Team:Yale/Project Bioassay
From 2013.igem.org
(Difference between revisions)
(→Aims for the Project) |
|||
| (12 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
<div id="header">{{Template:Team:Yale2013/Templates/Header}}</div> | <div id="header">{{Template:Team:Yale2013/Templates/Header}}</div> | ||
| - | + | <!-- Project Submenu adapted from cssmenumaker --> | |
| - | + | <html> | |
| - | + | <style type = "text/css"> | |
| - | + | #cssmenu1 ul, | |
| - | + | #cssmenu1 li, | |
| - | + | #cssmenu1 span, | |
| - | + | #cssmenu1 a { | |
| - | + | margin: 0; | |
| - | + | padding: 0; | |
| - | + | position: relative; | |
| - | + | } | |
| - | + | #cssmenu1 { | |
| - | = | + | height: 39px; |
| - | + | border-radius: 0px 0px 4px 4px; | |
| - | + | -moz-border-radius: 0px 0px 4px 4px; | |
| - | + | -webkit-border-radius: 0px 0px 4px 4px; | |
| - | + | background: #fefefe; | |
| - | + | background: -moz-linear-gradient(top, #fefefe 0%, #eee9f0 100%); | |
| + | background: -webkit-gradient(linear, left top, left bottom, color-stop(0%, #fefefe), color-stop(100%, #eee9f0)); | ||
| + | background: -webkit-linear-gradient(top, #fefefe 0%, #eee9f0 100%); | ||
| + | background: -o-linear-gradient(top, #fefefe 0%, #eee9f0 100%); | ||
| + | background: -ms-linear-gradient(top, #fefefe 0%, #eee9f0 100%); | ||
| + | background: linear-gradient(top, #fefefe 0%, #eee9f0 100%); | ||
| + | border-bottom: 2px solid #5763f0; | ||
| + | width: auto; | ||
| + | } | ||
| + | #cssmenu1:after, | ||
| + | #cssmenu1 ul:after { | ||
| + | content: ''; | ||
| + | display: block; | ||
| + | clear: both; | ||
| + | } | ||
| + | #cssmenu1 a { | ||
| + | background: #fefefe; | ||
| + | background: -moz-linear-gradient(top, #fefefe 0%, #ececec 100%); | ||
| + | background: -webkit-gradient(linear, left top, left bottom, color-stop(0%, #fefefe), color-stop(100%, #ececec)); | ||
| + | background: -webkit-linear-gradient(top, #fefefe 0%, #ececec 100%); | ||
| + | background: -o-linear-gradient(top, #fefefe 0%, #ececec 100%); | ||
| + | background: -ms-linear-gradient(top, #fefefe 0%, #ececec 100%); | ||
| + | background: linear-gradient(top, #fefefe 0%, #ececec 100%); | ||
| + | color: #000; | ||
| + | display: inline-block; | ||
| + | font-family: Helvetica, Arial, Verdana, sans-serif; | ||
