Team:Goettingen/Team/DAC
From 2013.igem.org
(→Navigation:) |
(→Results and Discussion) |
||
| (60 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | {{:Team:Goettingen/templates/ | + | {{:Team:Goettingen/templates/headerGray}} |
<html><script> | <html><script> | ||
$(document).ready(function(){ | $(document).ready(function(){ | ||
| Line 5: | Line 5: | ||
}); | }); | ||
</script> | </script> | ||
| - | </ | + | <img src="https://static.igem.org/mediawiki/2013/6/66/Goe-qNav.png" style="position: fixed;z-index: 199;bottom: 11%;right: 2.5%;_position: absolute;_top: expression(documentElement.scrollTop + documentElement.clientHeight-this.offsetHeight);cursor: pointer;opacity: 0.5;-moz-opacity: 0.5;-khtml-opacity: 0.5;-filter: alpha(opacity = 50 );" onclick="toggle(this.nextElementSibling || getnextElementSibling(this))" /> |
| - | < | + | <div style="position: fixed;z-index: 199;bottom: 19%;right: 2.5%;_position: absolute;_top: expression(documentElement.scrollTop + documentElement.clientHeight-this.offsetHeight);cursor: pointer;display: none;background: #fff;border: 1px solid #A3A3A3;padding: 20px; |
| + | border-radius: 5px;"> | ||
| + | <b>DAC Team:</b><br /> | ||
| + | <ul> | ||
| + | <li><a href="#Introduction">Introduction</a></li> | ||
| + | <li><a href="#Results_and_Discussion">Results and Discussion</a></li> | ||
| + | </ul> | ||
| + | </div> | ||
</div><!--close header--> | </div><!--close header--> | ||
<br /> | <br /> | ||
| Line 17: | Line 24: | ||
<div id="nav"> | <div id="nav"> | ||
</html> | </html> | ||
| - | + | {{:Team:Goettingen/templates/navProject}} | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
=== === | === === | ||
| Line 33: | Line 33: | ||
<div id="col-right" class="bl"> | <div id="col-right" class="bl"> | ||
<br /> | <br /> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/f/f4/Goe-christinaPaetz.jpg" style="display:inline;height:120px"> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/4/46/Goe-miriamLeonard.jpg" style="display:inline;height:120px"> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/5/53/Goe-navaneethanP.jpg" style="display:inline;height:120px"> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/0/09/Goe-soerenRindfleisch.jpg" style="display:inline;height:120px"> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/f/f2/Goe-stephanieSchaefer.jpg" style="display:inline;height:120px"> | ||
| + | <img src="https://static.igem.org/mediawiki/2013/6/6e/Goe-greenColi-crystal.png" style="display:inline;height:120px"> | ||
</html> | </html> | ||
===DAC Team=== | ===DAC Team=== | ||
| - | + | ==Introduction== | |
| - | + | In order to find a novel target that can be inhibited by new antibacterial compounds, we focused on the recently discovered signaling molecule bis-(3’,5’)-cyclic dimeric adenosine monophosphate (c-di-AMP). c-di-AMP was shown to be an essential second messenger in many important pathogenic Gram-positive bacteria (Witte et al., 2008). Moreover, c-di-AMP has a crucial function in cell wall synthesis and spore formation in ''Bacillus subtilis'' (Oppenheimer-Shaanan ''et al.'', 2011; Mehne ''et al.'', 2013). Interestingly, both lack and excess of c-di-AMP have detrimental effects on cell growth and morphology (Luo and Helmann, 2012; Mehne ''et al.'', 2013). | |
