Team:Newcastle/Project/shuffling endosymbiosis
From 2013.igem.org
| (15 intermediate revisions not shown) | |||
| Line 16: | Line 16: | ||
==Purpose== | ==Purpose== | ||
| - | <p>Over the past eight years | + | |
| - | + | <p>Over the past eight years iGEM teams have dreamt up exciting and innovative ways of harnessing Synthetic Biology. However this relatively new field faces challenges such as the complexity of biological systems, a public suspicious of genetic modification and the problem of producing a viable, high quality yield of the desired product. Using L-forms allows the use of a technique with the potential to solve this final obstacle. </p> | |
| - | + | ||
| - | + | <p>This technique is genome shuffling, and if harnessed it could improve the efficiency of hundreds of iGEM projects as well as cell factories across the field of Synthetic Biology. </p> | |
| - | + | ||
| - | <p>Processes in biology can be refined and improved by evolution, but in nature this occurs very slowly. We propose using L-forms to artificially evolve ''B. subtilis'' | + | <p>Processes in biology can be refined and improved by evolution, but in nature this occurs very slowly. We propose using genome shuffling and L-forms to artificially evolve ''B. subtilis''. This technique increases the rate of evolution, allowing the improvement of any biological system or phenotype in a feasible timeframe. </p> |
| - | </p> | + | |
<p>Selective breeding has been central to the success of agriculture. We are extending these principles to microorganisms. This is potentially very powerful, as microorganisms replicate very quickly, allowing numerous rounds of selection over short time periods. </p> | <p>Selective breeding has been central to the success of agriculture. We are extending these principles to microorganisms. This is potentially very powerful, as microorganisms replicate very quickly, allowing numerous rounds of selection over short time periods. </p> | ||
| Line 30: | Line 29: | ||
<p>''‘…two rounds of genome shuffling were sufficient to achieve results that had previously required 20 rounds of CSI. Theoretically, this is the difference between the asexual process of CSI and the hypersexual process of genome shuffling. Practically, this is the difference between 24,000 assays and approximately 1 year of effort, and roughly 1,000,000 assays and 20 years of effort.’''</p> | <p>''‘…two rounds of genome shuffling were sufficient to achieve results that had previously required 20 rounds of CSI. Theoretically, this is the difference between the asexual process of CSI and the hypersexual process of genome shuffling. Practically, this is the difference between 24,000 assays and approximately 1 year of effort, and roughly 1,000,000 assays and 20 years of effort.’''</p> | ||
<p>CSI refers to classical strain improvement (random mutagenesis and screening), currently the most popular method for the development of commercial microorganisms</p> | <p>CSI refers to classical strain improvement (random mutagenesis and screening), currently the most popular method for the development of commercial microorganisms</p> | ||
| - | <p>We believe that using L-forms rather than protoplast in genome shuffling is even more advantageous. Firstly protoplasting gram-negative bacteria such as '' | + | <p>We believe that using L-forms rather than protoplast in genome shuffling is even more advantageous. Firstly protoplasting bacteria, especially gram-negative bacteria such as ''E. coli'', can be difficult due to the complexity of the cell wall. Using L-forms circumvents this problem. Furthermore L-forms, whilst fragile, are more stable and therefore easier to work than protoplasts as they are less likely to burst due to increased membrane flexibility. Unlike protoplasts, L-forms can also grow and replicate, so can comfortably be kept in a cell wall-removed state for longer periods of time. Reintegration of the cell wall after experiments are complete is also easier with L-forms. </p> |
== Our Objectives == | == Our Objectives == | ||
| Line 37: | Line 36: | ||
We have constructed BioBricks that code for DNA tags. These are indiscriminate DNA binding proteins fused to GFP or RFP. The BioBricks are both under the control of an IPTG inducible promoter. This allows us to produce two types of cell: those which have a green fluorescing genome, and those with a red fluorescing genome. We place one of each at each end of a microfluidics chamber, and apply pressure. The cells will be pushed together and fuse, before splitting again into two cells. If recombination has taken place each cell should have both green and red fluorescence. This is because the DNA tags remain associated to the specific DNA they are bound to as bits of DNA are ‘swapped’ between genomes.</p> | We have constructed BioBricks that code for DNA tags. These are indiscriminate DNA binding proteins fused to GFP or RFP. The BioBricks are both under the control of an IPTG inducible promoter. This allows us to produce two types of cell: those which have a green fluorescing genome, and those with a red fluorescing genome. We place one of each at each end of a microfluidics chamber, and apply pressure. The cells will be pushed together and fuse, before splitting again into two cells. If recombination has taken place each cell should have both green and red fluorescence. This is because the DNA tags remain associated to the specific DNA they are bound to as bits of DNA are ‘swapped’ between genomes.</p> | ||