| + | font-size: 12px; | ||
| + | line-height: 39px; | ||
| + | padding: 0 10px; | ||
| + | text-decoration: none; | ||
| + | } | ||
| + | #cssmenu1 ul { | ||
| + | list-style: none; | ||
| + | } | ||
| + | #cssmenu1 > ul { | ||
| + | float: left; | ||
| + | } | ||
| + | #cssmenu1 > ul > li { | ||
| + | float: left; | ||
| + | } | ||
| + | #cssmenu1 > ul > li > a { | ||
| + | color: #000; | ||
| + | font-size: 12px; | ||
| + | } | ||
| + | #cssmenu1 > ul > li:hover:after { | ||
| + | content: ''; | ||
| + | display: block; | ||
| + | width: 0; | ||
| + | height: 0; | ||
| + | position: absolute; | ||
| + | left: 50%; | ||
| + | bottom: 0; | ||
| + | border-left: 10px solid transparent; | ||
| + | border-right: 10px solid transparent; | ||
| + | border-bottom: 10px solid #5763f0; | ||
| + | margin-left: -10px; | ||
| + | } | ||
| + | #cssmenu1 > ul > li:first-child > a { | ||
| + | border-radius: 0px 0px 4px 4px; | ||
| + | -moz-border-radius: 0px 0px 4px 4px; | ||
| + | -webkit-border-radius: 0px 0px 4px 4px; | ||
| + | } | ||
| + | #cssmenu1 > ul > li.active:after { | ||
| + | content: ''; | ||
| + | display: block; | ||
| + | width: 0; | ||
| + | height: 0; | ||
| + | position: absolute; | ||
| + | left: 50%; | ||
| + | bottom: 0; | ||
| + | border-left: 10px solid transparent; | ||
| + | border-right: 10px solid transparent; | ||
| + | border-bottom: 10px solid #5763f0; | ||
| + | margin-left: -10px; | ||
| + | } | ||
| + | #cssmenu1 > ul > li.active > a { | ||
| + | -moz-box-shadow: inset 0 0 2px rgba(0, 0, 0, 0.1); | ||
| + | -webkit-box-shadow: inset 0 0 2px rgba(0, 0, 0, 0.1); | ||
| + | box-shadow: inset 0 0 2px rgba(0, 0, 0, 0.1); | ||
| + | background: #ececec; | ||
| + | background: -moz-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: -webkit-gradient(linear, left top, left bottom, color-stop(0%, #ececec), color-stop(100%, #ffeeff ef)); | ||
| + | background: -webkit-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: -o-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: -ms-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | } | ||
| + | #cssmenu1 > ul > li:hover > a { | ||
| + | background: #ececec; | ||
| + | background: -moz-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: -webkit-gradient(linear, left top, left bottom, color-stop(0%, #ececec), color-stop(100%, #ffeeff ef)); | ||