| - | < | + | |
| - | < | + | Based on these observations, it makes sense that we take a closer look at the enzyme, the diadenylate cyclase (DAC), which produces c-di-AMP. The DAC CdaA is conserved among several Gram-positive bacteria like ''B. subtilis'', and in the important pathogenic bacteria ''Streptococcus pneumoniae, Staphylococcus aureus'' and ''Listeria monocytogenes'' (Corrigan and Gründling, 2013). So far, the DAC DisA from ''B. subtilis'' has been purified and crystallized (Witte ''et al.'', 2008). However, this protein is not present in all Gram-positive bacteria as well as in many pathogenic bacteria. Therefore we decided to concentrate on the DAC CdaA from ''L. monocytogenes'', which is - as mentioned above - well-conserved in all bacteria that need c-di-AMP for growth! |
| - | </html> | + | |
| + | ==Results and Discussion== | ||
| + | |||
| + | <html><img src="https://static.igem.org/mediawiki/2013/5/51/Goe-greenColi-labcoat.png" style="height: 200px;position: absolute;top: 1347px;left: 361px;" /></html> | ||
| + | <html><img src="https://static.igem.org/mediawiki/2013/6/6e/Goe-greenColi-crystal.png" style="position: absolute;top: 3565px;left: 500px;z-index: 3;" /></html> | ||
| + | Unfortunately, cloning of the full-length gene, encoding the membrane-bound DAC, DacA (Lmo2120) using ''Escherichia coli'' failed. Therefore, we decided to clone a truncated ''dacA'' gene, that encodes a DAC enzyme, lacking the trans-membrane domain. In this protein the first 100 amino acids are missing. Nevertheless, the resulting truncated part still included the essential DAC domain, and therefore represents one of our favorite BioBricks: [http://parts.igem.org/Part:BBa_K1045003 BBa_K1045003]! | ||
| + | |||
| + | By conducting several experiments, we proved that the truncated DacA protein (BBa_K1045003) was not only active ''in vivo'', but also did it's job ''in vitro''. Moreover, we were able to purify the DAC domain in large scale for determining its 3D structure! Using the structure data, one can now search for chemical compounds that interfere with the activity of the cyclase either by testing with available chemical libraries or by computational modeling. In the following text, the experiments will be explained in more detail. However, if you wish to get even more details, please visit the [[Team:Goettingen/Parts|Parts Registry]] or [[Team:Goettingen/NoteBook|our LabBook.]] | ||
| + | |||
