µl | 10 µl | 10 µl | 10 µl |
| ddH2O | 5 µl | 5 µl | 5 µl | 5 µl | 5 µl |
| DMSO | 1 µl | 1 µl | 1 µl | 1 µl | 1 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 5 |
| 30 | 98 | 1 |
| 65 | 5 | |
| 72 | 4:45 min | |
| 1 | 72 | 10 min |
| 1 | 12 | inf |
- Lid preheated at 98°C
- No hot start used
Result
Expected bands: 18 Kb (G0), 13 Kb (G1/2a)
Gel shows expected bands.
- => They were cut and gel isolated.
Generation of Plasmid DelH 11-08
Colony-PCR CP.W16.A
| Reagent | Cells + 1 µl Gibson Assembly | Cells + 14 µl Gibson Assembly | Cells + 14 µl Gibson Assembly (ONLY Backbone) | Cells + 14 µl Gibson Assembly (ONLY Backbone) |
|---|---|---|---|---|
| Expected length [bp] | 663 | 663 | - | - |
| Named | D1 | D14 | B14(1) | B14(2) |
| Primer fw 10 µM | 2 µl VF2 | 2 µl VF2 | 2 µl VF2 | 2 µl VF2 |
| Primer rev 10 µM | 2 µl Screen_delH_rev | 2 µl Screen_delH_rev | 2 µl Screen_delH_rev | 2 µl Screen_delH_rev |
| Dream-Taq Polymerase (2x) | 10 µl | 10 µl | 10 µl | 10 µl |