| + | background: -webkit-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: -o-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: -ms-linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | background: linear-gradient(top, #ececec 0%, #ffeeff ef 100%); | ||
| + | -moz-box-shadow: inset 0 0 2px rgba(0, 0, 0, 0.1); | ||
| + | -webkit-box-shadow: inset 0 0 2px rgba(0, 0, 0, 0.1); | ||
| + | box-shadow: inset 0 0 2px rgba(0, 0, 0, 0.1); | ||
| + | } | ||
| + | #cssmenu1 .has-sub { | ||
| + | z-index: 1; | ||
| + | } | ||
| + | #cssmenu1 .has-sub:hover > ul { | ||
| + | display: block; | ||
| + | } | ||
| + | #cssmenu1 .has-sub ul { | ||
| + | display: none; | ||
| + | position: absolute; | ||
| + | width: 200px; | ||
| + | top: 100%; | ||
| + | left: 0; | ||
| + | } | ||
| + | #cssmenu1 .has-sub ul li { | ||
| + | *margin-bottom: -1px; | ||
| + | } | ||
| + | #cssmenu1 .has-sub ul li a { | ||
| + | background: #5763f0; | ||
| + | border-bottom: 1px dotted #868ef4; | ||
| + | filter: none; | ||
| + | font-size: 11px; | ||
| + | display: block; | ||
| + | line-height: 120%; | ||
| + | padding: 10px; | ||
| + | color: #ffffff; | ||
| + | } | ||
| + | #cssmenu1 .has-sub ul li:hover a { | ||
| + | background: #2838ec; | ||
| + | } | ||
| + | #cssmenu1 .has-sub .has-sub:hover > ul { | ||
| + | display: block; | ||
| + | } | ||
| + | #cssmenu1 .has-sub .has-sub ul { | ||
| + | display: none; | ||
| + | position: absolute; | ||
| + | left: 100%; | ||
| + | top: 0; | ||
| + | } | ||
| + | #cssmenu1 .has-sub .has-sub ul li a { | ||
| + | background: #2838ec; | ||
| + | border-bottom: 1px dotted #868ef4; | ||
| + | } | ||
| + | #cssmenu1 .has-sub .has-sub ul li a:hover { | ||
| + | background: #1525e6; | ||
| + | } | ||
| + | </style> | ||
| + | <div id="cssmenu1" style="width:830px; margin:0 auto;"> | ||
| + | <ul> | ||
| + | <li><a href='https://2013.igem.org/Team:Yale/Project_Overview'><span>Project Overview</span></a></li> | ||
| + | <li><a href='https://2013.igem.org/Team:Yale/Project_Validate'><span>Validate PLA Synthesis</span></a></li> | ||
| + | <li class='active'><a href='https://2013.igem.org/Team:Yale/Project_Bioassay'><span>Develop Bioassay</span></a></li> | ||