| + | The truncated DacA protein ([http://parts.igem.org/Part:BBa_K1045003 BBa_K1045003]) was extended with an N-terminal ''Step''-tag allowing the rapid purification using the ''Strep''-tag:Streptactin purification system. Synthesis of our protein is driven by a T7 promoter. This promoter is recognized by the T7 polymerase, which is encoded in the genome of the <i>E. coli</i> strain BL21. Synthesis of the T7 polymerase can be controlled by Isopropyl-β-D-thiogalactopyranoside (IPTG). The generated plasmid was then used to transform the <i>E. coli</i> strain BL21, which is a powerful strain for synthesis of recombinant proteins. In contrast to Gram-positive bacteria, the Gram-negative bacterium ''E. coli'' does not produce c-di-AMP and growth is not affected by the signaling molecule. | ||
| + | |||
| + | In order to analyze the DAC activity ''in vivo'', DAC production by the ''E. coli'' clones was induced by adding IPTG. The cells were then lysed to extract c-di-AMP from the cells. | ||
| + | |||
| + | By performing the SDS PAGE, we could nicely show that the desired protein was well-produced (Fig. 1). Furthermore, the presence of c-di-AMP in the supernatant of the lysed bacteria was confirmed using LC-MS/MS. Thus, we can conclude that the truncated DacA protein codes for an DAC domain that is active ''in vivo''. | ||
| + | |||
| + | https://static.igem.org/mediawiki/2013/7/70/Goe-dac-fig-1.png | ||
| + | |||
| + | Fig. 1.''' Confirmation of DAC production.''' The SDS PAGE revealed high-level DAC production by three biological replicates. As expected, the DAC domain has a molecular wight of about 20 kDa; Lane 1: Thermo Scientific Page Ruler Plus Prestained Protein Ladder. | ||
| + | |||
| + | |||
| + | DACs catalyze the condensation reaction of two molecules of ATP to a single molecule c-di-AMP, while releasing two pyrophosphate (PP) molecules, consisting of the β-γ-phosphates (P) of each ATP (Fig. 2). In order to analyze the DAC activity ''in vitro'', we performed an assay in which the generation of PP molecules served as an indicator for c-di-AMP production. Among other things such as malachite-green and molybdate, the assay included a PP phosphatase that cleaves the PP molecule into free P molecules. Malachite-green forms a complex with free P molecules and molybdate stabilizes the complex. The product absorption can be measured at a wavelength of 630 nm. With this ''in vitro'' assay, the concentration of free P molecules and thus the conversion rate of ATP to c-di-AMP was analyzed. | ||
| + | |||
| + | Our results confirmed that the DAC domain of the DacA enzyme (Lmo2120) is active in the presence of a divalent cation, ATP and a buffer system at pH 8 (''in vitro''), however, the catalysis rate ''in vivo'' appears to be much more efficient. | ||
| + | |||
| + | <html><a href="https://static.igem.org/mediawiki/2013/7/7f/Goe-dac-fig-2.png" target="_blank"><img src="https://static.igem.org/mediawiki/2013/7/7f/Goe-dac-fig-2.png" width="750"/></a></html> | ||
| + | |||
| + | Fig. 2. '''c-di-AMP production and degradation.''' c-di-AMP is produced from two molecules of ATP by DACs and degraded to pApA by phosphodiasterases. (Edited from Corrigan and Gründling, 2013) | ||