| ddH2O | 6 µl | 6 µl | 6 µl | 6 µl |
| Cycles | Temperature DelH-G0 [°C] | Time [s] |
|---|---|---|
| 1 | 95 | 120 |
| 12 | 95 | 60 |
| 68 (touchdown -0.5°C) | 30 | |
| 72 | 45 | |
| 18 | 95 | 60 |
| 65 (touchdown -0.5°C) | 30 | |
| 72 | 45 | |
| 1 | 12 | inf |
Result
Expected band: 663 bp
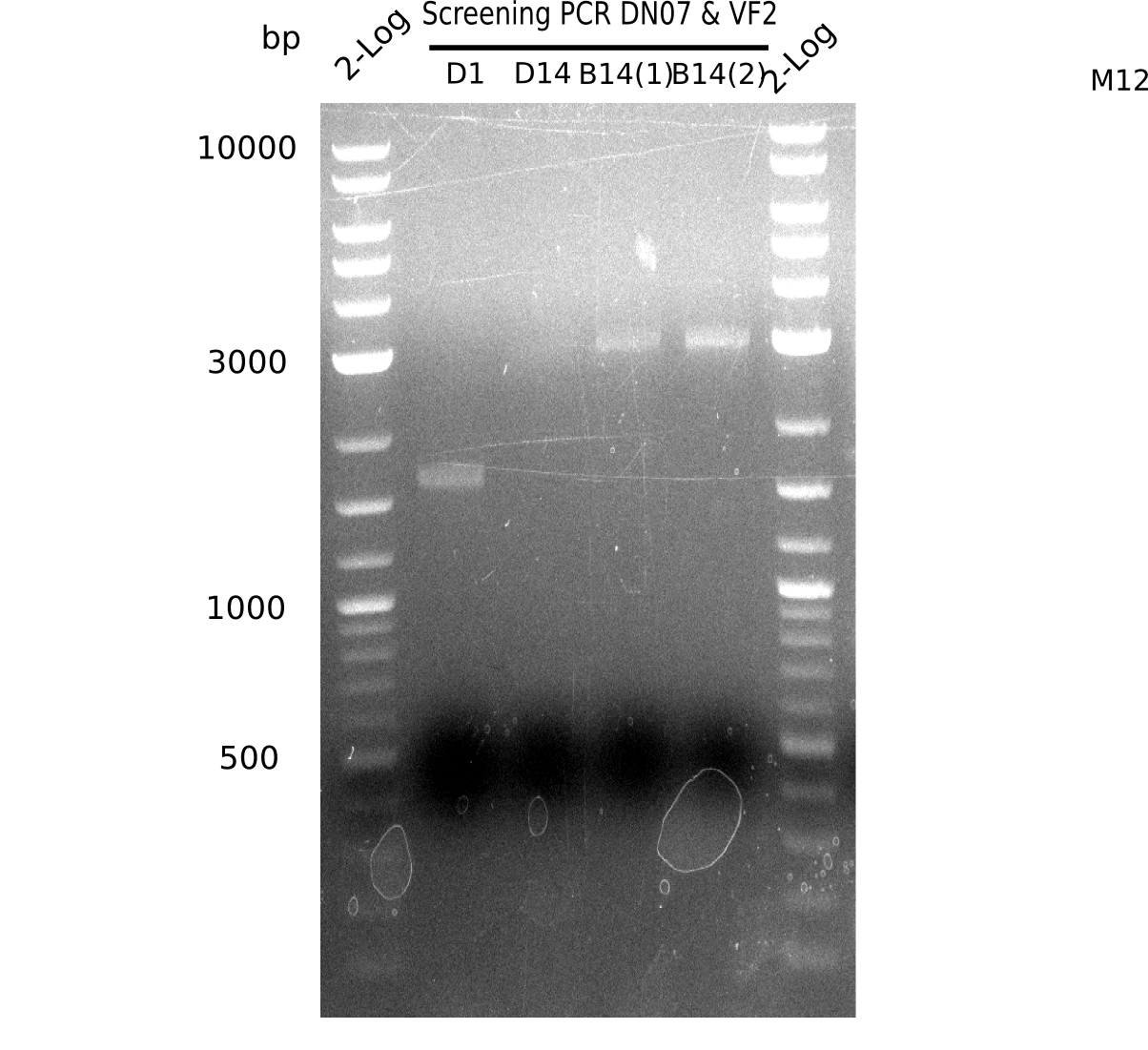
l1: 2log ladder, l2: colony PCR of 1 µl Gibson transformed,l3: colony PCR of 14 µl Gibson transformed,l4-5: 2 colony PCRs of BB - 14 µl Gibson transformed, l6: 2log ladder
none of the colonies is positive, of course the BB shows not the expected band of 663 bp (for screening). The first colony shows a band at ~1.7 kb.
None of the colonies is positive, of course the BB shows not the expected band of 663 bp (for screening). The first colony shows a band at ~1.7 kb.
- => None of the colonies harbours DelH plasmid.
Colony-PCR CP.W16.B