| + | <li><a href='https://2013.igem.org/Team:Yale/Project_MAGE'><span>Apply Mage</span></a></li> | ||
| + | <li><a href='https://2013.igem.org/Team:Yale/Project_Export'><span>Introduce Export System</span></a></li> | ||
| + | <li><a href='https://2013.igem.org/Team:Yale/Project_Bioplastic'><span>Make a Bioplastic</span></a></li> | ||
| + | <li class='last'><a href='https://2013.igem.org/Team:Yale/Project_Collaboration'><span>Collaboration</span></a></li> | ||
| + | </ul> | ||
| + | </div> | ||
| + | </html> | ||
== Develop bioassay to screen PLA production == | == Develop bioassay to screen PLA production == | ||
*We needed a way to detect the PLA once we produced it using the heterologous enzymes | *We needed a way to detect the PLA once we produced it using the heterologous enzymes | ||
**We decided to use the fluorescent dye Nile red, a intercellular lipid strain | **We decided to use the fluorescent dye Nile red, a intercellular lipid strain | ||
| - | **Nile red does not affect the growth of bacteria, and its fluorescence is quenched in water <br> | + | **Nile red does not affect the growth of bacteria, and its fluorescence is quenched in water |
| - | <center>[[File:Nile_red_PHB_stain.jpg|200px]]</center><br><br> | + | <br> |
| + | <center>[[File:Nile_red_PHB_stain.jpg|200px]]<br> | ||
| + | This is a figure from Spiekermann et al. 1996 demonstrating Nile red staining of both PHB+ ''E. coli'' and PHB negative ''E. coli'' </center><br><br> | ||
===Positive Control=== | ===Positive Control=== | ||
*We used the 2012 Tokyo Tech Biobrick (BBa_K934001) as a positive control to attempt to detect Nile red fluorescence on our plate reader (ex. 530nm, em. 590nm) | *We used the 2012 Tokyo Tech Biobrick (BBa_K934001) as a positive control to attempt to detect Nile red fluorescence on our plate reader (ex. 530nm, em. 590nm) | ||
| - | **This plasmid had | + | **This plasmid had three enzymes which together give ''E. coli'' that ability to synthesize P(3HB) a similar plastic PLA |
**Cells were grown for 24 hours in the presence of Nile red. The cells were washed and resuspended in PBS. | **Cells were grown for 24 hours in the presence of Nile red. The cells were washed and resuspended in PBS. | ||
<center>[[File:Tokyo Tech plasmid.jpg|400px]]</center><br><br> | <center>[[File:Tokyo Tech plasmid.jpg|400px]]</center><br><br> | ||
