Team:Tokyo Tech
From 2013.igem.org
| (186 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{tokyotechmenudark}} | {{tokyotechmenudark}} | ||
<html> | <html> | ||
| - | + | <script language="javascript"> | |
| + | function show(k) | ||
| + | { | ||
| + | if(k==1){ | ||
| + | document.getElementById("circuit").src="https://static.igem.org/mediawiki/2013/f/fc/Titech2013_home_step1circuitanime.gif"; | ||
| + | document.getElementById("step1").src="https://static.igem.org/mediawiki/2013/4/47/Titech2013_home_step1active.gif"; | ||
| + | document.getElementById("step2").src="https://static.igem.org/mediawiki/2013/b/b0/Titech2013_home_step2.gif"; | ||
| + | document.getElementById("step3").src="https://static.igem.org/mediawiki/2013/0/0f/Titech2013_home_step3.gif"; | ||
| + | document.getElementById("step4").src="https://static.igem.org/mediawiki/2013/9/9e/Titech2013_home_step4.gif"; | ||
| + | document.getElementById("step5").src="https://static.igem.org/mediawiki/2013/e/e6/Titech2013_home_step5.gif"; | ||
| + | } | ||
| + | if(k==2){ | ||
| + | document.getElementById("circuit").src="https://static.igem.org/mediawiki/2013/b/bc/Titech2013_home_step2circuitanime.gif"; | ||
| + | document.getElementById("step1").src="https://static.igem.org/mediawiki/2013/d/d8/Titech2013_home_step1.gif"; | ||
| + | document.getElementById("step2").src="https://static.igem.org/mediawiki/2013/0/07/Titech2013_home_step2active.gif"; | ||
| + | document.getElementById("step3").src="https://static.igem.org/mediawiki/2013/0/0f/Titech2013_home_step3.gif"; | ||
| + | document.getElementById("step4").src="https://static.igem.org/mediawiki/2013/9/9e/Titech2013_home_step4.gif"; | ||
| + | document.getElementById("step5").src="https://static.igem.org/mediawiki/2013/e/e6/Titech2013_home_step5.gif"; | ||
| + | } | ||
| + | if(k==3){ | ||
| + | document.getElementById("circuit").src="https://static.igem.org/mediawiki/2013/a/af/Titech2013_home_step3circuitanime.gif"; | ||
| + | document.getElementById("step1").src="https://static.igem.org/mediawiki/2013/d/d8/Titech2013_home_step1.gif"; | ||
| + | document.getElementById("step2").src="https://static.igem.org/mediawiki/2013/b/b0/Titech2013_home_step2.gif"; | ||
| + | document.getElementById("step3").src="https://static.igem.org/mediawiki/2013/9/9c/Titech2013_home_step3active.gif"; | ||
| + | document.getElementById("step4").src="https://static.igem.org/mediawiki/2013/9/9e/Titech2013_home_step4.gif"; | ||
| + | document.getElementById("step5").src="https://static.igem.org/mediawiki/2013/e/e6/Titech2013_home_step5.gif"; | ||
| + | } | ||
| + | if(k==4){ | ||
| + | document.getElementById("circuit").src="https://static.igem.org/mediawiki/2013/5/59/Titech2013_home_step4circuitanime.gif"; | ||
| + | document.getElementById("step1").src="https://static.igem.org/mediawiki/2013/d/d8/Titech2013_home_step1.gif"; | ||
| + | document.getElementById("step2").src="https://static.igem.org/mediawiki/2013/b/b0/Titech2013_home_step2.gif"; | ||
| + | document.getElementById("step3").src="https://static.igem.org/mediawiki/2013/0/0f/Titech2013_home_step3.gif"; | ||
| + | document.getElementById("step4").src="https://static.igem.org/mediawiki/2013/7/71/Titech2013_home_step4active.gif"; | ||
| + | document.getElementById("step5").src="https://static.igem.org/mediawiki/2013/e/e6/Titech2013_home_step5.gif"; | ||
| + | } | ||
| + | if(k==5){ | ||
| + | document.getElementById("circuit").src="https://static.igem.org/mediawiki/2013/3/37/Titech2013_home_step5circuitanime.gif"; | ||
| + | document.getElementById("step1").src="https://static.igem.org/mediawiki/2013/d/d8/Titech2013_home_step1.gif"; | ||
| + | document.getElementById("step2").src="https://static.igem.org/mediawiki/2013/b/b0/Titech2013_home_step2.gif"; | ||
| + | document.getElementById("step3").src="https://static.igem.org/mediawiki/2013/0/0f/Titech2013_home_step3.gif"; | ||
| + | document.getElementById("step4").src="https://static.igem.org/mediawiki/2013/9/9e/Titech2013_home_step4.gif"; | ||