| Reagent | Cells + 10 µl Gibson Assembly pink | Cells + 10 µl Gibson Assembly pink | Cells + 10 µl Gibson Assembly white | Cells + 10 µl Gibson Assembly white |
|---|---|---|---|---|
| Expected length [bp] | 663 | 663 | 663 | 663 |
| Named | A | B | C | D |
| Primer fw 10 µM | 2 µl VF2 | 2 µl VF2 | 2 µl VF2 | 2 µl VF2 |
| Primer rev 10 µM | 2 µl Screen_delH_rev | 2 µl Screen_delH_rev | 2 µl Screen_delH_rev | 2 µl Screen_delH_rev |
| Dream-Taq Polymerase (2x) | 10 µl | 10 µl | 10 µl | 10 µl |
| ddH2O | 6 µl | 6 µl | 6 µl | 6 µl |
| Cycles | Temperature DelH-G0 [°C] | Time [s] |
|---|---|---|
| 1 | 95 | 120 |
| 12 | 95 | 60 |
| 68 (touchdown -0.5°C) | 30 | |
| 72 | 45 | |
| 18 | 95 | 60 |
| 65 (touchdown -0.5 °C) | 30 | |
| 72 | 45 | |
| 1 | 12 | inf |
Result
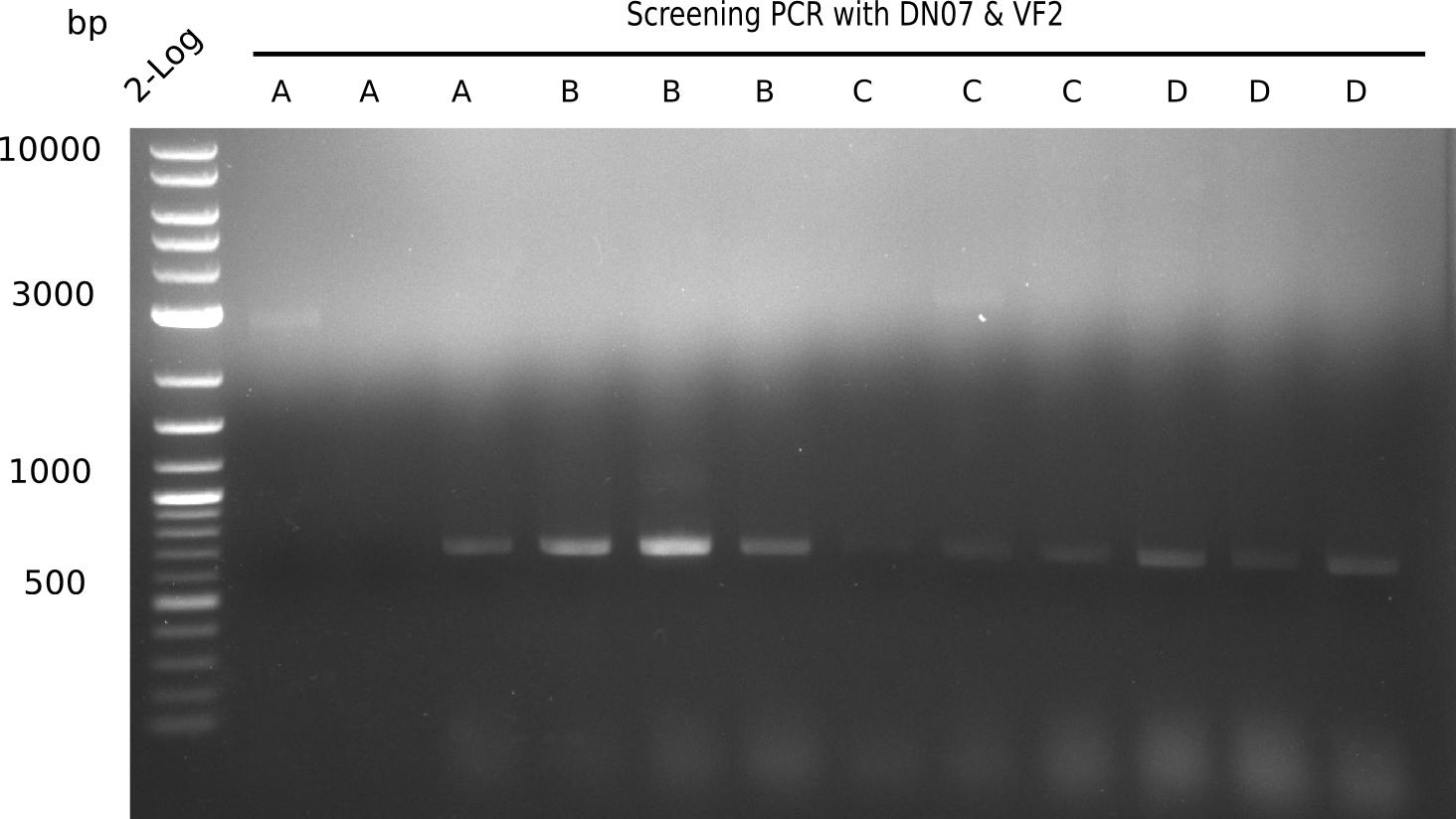
Expected band: 663 bp
Testing of Screening Primers
To test the reverse screening primer DN07 we amplify with the DreamTaqPolymerase and DN11 primer. As template, we use the complete DelH (18 Kb) and the first 3/4 of DelH (14 Kb).