===Our Construct=== | ===Our Construct=== | ||
| - | *We then proceeded to test | + | *We then proceeded to test our plasmid in the plate reader. |
| - | **Cells were grown for 24 hours with both enzymes induced and in the presence of Nile red. The cells were washed and | + | **Cells were grown for 24 hours with both enzymes induced and in the presence of Nile red. The cells were washed and re-suspended in PBS. The readings were normalized for optical density. |
<center>[[File:EcNR2 LB x20 wiki.jpg|400px]]</center><br><br> | <center>[[File:EcNR2 LB x20 wiki.jpg|400px]]</center><br><br> | ||
== FACS Sorting == | == FACS Sorting == | ||
| - | *In order to quickly screen the large diversity | + | *In order to quickly screen the large diversity created using MAGE, we employed Fluorescence-activated cell sorting (FACS) |
**FACS could sort cells based on the Nile red fluorescence, thus indicating those cells that have produced larger quantities of PLA | **FACS could sort cells based on the Nile red fluorescence, thus indicating those cells that have produced larger quantities of PLA | ||
<center>[[File:FACS.jpg|400px]]</center><br><br> | <center>[[File:FACS.jpg|400px]]</center><br><br> | ||
| Line 45: | Line 203: | ||
*Nile red has an emission maximum at 598nm when bound to p(3HB) granules according to Spiekermann et al. 1998 | *Nile red has an emission maximum at 598nm when bound to p(3HB) granules according to Spiekermann et al. 1998 | ||
*Thus we decided to use PE-Texas Red which has an emission maximum at 615nm in order pick the best colonies. | *Thus we decided to use PE-Texas Red which has an emission maximum at 615nm in order pick the best colonies. | ||
| + | **PE-Texas Red is a covalently linked fluorochrome used as the detector. The FACS machine measure the emission of this fluorochrome because Nile Red is not a common fluorochrome that a FACS machine would have detector specific to Nile red. | ||
{| | {| | ||
|- | |- | ||
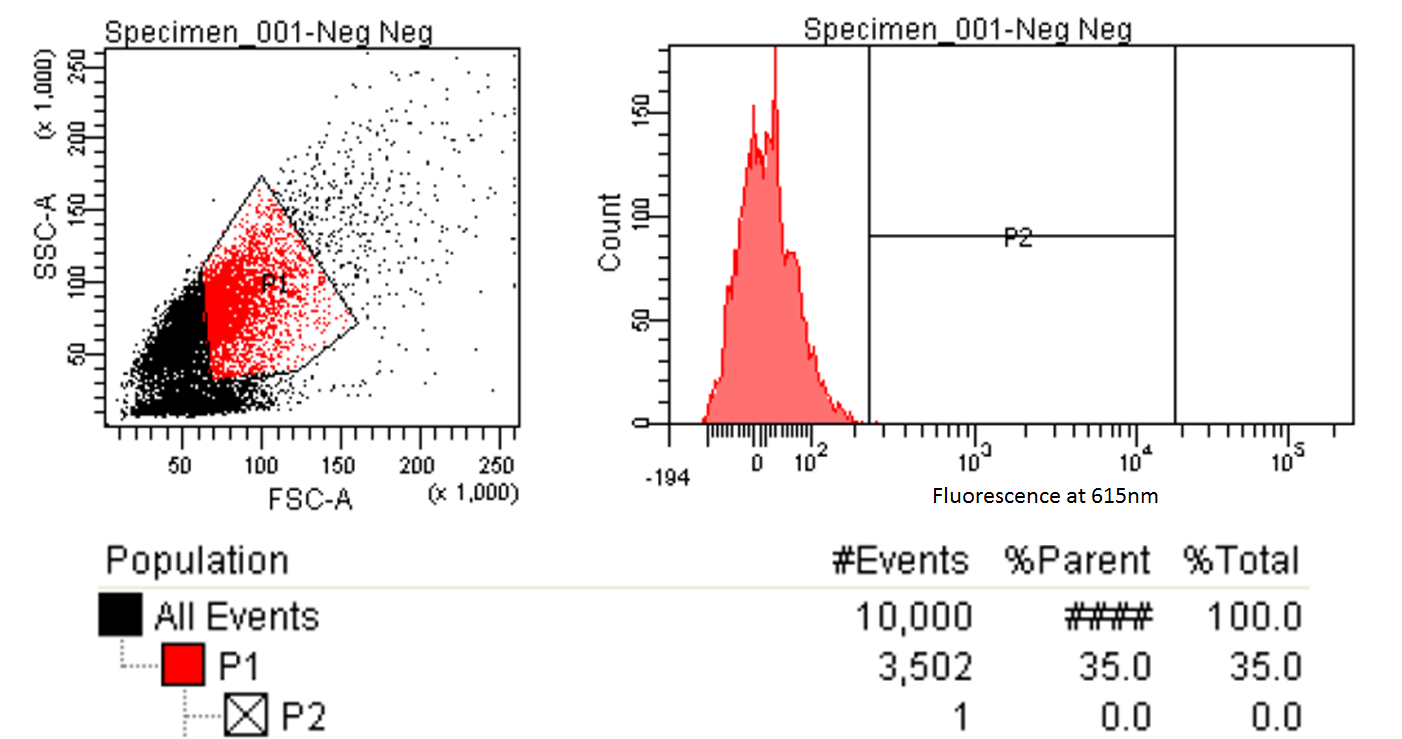
|style="padding-left: 20px; padding-right: 20px;"|[[File:FACScontrol.png|600px]] | |style="padding-left: 20px; padding-right: 20px;"|[[File:FACScontrol.png|600px]] | ||