| + | |||
| + | |||
| + | Finally, we are coming to the core of our project, the protein structure of the DacA DAC domain! It is very helpful to have the molecular structure of a DAC in hands because it allows to perform ''in silico'' experiments. These ''in silico'' experiments can lead to the discovery of new antibacterial substance classes, that specifically inhibit DAC activity. The final goal is to use potential antibiotics to treat patients that suffer from infection with pathogenic Gram-positive bacteria. | ||
| + | |||
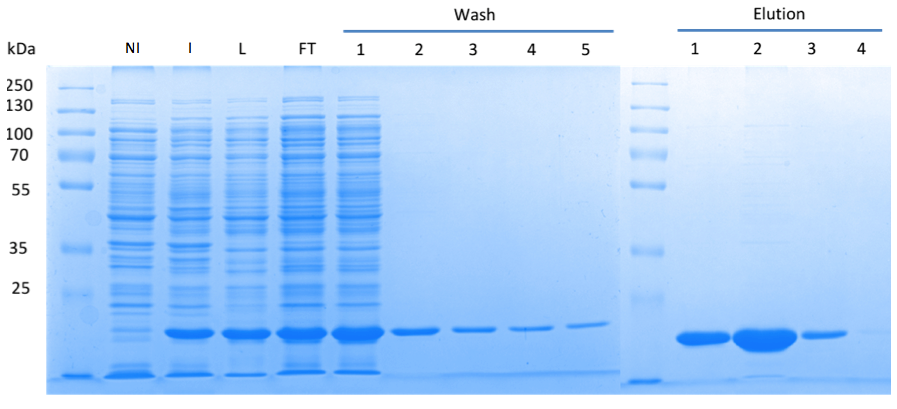
| + | In order to purify a large amount of this protein, we used our BioBrick [http://partsregistry.org/Part:BBa_K1045003 BBa_K1045003] to produce the N-terminally ''Strep''-tagged DAC domain in ''E. coli''. Expression was confirmed by SDS PAGE and the protein showed a molecular weight of about 20 kDa (Fig. 3). The ''E. coli'' strain was grown in a large scale (10 liters), the cells were harvested by centrifugation, lysed and the protein was purified by affinity purification (Fig. 3). | ||
| + | |||
| + | <html><a href="https://static.igem.org/mediawiki/2013/f/fe/Goe-dac-fig-3.png" target="_blank"><img src="https://static.igem.org/mediawiki/2013/f/fe/Goe-dac-fig-3.png" width="750" /></a></html> | ||
| + | |||
| + | Fig. 3. '''SDS PAGE showing high-level production and purification of the DAC domain.''' (A) Lane 1: Thermo Scientific PageRuler Plus Prestained Protein Ladder; NI: Non-induced clone, the cells did not produce the DAC domain and c-di-AMP; I: Induction of ''dac'' expression with IPTG, the clone synthesized the DAC domain and c-di-AMP was detected; L: Lysate; FT: Flow-through; the protein eluted in the first elution fraction. | ||
| + | |||
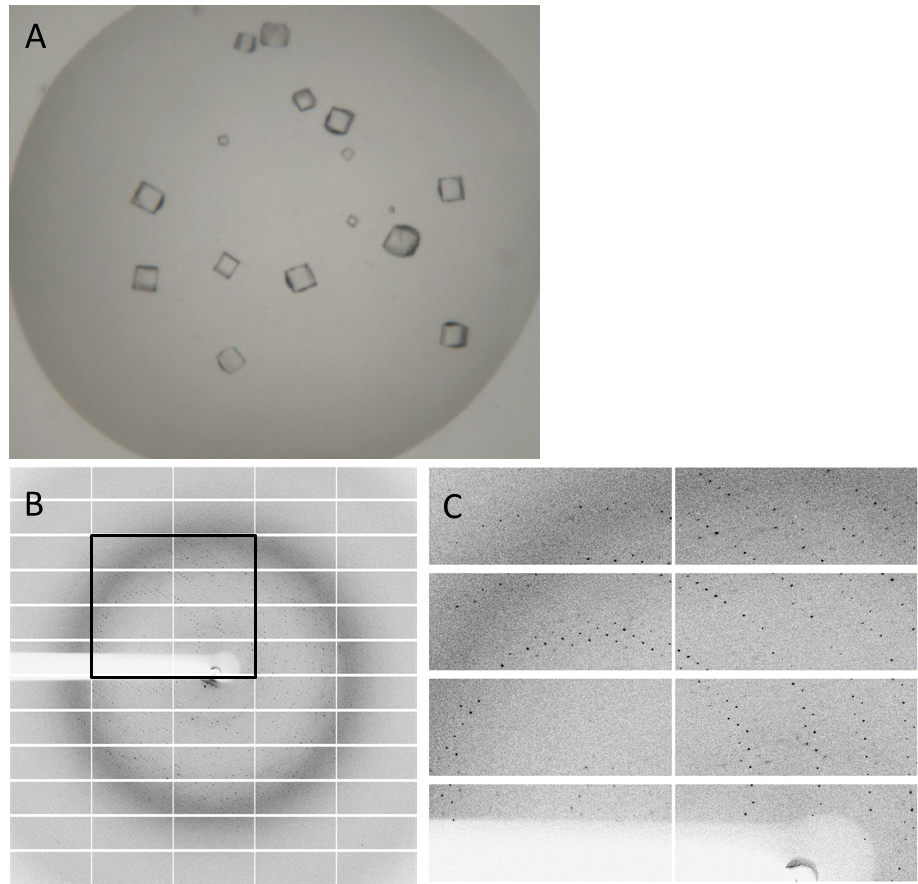
| + | Having dialyzed and concentrated the purified protein, we obtained a protein solution with a concentration of about 10 mg/ml, which was sufficient to perform a crystallization screening. Luckily, we have obtained very nice crystals in the initial crystallization screen (Fig. 4A). In order to find the perfect supplements for growing our crystals, the whole procedure was repeated and the conditions were refined. The crystals yielded an x-ray diffraction pattern, with a resolution of 2,8 Å (Fig. 4B,C). The dataset was measured at the EMBL Hamburg Beamline P13 at the PETRA III synchrotron on the DESY campus, using a PILATUS2 6M X-ray detector (https://www.dectris.com/). | ||
| + | |||
| + | Finally, we were also able to obtain the protein structure (Fig. 5)! The structure shows a globular protein with a distinct ATP-binding cleft. The ribbon model demonstrates the secondary structure composed of α-helices and β-sheets. | ||
| + | |||
| + | <html><a href="https://static.igem.org/mediawiki/2013/a/a8/Goe-dac-fig-4.png" target="_blank"><img src="https://static.igem.org/mediawiki/2013/a/a8/Goe-dac-fig-4.png" width="750"></a></html> | ||
