Team:UC Davis/Data
From 2013.igem.org
(Difference between revisions)
Alferreiro (Talk | contribs) |
|||
| Line 29: | Line 29: | ||
<div class="floatboxwide"> | <div class="floatboxwide"> | ||
| + | <h1>Fluorescence is modulated by theophylline concentrations</h1> | ||
| + | <br>We subjected our testing construct to the induction condition of 10 mM aTc, which would result in constitutive and maximal expression of GFP given no repression. We also subjected the construct to the induction condition of 0.1% arabinose, which would produce a nominal level of RiboTALe transcript. We varied the only the concentration of theophylline, over a range of 0 mM to 10 mM. Thus, difference in fluorescence between induction conditions would be due only to the RiboTALe repression activity. We measured the fluorescence of our construct in E. Coli MG1655Z1 over a course of 12 hrs using the Tecan 200Pro microplate reader. Please refer to the <a href="https://2013.igem.org/Team:UC_Davis/Protocols">Protocols</a></hi> page for details on our culture preparation and Tecan testing parameters. </br> | ||
| + | <center><img src="http://31.media.tumblr.com/tumblr_llc8j1LWBF1qzamioo1_500.jpg"></center> | ||
| + | <br>The image above illustrates that GFP fluorescence is inversely related to theophylline concentrations, indicating that the TAL repressor is in fact being translated at rates corresponding to the theophylline induction levels, and effectively binding to its target site. At maximal theophylline concentrations, the expression of GFP is reduced (X) fold.</br> | ||
| + | <br>Next, we investigated what difference in system response we could achieve by altering the binding affinity of the TAL repressor protein.</br> | ||
| + | <h1>Varying the binding affinity of the TAL repressor allows for system tunability</h1> | ||
<h1 id="widget">KO3D</h1> | <h1 id="widget">KO3D</h1> | ||
<div id="mutantwidget" class="floatbox"> | <div id="mutantwidget" class="floatbox"> | ||
Revision as of 19:21, 24 September 2013

Proof of Concept: Our Testing Construct
To characterize the behavior of a RiboTALe device, we acquired cells containing the sequences for TAL repressors from the Tagkopoulos Lab at UC Davis, with which we have worked closely. We placed the TAL repressors downstream of theophylline-responsive riboswitches, the sequences of which were taken from the studies A high-throughput screen for synthetic riboswitches reveals mechanistic insights into their function (Lynch et al, Chem Biol. 2007 Feb;14(2):173-84.), and Synthetic Riboswitches That Induce Gene Expression in Diverse Bacterial Species (Topp et al, Appl. Environ. Microbiol. December 2010 vol. 76 no. 23 7881-7884). The riboswitch-TALe sequences were placed under the regulation of a pBAD promoter.
We inserted previously engineered TALe binding sites corresponding to the TAL repressors used in our characterization experiments upstream of a reporter, GFP. This target sequence was placed under the regulation of a pTET promoter.

We tested our construct by subjecting the pBAD promoter, the theophylline riboswitch, and the pTET promoter to a range of induction levels with arabinose, theophylline, and aTc, respectively. It was expected that at low levels of arabinose and theophylline, but at high levels of aTc, GFP expression would be maximal due to the very low production of TAL repressor protein. On the other hand, at high levels of arabinose and theophylline it was expected that fluorescence levels would be greatly reduced due the higher rate of TAL repressor production. We also expected to see many instances of neither total GFP expression or total GFP repression, depending on the relative states of induction of the pBAD promoter, the theophylline riboswitch, and the pTET promoter.
Data Links
- TAL repressor modularity
- Riboswitch modularity
- KO3D
- TAL repressor modularity
- Riboswitch modularity
- KO3D
Fluorescence is modulated by theophylline concentrations
We subjected our testing construct to the induction condition of 10 mM aTc, which would result in constitutive and maximal expression of GFP given no repression. We also subjected the construct to the induction condition of 0.1% arabinose, which would produce a nominal level of RiboTALe transcript. We varied the only the concentration of theophylline, over a range of 0 mM to 10 mM. Thus, difference in fluorescence between induction conditions would be due only to the RiboTALe repression activity. We measured the fluorescence of our construct in E. Coli MG1655Z1 over a course of 12 hrs using the Tecan 200Pro microplate reader. Please refer to the Protocols page for details on our culture preparation and Tecan testing parameters.
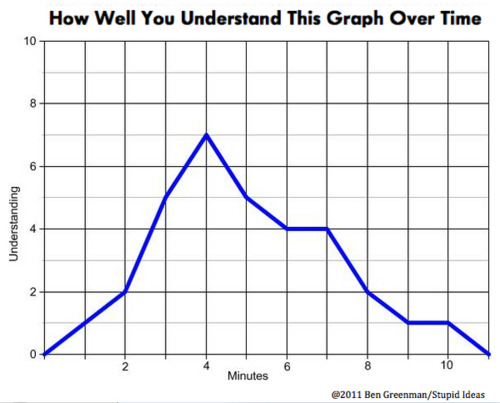
The image above illustrates that GFP fluorescence is inversely related to theophylline concentrations, indicating that the TAL repressor is in fact being translated at rates corresponding to the theophylline induction levels, and effectively binding to its target site. At maximal theophylline concentrations, the expression of GFP is reduced (X) fold.
Next, we investigated what difference in system response we could achieve by altering the binding affinity of the TAL repressor protein.
Varying the binding affinity of the TAL repressor allows for system tunability
KO3D
0 ng/mL aTc
caatacgcaaaccgcctctccccgcgcgttggccgattcattaatgcagctggcacgacaggtttcccgactggaaagcgggcagtgagcgcaacgcaattaatgtgagttagctcactcattaggcaccccaggctttacactttatgcttccggctcgtatgttgtgtggaattgtgagcggataacaatttcacaca 0, 1, 2, 5, 10 599, 647, 690, 684, 494 0.0031579, 0.0052541, 0.0059555, 0.0069092, 0.0041919 6 0, .01, .1, .25, .5, 1 0, 1, 2, 5, 10 688, 634, 630, 567, 613, 599, 697, 690, 672, 686, 671, 647, 691, 709, 699, 689, 703, 690, 517, 719, 700, 699, 717, 684, 448, 510, 493, 500, 512, 494100 ng/mL aTc
caatacgcaaaccgcctctccccgcgcgttggccgattcattaatgcagctggcacgacaggtttcccgactggaaagcgggcagtgagcgcaacgcaattaatgtgagttagctcactcattaggcaccccaggctttacactttatgcttccggctcgtatgttgtgtggaattgtgagcggataacaatttcacaca 0, 1, 2, 5, 10 19281, 14779, 18481, 16665, 7100 .017132, .016563, .010138, .0083723, .0036696, .0013295, .00029933 0, .01, .1, .25, .5, 1 0, 1, 2, 5, 10 27676, 27805, 27124, 20736, 22057, 19281, 25713, 27355, 25079, 18345, 15425, 14779, 25355, 26322, 26972, 17362, 17072, 18481, 18771, 19210, 21053, 10468, 11105, 16665, 5461, 6111, 7360, 2909, 2919, 7100

Project BackgroundLearn about how we combine riboswitches and TAL's into robust orthogonal mechanisms for inducible repression. |
ResultsCheck out the results of our experiments. |

Human PracticesTake a look at how we designed a new database for better raw data characterization of Biobricks. |

Judging CriteriaHere's the criteria that we met for this year's team. |
 "
"