| - | |The left graph shows the distribution of cells entering the FACS. A gate is made to only focus on those which are relatively round and average size (the enclosed red shape). The right graph is the fluorescence of those selected cells. A gate is made and 1/10,000 of the cells are within this gate. Ideally we would want no control cells but this one cells | + | |style="text-align:justify; padding: 5px;"|The left graph shows the distribution of cells entering the FACS. A gate is made to only focus on those which are relatively round and average size (the enclosed red shape). The right graph is the fluorescence of those selected cells. A gate (labeled P2) is made and 1/10,000 of the cells are within this gate. Ideally we would want the machine to select only those with fluorescence above the control cells (and thus no control cells in the P2 gate) but we believe this one cells is an outlier due to its abnormally high levels of fluorescence. |
|} | |} | ||
<br><br> | <br><br> | ||
| Line 54: | Line 213: | ||
|- | |- | ||
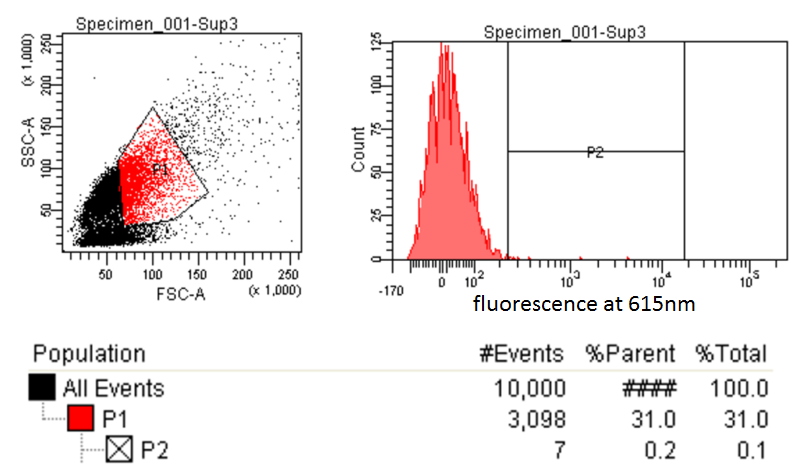
|style="padding-left: 20px; padding-right: 20px;"|[[File:FACSsample.png|600px]] | |style="padding-left: 20px; padding-right: 20px;"|[[File:FACSsample.png|600px]] | ||
| - | |This is the experimental sample. The same gate is applied to attempt to sort the desired cells (those with higher levels of Nile red fluorescence) compared to the control. | + | |style="text-align:justify; padding: 5px;"|This is the experimental sample. The same gate is applied to attempt to sort the desired cells (those with higher levels of Nile red fluorescence) compared to the control. These 7/10,000 cells were sorted and a culture was made in a matter of minutes. |
|} | |} | ||
<br><br> | <br><br> | ||
| - | *Next we tested these FACS sorted cells on the plate reader to see if we could detect a difference from the FACS sorted and non-FACS sorted cells | + | *Next we tested these FACS sorted cells on the plate reader to see if we could detect a difference from the FACS sorted and non-FACS sorted cells <br><br> |
<center>[[File:FACSsortingBandA.jpg|400px]]</center> | <center>[[File:FACSsortingBandA.jpg|400px]]</center> | ||
| + | *This graph demonstrates that our FACS bioassay method was successful in selecting for cells that exhibit higher levels of PLA-induced fluorescence. | ||