<p>Dual expression will not be due to our fusion protein binding to DNA during the fused cell state as the cells will be removed from an IPTG containing media the night before we fuse. This is acceptable due to the long (18-24 hour) half-life of the protein. Furthermore, the protein binds strongly to DNA, so there isn’t a significant risk of it unbinding and rebinding to another DNA strand.</p> | <p>Dual expression will not be due to our fusion protein binding to DNA during the fused cell state as the cells will be removed from an IPTG containing media the night before we fuse. This is acceptable due to the long (18-24 hour) half-life of the protein. Furthermore, the protein binds strongly to DNA, so there isn’t a significant risk of it unbinding and rebinding to another DNA strand.</p> | ||
| + | |||
| + | ==Modelling== | ||
| + | |||
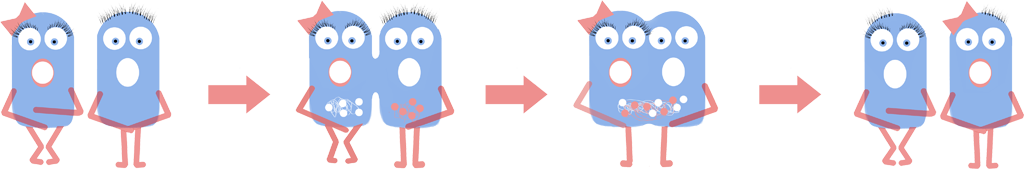
| + | We created an animation to model genomic fusion visually. The simulation shows the expected behaviour of fusion events of L-forms. The L-forms express one of the two different BioBricks BBa_K1185001 and BBa_K1185002 which tag the genome with green or red fluorescence. This simulation also shows the predicted colours of the L-forms after fusion. For more information of the model, please see the [https://2013.igem.org/Team:Newcastle/Modelling/Cell_Fusion cell fusion modelling page]. | ||
| Line 57: | Line 60: | ||
[[File:BareCillus GFP.png|centre|200px]] | [[File:BareCillus GFP.png|centre|200px]] | ||
| - | ''Figure 2. L-forms producing HBsu-GFP. This fusion protein has bound to the genome of these L-forms and emits a green light upon excitation '' | + | ''Figure 2. L-forms producing HBsu-GFP. This fusion protein has bound to the genome of these L-forms and emits a green light upon excitation.'' |
[[File:RFP.png|centre|200px]] | [[File:RFP.png|centre|200px]] | ||
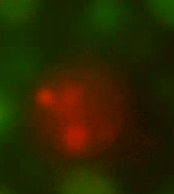
| - | ''Figure 3. An L-form producing HBsu-RFP (central), surrounded by L-forms producing HBsu-GFP. HBsu- | + | ''Figure 3. An L-form producing HBsu-RFP (central), surrounded by L-forms producing HBsu-GFP. HBsu-RFP has bound to the genome of this L-form and is emitting a red light upon excitation.'' |
[[File:MIXED.png|centre|200px]] | [[File:MIXED.png|centre|200px]] | ||
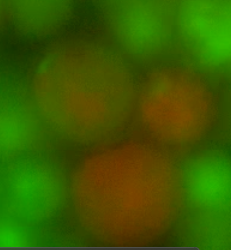
| - | ''Figure 4. Fusants of L-forms expressing the two different HBsu-xFP BioBricks. It can be seen that the fluorescent signals from the two different fusion proteins are mixed together. This shows that recombination has occurred '' | + | ''Figure 4. Fusants of L-forms expressing the two different HBsu-xFP BioBricks. It can be seen that the fluorescent signals from the two different fusion proteins are mixed together. This shows that recombination has occurred.'' |
[[File:MIXED 1.png|centre|200px]] | [[File:MIXED 1.png|centre|200px]] | ||
| - | ''Figure 5. Another fusant of L-forms expressing the two different HBsu-xFP BioBricks. It can be seen that the fluorescent signals from the two different fusion proteins are mixed together. This shows that recombination has occurred'' | + | ''Figure 5. Another fusant of L-forms expressing the two different HBsu-xFP BioBricks. It can be seen that the fluorescent signals from the two different fusion proteins are mixed together. This shows that recombination has occurred.'' |
==References== | ==References== | ||
[http://www.nature.com/nature/journal/v415/n6872/full/415644a.html Zhang, Y. X., Perry, K., Vinci, V. A., Powell, K., Stemmer, W. P. C., & del Cardayre, S. B. (2002). Nature, 415(6872), 644–646.- Genome shuffling leads to rapid phenotypic improvement in bacteria] | [http://www.nature.com/nature/journal/v415/n6872/full/415644a.html Zhang, Y. X., Perry, K., Vinci, V. A., Powell, K., Stemmer, W. P. C., & del Cardayre, S. B. (2002). Nature, 415(6872), 644–646.- Genome shuffling leads to rapid phenotypic improvement in bacteria] | ||
Latest revision as of 18:33, 28 October 2013

Contents |
Genome shuffling
B. subtilis L-forms were squeezed together using microfluidics in the presence of polyethylene glycol (PEG), which facilitates cellular fusion. Fusion will result in the genomes coming into close proximity with one another. The L-forms being fused are from the same strain; therefore their genomes will be almost identical. This will allow homologous recombination, where similar sections of DNA are ‘swapped’ between genomes, to take place. Subsequent splitting of the fusion product will give two new cells, neither of which is identical to their ‘parents’. The cells which exhibit improved phenotypes can be selected and subjected to this process again and again. In this way, improved bacteria can be created!