| Reagent | 1 µl G0 of 23-07 | 1 µl G1/2a of 23-07 |
|---|---|---|
| Expected length [bp] | 323 | 323 |
| Named | 0 | 1 |
| Primer fw 10 µM | 2 µl DN11 | 2 µl DN11 |
| Primer rev 10 µM | 2 µl Screen_delH_rev | 2 µl Screen_delH_rev |
| Dream-Taq Polymerase (2x) | 10 µl | 10 µl |
| ddH2O | 5 µl | 5 µl |
| Cycles | Temperature DelH-G0 [°C] | Time [s] |
|---|---|---|
| 1 | 95 | 120 |
| 12 | 95 | 60 |
| 68 (touchdown -0.5°C) | 30 | |
| 72 | 30 | |
| 18 | 95 | 60 |
| 65 (touchdown -0.5°C) | 30 | |
| 72 | 30 | |
| 1 | 12 | inf |
Result
Expected length= 323 bp for positive clones
Gel shows expected bands.
- => Both PCRs were positive. Proof of functionality of DN07 screening primer and that both fragments are the expected DelH fragments.
Amplification of DelH G0
Gradient PCR Conditions G0.W16.B
To improve the PCR of the fragment with the length of 18 Kb, we run a gradient PCR between 68°C and 65°C.
| Reagent | G0 | G0 | G0 | G0 |
|---|---|---|---|---|
| Expected length [Kb] | 18 | 18 | 18 | 18 |
| Named | G0 1 | G0 2 | G0 3 | G0 4 |
| Template | Picked colony | Picked colony | Picked colony | Picked colony |
| Primer fw 10 µM | 2 µl short2 | 2 µl short2 | 2 µl short2 | 2 µl short2 |
| Primer rev 10 µM | 2 µl HM08 | 2 µl HM08 | 2 µl HM08 | 2 µl HM08 |
| Phusion Flash Ready Mix | 10 µl | 10 µl | 10 µl | 10 µl |
| ddH2O | 5 µl | 5 µl | 5 µl | 5 µl |
| DMSO | 1 µl | 1 µl | 1 µl | 1 µl |
| Cycles | Temperature DelH-G0 [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 5 |
| 30 | 98 | 1 |
| 68 - 65 | 5 | |
| 72 | 4:45 min | |
| 1 | 72 | 10 min |
| 1 | 12 | inf |
- Lid preheated at 98°C
- No hot start
Result
Expected band: 18 Kb
Gel does not show expected band.
- => Further optimize PCR conditions.
PCR Conditions G0.W16.C
Because the amplification of DelH GO didn't work yesterday, we try it again with one picked colony and 1 µl of a glycerol stock at a constant annealing temperature of 65°C. 65°C was the temperature on which it worked the last time.
| Reagent | G0 | G0 |
|---|---|---|
| Expected length [Kb] | 18 | 18 |
| Named | G0 1 | G0 2 |
| Template | Picked colony SPH1 | 1 µl of glycerol stock SPH1 |
| Primer fw 10 µM | 2 µl short2 | 2 µl short2 |
| Primer rev 10 µM | 2 µl HM08 | 2 µl HM08 |
| Phusion Flash Ready Mix | 10 µl | 10 µl |
| ddH2O | 5 µl | 4 µl |
| DMSO | 1 µl | 1 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 5 |
| 30 | 98 | 1 |
| 65 | 5 | |
| 72 | 4:45 min | |
| 1 | 72 | 10 min |
| 1 | 12 | inf |
- Lid preheated at 98°C
- No hot start
Result
Expected band: 18 Kb
Gel shows expected band.
- => It was cut and gel extracted. Also, we will run again 3 samples with the same conditions.