| + | document.getElementById("step5").src="https://static.igem.org/mediawiki/2013/e/e8/Titech2013_home_step5active.gif"; | ||
| + | } | ||
| + | } | ||
| + | |||
| + | </script> | ||
| + | <a name="top"></a> | ||
<div id="text-area"><br> | <div id="text-area"><br> | ||
| - | + | <div class="box" id="title"> | |
| - | + | Home | |
| - | + | </div> | |
| - | + | <div class="box"> | |
| + | <br> | ||
| + | <table align="center"> | ||
<tr> | <tr> | ||
| - | <td colspan=2 | + | <td colspan=2> |
| - | + | ||
| - | + | <h3><font size="6">P</font>roject Background</h3> | |
| - | + | <h2><p> | |
| - | + | In this iGEM Competition, we intended to tell the public about the development of synthetic biology, especially about the network programming, as well as we enjoyed our activity for iGEM. Tokyo Tech 2013 assisted with an experiment workshop for high school students, participated in a poster session and collected feedback from public people as human practice (Fig. 1-1-1). We learned that an interesting story is an easier way to make general people | |
| - | + | understand the importance of genetic programming in synthetic biology. | |
| - | <td | + | To respond further to the feedback that we received, we also address a farming issue. Thus, we designed a story which contains a farming issue and a state switching circuit in <i>E. coli</i>, the life of ninja: battle and farming. |
| - | + | ||
| - | <h4><br> | + | |
| - | + | <div align="right"><a href="https://2013.igem.org/Team:Tokyo_Tech/Human_Practice#1._Introduction">(Go to Human Practice page)</a></div></p></h2> | |
| - | + | <td> | |
| + | <a href="https://2013.igem.org/Team:Tokyo_Tech/Human_Practice#2._Human_Practice_1:_Experiment_Workshop"><img src="https://static.igem.org/mediawiki/2013/0/06/Titech2013_home_Fig1_human_experiment.jpg" width="300"></a><br> | ||
| + | <h4>[Fig. 1-1-1. Poster session]<br> | ||
| + | Yellow labels and blue labels are sticked on our posters. | ||
</h4> | </h4> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| + | |||
<tr> | <tr> | ||
| - | <td | + | <td colspan=1><iframe width="300" height="220" src="//www.youtube.com/embed/-zCe53lq09U" frameborder="0" allowfullscreen></iframe> |
| - | + | <br> | |
| - | <h4>[ | + | <h4>[Fig. 1-1-2. Ninja vs. Samurai in Tokyo Tech]<br></h4> |
| - | + | ||
| - | + | ||
| - | + | ||
</td> | </td> | ||
| - | <td colspan=2 | + | <td colspan=2> |
| + | <h3><font size="6">S</font>tory</h3> | ||
<h2><p> | <h2><p> | ||
| - | Ninja is a | + | Ninja is a Japan's ancient spy-warrior. Ninja usually mimics an ordinary civilian. Once he detects samurai who is the target of assassination, he immediately gets ready for battle. He attacks samurai by throwing so-called "ninja stars". These weapons are called "shuriken" in Japanese (Fig. 1-1-2). |
</p></h2> | </p></h2> | ||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td colspan=3> | ||
| + | <br><br> | ||
| + | <div id="marginbox"> | ||
| + | <h3 style="color:#d3381c;">Click a step shown below to know what happens in our circuit!</h3> | ||
| + | <table align="center" border=0> | ||
| + | <tr><td> | ||
| + | <span onclick="show(1)"><img id="step1" src="https://static.igem.org/mediawiki/2013/4/47/Titech2013_home_step1active.gif" width="400" border=0 onmouseover=this.src="https://static.igem.org/mediawiki/2013/4/47/Titech2013_home_step1active.gif" onmouseout=this.src="https://static.igem.org/mediawiki/2013/d/d8/Titech2013_home_step1.gif"></span></td> | ||
| + | |||
| + | <td rowspan=5 align="center"> | ||
| + | <img id="circuit" src="https://static.igem.org/mediawiki/2013/f/fc/Titech2013_home_step1circuitanime.gif" width="400" height="150" border=0> | ||
| + | |||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr><td><span onclick="show(2)"><img id="step2" src="https://static.igem.org/mediawiki/2013/b/b0/Titech2013_home_step2.gif" width="400" border=0 onmouseover=this.src="https://static.igem.org/mediawiki/2013/0/07/Titech2013_home_step2active.gif" onmouseout=this.src="https://static.igem.org/mediawiki/2013/b/b0/Titech2013_home_step2.gif"></span></td></tr> | ||