We visualised this 'genome shuffling' by fluorescently tagging our L-form genomes with two different colours, fusing the cells and observing whether the two colours mixed. We created and used the parts [http://parts.igem.org/Part:BBa_K1185001 BBa_K1185001] and [http://parts.igem.org/Part:BBa_K1185002 BBa_K1185002], which marked bacterial genomes with HBsu-sfGFP (a green colour) and HBsu-RFP (a red colour) respectively.
This technique exploits the diversity that naturally exists between members of the same species. Parallels can be drawn between this process and meiosis, and could even be viewed as making the bacteria procreate. Our new cells will be phenotypically distinct, in the same way that a child is different from either of its parents. Some of the cells produced will have enhanced function of the trait of interest. This can be assayed for, these cells isolated, and the process repeated multiple times, giving us new and improved (for the phenotype of interest) cells. In theory, different strains, or even species, of bacteria could be fused and exhibit shuffling, increasing variation further.
Purpose
Over the past eight years iGEM teams have dreamt up exciting and innovative ways of harnessing Synthetic Biology. However this relatively new field faces challenges such as the complexity of biological systems, a public suspicious of genetic modification and the problem of producing a viable, high quality yield of the desired product. Using L-forms allows the use of a technique with the potential to solve this final obstacle.
This technique is genome shuffling, and if harnessed it could improve the efficiency of hundreds of iGEM projects as well as cell factories across the field of Synthetic Biology.
Processes in biology can be refined and improved by evolution, but in nature this occurs very slowly. We propose using genome shuffling and L-forms to artificially evolve B. subtilis. This technique increases the rate of evolution, allowing the improvement of any biological system or phenotype in a feasible timeframe.
Selective breeding has been central to the success of agriculture. We are extending these principles to microorganisms. This is potentially very powerful, as microorganisms replicate very quickly, allowing numerous rounds of selection over short time periods.
Directed evolution of bacteria and even individual genes is being increasingly explored. For example GFP, now essential to researchers, initially needed to be modified by directed evolution to produce enough fluorescence to be useful.
In this case modification was accomplished using mutagenic PCR, which mutates a single gene. It relies on the researcher knowing, and having sequence data on what genes govern the phenotype of interest. But many genetic circuits are poorly understood or controlled by many different genes, each of which could potentially have its function improved. Or it may even be desirable to engineer multiple phenotypes simultaneously. Genome shuffling is therefore a powerful tool for refining complex systems, as it effects the whole genome and requires no knowledge of how the system being engineered works.
Protoplasts, cells in which the cell wall has been removed with an enzyme such as lysozyme, have already been used in fusion and genome shuffling. In the paper [http://www.nature.com/nature/journal/v415/n6872/full/415644a.html Genome shuffling leads to rapid phenotypic improvement in bacteria], the authors claim that:
‘…two rounds of genome shuffling were sufficient to achieve results that had previously required 20 rounds of CSI. Theoretically, this is the difference between the asexual process of CSI and the hypersexual process of genome shuffling. Practically, this is the difference between 24,000 assays and approximately 1 year of effort, and roughly 1,000,000 assays and 20 years of effort.’