Latest revision as of 00:25, 27 September 2013
Contents |
Develop bioassay to screen PLA production
- We needed a way to detect the PLA once we produced it using the heterologous enzymes
- We decided to use the fluorescent dye Nile red, a intercellular lipid strain
- Nile red does not affect the growth of bacteria, and its fluorescence is quenched in water

This is a figure from Spiekermann et al. 1996 demonstrating Nile red staining of both PHB+ E. coli and PHB negative E. coli
Positive Control
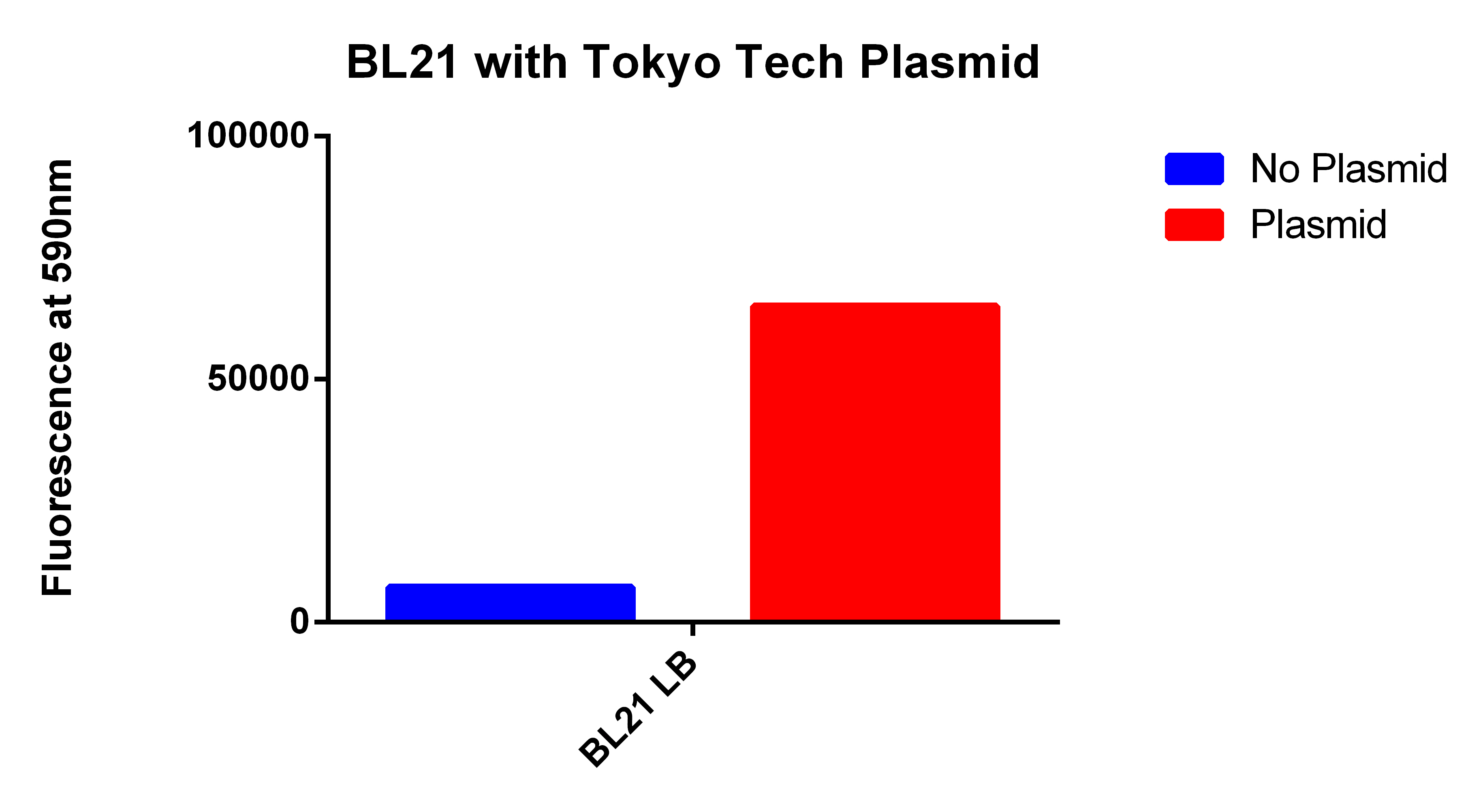
- We used the 2012 Tokyo Tech Biobrick (BBa_K934001) as a positive control to attempt to detect Nile red fluorescence on our plate reader (ex. 530nm, em. 590nm)
- This plasmid had three enzymes which together give E. coli that ability to synthesize P(3HB) a similar plastic PLA
- Cells were grown for 24 hours in the presence of Nile red. The cells were washed and resuspended in PBS.
Our Construct
- We then proceeded to test our plasmid in the plate reader.
- Cells were grown for 24 hours with both enzymes induced and in the presence of Nile red. The cells were washed and re-suspended in PBS. The readings were normalized for optical density.

FACS Sorting
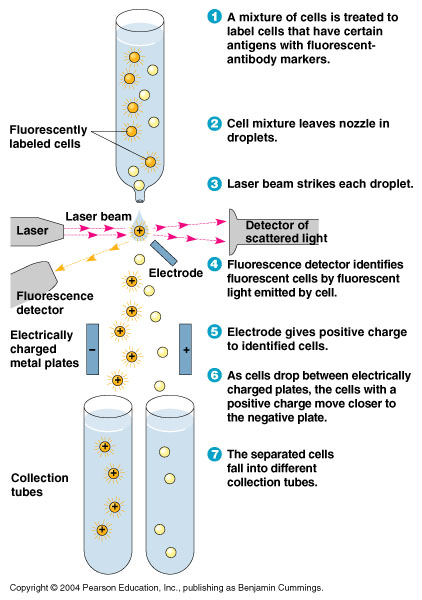
- In order to quickly screen the large diversity created using MAGE, we employed Fluorescence-activated cell sorting (FACS)
- FACS could sort cells based on the Nile red fluorescence, thus indicating those cells that have produced larger quantities of PLA
Testing for PLA
- Nile red has an emission maximum at 598nm when bound to p(3HB) granules according to Spiekermann et al. 1998
- Thus we decided to use PE-Texas Red which has an emission maximum at 615nm in order pick the best colonies.
- PE-Texas Red is a covalently linked fluorochrome used as the detector. The FACS machine measure the emission of this fluorochrome because Nile Red is not a common fluorochrome that a FACS machine would have detector specific to Nile red.
- Next we tested these FACS sorted cells on the plate reader to see if we could detect a difference from the FACS sorted and non-FACS sorted cells

- This graph demonstrates that our FACS bioassay method was successful in selecting for cells that exhibit higher levels of PLA-induced fluorescence.
 "
"