PCR Conditions G0.W16.D
| Reagent | G0 | G0 | G0 |
|---|---|---|---|
| Expected length [Kb] | 18 | 18 | 18 |
| Named | G0 1 | G0 2 | G0 2 |
| Template | 1 µl of glycerol stock SPH1 | 1 µl of glycerol stock SPH1 | 1 µl of glycerol stock SPH1 |
| Primer fw 10 µM | 2 µl short2 | 2 µl short2 | 2 µl short2 |
| Primer rev 10 µM | 2 µl HM08 | 2 µl HM08 | 2 µl HM08 |
| Phusion Flash Ready Mix | 10 µl | 10 µl | 10 µl |
| ddH2O | 4 µl | 4 µl | 4 µl |
| DMSO | 1 µl | 1 µl | 1 µl |
| Cycles | Temperature DelH-G0 [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 5 |
| 30 | 98 | 1 |
| 65 | 5 | |
| 72 | 4:45 min | |
| 1 | 72 | 10 min |
| 1 | 12 | inf |
- Lid preheated at 98°C
- No hot start
Result
Expected band: 18 Kb
Gel shows expected fragments.
- => They were cut and gel extracted.
Amplification of DelH G1/2a
Gradient PCR Conditions G1/2a.W16.B
To improve the PCR of the fragment with the length of 18 Kb, we run a gradient PCR between 68°C and 65°C.
| Reagent | G1/2a | G1/2a | G1/2a | G1/2a |
|---|---|---|---|---|
| Expected length [Kb] | 13.083 | 13.083 | 13.083 | 13.083 |
| Named | 1 1 | 1 2 | 1 3 | 1 4 |
| Template | Picked colony | Picked colony | Picked colony | Picked colony |
| Primer fw 10 µM | 2 µl short2 | 2 µl short2 | 2 µl short2 | 2 µl short2 |
| Primer rev 10 µM | 2 µl HM06 | 2 µl HM06 | 2 µl HM06 | 2 µl HM06 |
| Phusion Flash Ready Mix | 10 µl | 10 µl | 10 µl | 10 µl |
| ddH2O | 5 µl | 5 µl | 5 µl | 5 µl |