CSI refers to classical strain improvement (random mutagenesis and screening), currently the most popular method for the development of commercial microorganisms
We believe that using L-forms rather than protoplast in genome shuffling is even more advantageous. Firstly protoplasting bacteria, especially gram-negative bacteria such as E. coli, can be difficult due to the complexity of the cell wall. Using L-forms circumvents this problem. Furthermore L-forms, whilst fragile, are more stable and therefore easier to work than protoplasts as they are less likely to burst due to increased membrane flexibility. Unlike protoplasts, L-forms can also grow and replicate, so can comfortably be kept in a cell wall-removed state for longer periods of time. Reintegration of the cell wall after experiments are complete is also easier with L-forms.
Our Objectives
The timescale of iGEM means that it isn’t feasible to actually evolve and show improvement of a phenotype of interest. However we can show that recombination is taking place, and then directed evolution would be possible. We have constructed BioBricks that code for DNA tags. These are indiscriminate DNA binding proteins fused to GFP or RFP. The BioBricks are both under the control of an IPTG inducible promoter. This allows us to produce two types of cell: those which have a green fluorescing genome, and those with a red fluorescing genome. We place one of each at each end of a microfluidics chamber, and apply pressure. The cells will be pushed together and fuse, before splitting again into two cells. If recombination has taken place each cell should have both green and red fluorescence. This is because the DNA tags remain associated to the specific DNA they are bound to as bits of DNA are ‘swapped’ between genomes.
Dual expression will not be due to our fusion protein binding to DNA during the fused cell state as the cells will be removed from an IPTG containing media the night before we fuse. This is acceptable due to the long (18-24 hour) half-life of the protein. Furthermore, the protein binds strongly to DNA, so there isn’t a significant risk of it unbinding and rebinding to another DNA strand.
Modelling
We created an animation to model genomic fusion visually. The simulation shows the expected behaviour of fusion events of L-forms. The L-forms express one of the two different BioBricks BBa_K1185001 and BBa_K1185002 which tag the genome with green or red fluorescence. This simulation also shows the predicted colours of the L-forms after fusion. For more information of the model, please see the cell fusion modelling page.
Results
In order to observe cell fusion and recombination, we used an agarose based single-cell chemostat microfluidic chamber designed by Harvard University technology. This chip allows us to have fine control over single cells movement, which is beneficial in bacterial single-cell and communities’ studies. Figure 1 shows the design of this chamber. By using the BioBricks we created, we were able to produce two types of L-forms: those which have a green fluorescing genome, and those with a red fluorescing genome. We placed one of each at each end of a microfluidic chamber, and used air pressure to apply the physical pressure required to trigger cell fusion and control their directionality. Once the cells fused, genome shuffling between the two chromosomes was allowed to occur. The fused cells split back into two after a while.
Figure 1. The design for the microfluidics chip used to manipulate our L-forms
In order to observe whether recombination had taken place, we exposed the cells to the excitation wavelengths of both GFP and RFP and observed for fluorescence at the emittance wavelengths of these two proteins. If recombination had occurred each ‘new’ cell should either exhibit a mosaic of both green and red fluorescence or show a brownish colour (if pictures from both wavelengths are stacked on top of one another).
Figures 2 and 3 shows cells prior to fusion. Figure 2 shows L-forms with GFP attached to their genome via HBsu proteins. Likewise, Figure 3 depicts an L-form with RFP bound to its genome via HBsu (surrounded by GFP-tagged L-forms).
Figures 4 and 5 show the recombined genomes of L-forms after they have undergone fusion events. A brownish colour, or green/red mosaic, results from these cells (as predicted) due to their genomes recombining which has caused the mixing of the fluorescent signals.
Here we have proved that through the use of L-forms we were able to perform genome shuffling. This opens up lots of new possible applications with L-forms as mentioned previously.
Figure 2. L-forms producing HBsu-GFP. This fusion protein has bound to the genome of these L-forms and emits a green light upon excitation.
Figure 3. An L-form producing HBsu-RFP (central), surrounded by L-forms producing HBsu-GFP. HBsu-RFP has bound to the genome of this L-form and is emitting a red light upon excitation.
Figure 4. Fusants of L-forms expressing the two different HBsu-xFP BioBricks. It can be seen that the fluorescent signals from the two different fusion proteins are mixed together. This shows that recombination has occurred.
Figure 5. Another fusant of L-forms expressing the two different HBsu-xFP BioBricks. It can be seen that the fluorescent signals from the two different fusion proteins are mixed together. This shows that recombination has occurred.
References
[http://www.nature.com/nature/journal/v415/n6872/full/415644a.html Zhang, Y. X., Perry, K., Vinci, V. A., Powell, K., Stemmer, W. P. C., & del Cardayre, S. B. (2002). Nature, 415(6872), 644–646.- Genome shuffling leads to rapid phenotypic improvement in bacteria]
 "
"