| + | |||
| + | Fig. 4. '''Crystals and diffraction pattern.''' Nice crystals were obtained with a medium concentration of alcohol and other supplements (confidential :-]) '''(A).''' X-ray diffraction image of the DacA DAC domain crystals '''(B)'''; the highlighted box is shown enlarged '''(C)'''. The dataset was measured at the EMBL Hamburg Beamline P13 in the PETRA III synchrotron on the DESY campus. | ||
| + | |||
| + | https://static.igem.org/mediawiki/2013/3/36/Goe-dac-fig-5.png | ||
| + | |||
| + | Fig. 6.''' Protein structure of the DacA DAC domain.''' (A, B) Ribbon model of the DAC domain in its ATP-bound state. (C, D) Surface structure of the DAC domain and the ATP-binding pocket. (E) Magnified view into the ATP-binding pocket | ||
| + | |||
| + | <html><object width="420" height="315"><param name="movie" value="//www.youtube.com/v/9BQOEIVsF-Y?hl=zh_CN&version=3&rel=0"></param><param name="allowFullScreen" value="true"></param><param name="allowscriptaccess" value="always"></param><embed src="//www.youtube.com/v/9BQOEIVsF-Y?hl=zh_CN&version=3&rel=0" type="application/x-shockwave-flash" width="420" height="315" allowscriptaccess="always" allowfullscreen="true"></embed></object></html> | ||
| + | |||
| + | '''References''' | ||
| + | |||
| + | Corrigan RM and Gründling A. (2013) Cyclic di-AMP: another second messenger enters the fray. ''Nat Rev Microbiol''. 11(8):513-524 | ||
| + | |||
| + | Luo Y and Helmann JD. (2012) Analysis of the role of ''Bacillus subtilis'' σ(M) in β-lactam resistance reveals an essential role for c-di-AMP in peptidoglycan homeostasis. ''Mol Microbiol.'' 83(3):623-639 | ||
| + | |||
| + | Mehne FM, Gunka K, Eilers H, Herzberg C, Kaever V, Stülke J. (2013) Cyclic di-AMP homeostasis in ''Bacillus subtilis'': both lack and high level accumulation of the nucleotide are detrimental for cell growth.'' J Biol Chem.'' 288(3):2004-2017 | ||
| + | |||
| + | Oppenheimer-Shaanan Y, Wexselblatt E, Katzenhendler J, Yavin E, Ben-Yehuda S. (2011) c-di-AMP reports DNA integrity during sporulation in ''Bacillus subtilis''. ''EMBO Rep.'' 12(6):594-601 | ||
| + | |||
| + | Witte G, Hartung S, Büttner K, Hopfner KP. (2008) Structural biochemistry of a bacterial checkpoint protein reveals diadenylate cyclase activity regulated by DNA recombination intermediates.'' Mol Cell.'' 30(2):167-178 | ||
=== === | === === | ||
<html> | <html> | ||
<a href="/Team:Goettingen/Team/Array" class="moreinfo fl"><b>Previous</b></a> | <a href="/Team:Goettingen/Team/Array" class="moreinfo fl"><b>Previous</b></a> | ||
| - | <a href="/Team:Goettingen/Project/ | + | <a href="/Team:Goettingen/Project/Outlook" class="moreinfo fr"><b>Next</b></a> |
<br /><br /> | <br /><br /> | ||
</div><!--close col-right--> | </div><!--close col-right--> | ||
<br /> | <br /> | ||
| + | </div><!--close f-box--> | ||
</html> | </html> | ||
Latest revision as of 09:45, 28 October 2013
 "
"