| DMSO | 1 µl | 1 µl | 1 µl | 1 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 98 | 5 |
| 30 | 98 | 1 |
| 68 - 65 | 5 | |
| 72 | 4:45 min | |
| 1 | 72 | 10 min |
| 1 | 12 | inf |
- Lid preheated at 98°C
- No hot start
Result
Expected band: 13 Kb
Gel shows expected band of G1/2a, but also additional uspecific bands.
- => Fragment was cut and gel extracted.
Generation of Plasmid DelH 17-08
Electroporation
Remaining Gibson mix from 01.08 was electroporated.
Colony-PCR CP.W16.B
5 colony per PCR tube (but in different LB Amp Eppis)
| Reagent | Electroporation from 17-08 10 µl | Electroporation from 17-08 100 µl |
|---|---|---|
| Expected length [bp] | 663 | 663 |
| Named | 1 - 2 | 3 - 12 |
| Template | 2 colonies of 10 µl | 10 colonies of 100 µl |
| Primer fw 10 µM | 1 µl VF2 | 1 µl VF2 |
| Primer rev 10 µM | 1 µl Screen_delH_rev | 1 µl Screen_delH_rev |
| Dream-Taq Polymerase (2x) | 10 µl | 10 µl |
| ddH2O | 8 µl | 8 µl |
| Cycles | Temperature [°C] | Time [s] |
|---|---|---|
| 1 | 95 | 120 |
| 12 | 95 | 60 |
| 68 (touchdown -0.5°C) | 30 | |
| 72 | 45 | |
| 18 | 95 | 60 |
| 65 (touchdown -0.5°C) | 30 | |
| 72 | 45 | |
| 1 | 12 | inf |
Result
Expected band: 663 bp
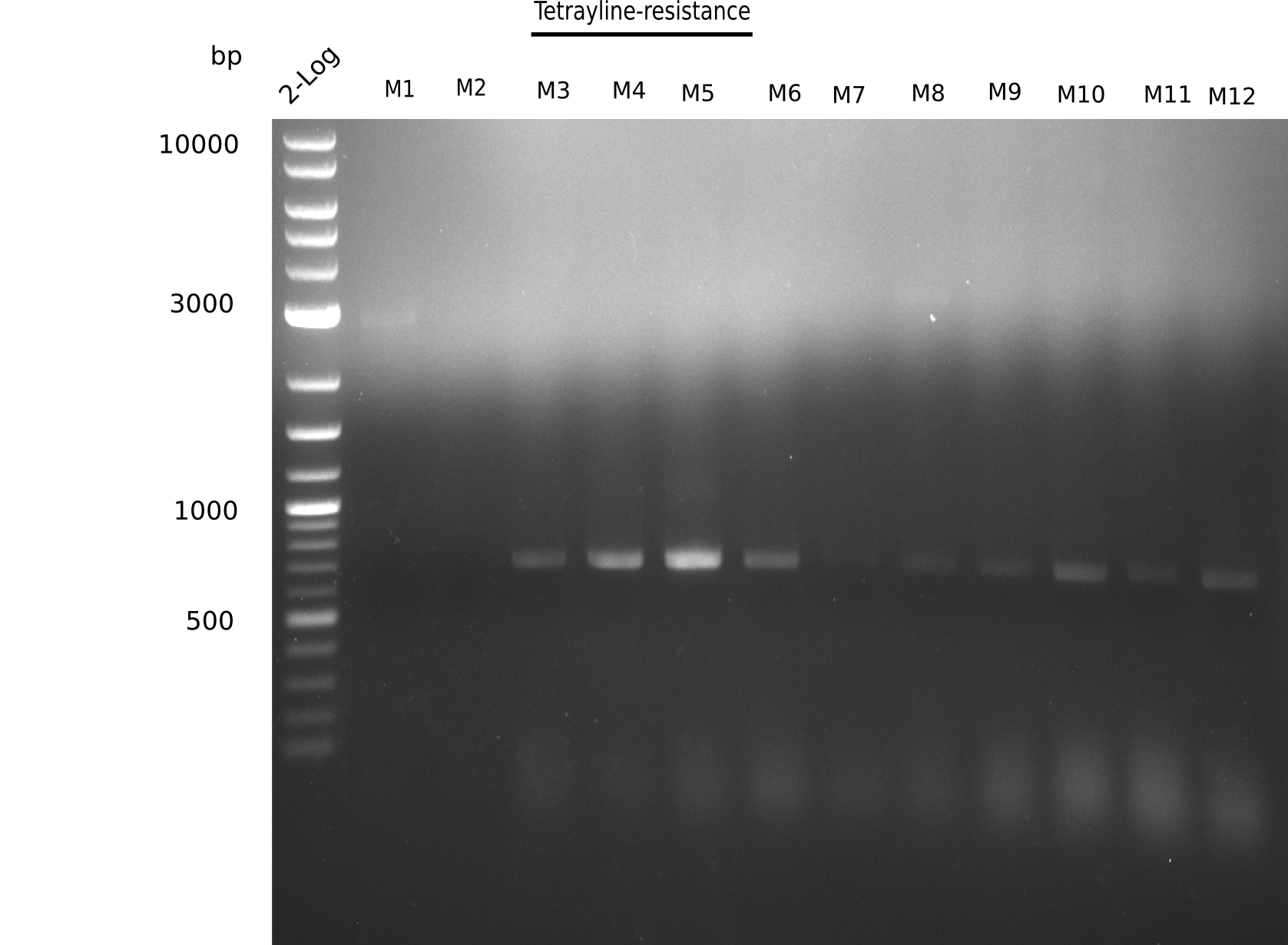
l1: 2log ladder, l2: 1, l3: 2, l4: 3, l5: 4, l6: 5, l7: 6, l8: 7, l9: 8, l10: 9, l11: 10, l12: 11, l13: 12
l4-13 show expected band = positive?
Majority of colonies show expected band.
- => Grow ON, perform mini prep and test digests as well as PCRs of fragments from isolated DNA.
 "
"