| + | |||
| + | <tr><td><span onclick="show(3)"><img id="step3" src="https://static.igem.org/mediawiki/2013/0/0f/Titech2013_home_step3.gif" width="400" border=0 onmouseover=this.src="https://static.igem.org/mediawiki/2013/9/9c/Titech2013_home_step3active.gif" onmouseout=this.src="https://static.igem.org/mediawiki/2013/0/0f/Titech2013_home_step3.gif"></span></td></tr> | ||
| + | |||
| + | <tr><td><span onclick="show(4)"><img id="step4" src="https://static.igem.org/mediawiki/2013/9/9e/Titech2013_home_step4.gif" width="400" border=0 onmouseover=this.src="https://static.igem.org/mediawiki/2013/7/71/Titech2013_home_step4active.gif" onmouseout=this.src="https://static.igem.org/mediawiki/2013/9/9e/Titech2013_home_step4.gif"></span></td></tr> | ||
| + | |||
| + | <tr><td><span onclick="show(5)"><img id="step5" src="https://static.igem.org/mediawiki/2013/e/e6/Titech2013_home_step5.gif" width="400" border=0 onmouseover=this.src="https://static.igem.org/mediawiki/2013/e/e8/Titech2013_home_step5active.gif" onmouseout=this.src="https://static.igem.org/mediawiki/2013/e/e6/Titech2013_home_step5.gif"></span></td></tr> | ||
| + | |||
| + | </table> | ||
| + | </div><br><br> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td colspan=2> | ||
| + | <h3><font size="6">P</font>roject Overview</h3> | ||
| + | <h2><p>In our programming of artificial genetic circuit, <i>E. ninja</i> heads casts. In response to <i>E. civilian</i> signal or <i>E. samurai</i> signal, <i>E. ninja</i> changes its state: "Mimic state" or "Attack state". The circuit of <i>E. ninja</i> contains a bi-stable switch part and a signal dependent switching part. We decided to use 3OC6HSL and 3OC12HSL as the signals. The crosstalk between these two signals is well known as a significant problem in synthetic biology. To realize an accurate switching, we designed a circuit which achieved the circumvention of crosstalk that occurs in bacterial cell-cell communication system (Fig. 1-1-4). | ||
| + | <div align="right"><a href="https://2013.igem.org/Team:Tokyo_Tech/Project/Ninja_State_Switching#1._Introduction">(Go to State Switching page)</a></div> | ||
| + | </p></h2> | ||
| + | </td> | ||
| + | <td> | ||
| + | <div align="center"><a href="https://2013.igem.org/Team:Tokyo_Tech/Experiment/Crosstalk_Circumvention_Assay#1._Introduction"><img src="https://static.igem.org/mediawiki/2013/6/6f/Titech2013_home_Fig3_crosstalk_assay.png" width="120" height="200" style="border: #E7E5F0 2px"></a></div><br> | ||
| + | <h4>[Fig. 1-1-3. The result of our wet experiment for the circumvention of the crosstalk]<br> | ||
| + | The level of GFP expression in cells, when TetR is active, was clearly lower than that when TetR is inhibited. Even with activated LasR, <i>lux/tet</i> hybrid promoter (<a href="http://parts.igem.org/Part:BBa_K1139110">BBa_K1139110</a>) was repressed by TetR precisely. This result showed that our network including TetR can circumvent crosstalk by the activated LasR. | ||
| + | </h4><br> | ||
| + | |||
</td> | </td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
<td colspan=3 align="center"> | <td colspan=3 align="center"> | ||
| - | < | + | <a href="https://2013.igem.org/Team:Tokyo_Tech/Project/Ninja_State_Switching#2-2._The_problem:_crosstalk"><img src="https://static.igem.org/mediawiki/2013/6/6c/Titech2013_home_Fig4_genetic_circuit.png" width="600" height="300"></a><br> |
| - | + | <h4>[Fig. 1-1-4. Our designed circuit for the circumvention of crosstalk]<br> | |
| - | </ | + | We designed the circumvention of crosstalk by network engineering. |
| + | |||
| + | </h4> | ||
</td> | </td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td colspan= | + | <td colspan=3> |
| - | + | <h2><p> | |
| - | + | Our wet experiment results showed that the combination of <i>lux/tet</i> hybrid promoter and TetR protein circumvented the crosstalk by preventing the LasR protein from acting on LuxR-binding sequences (Fig. 1-1-3). Our mathematical model based on these results showed the circumvention of crosstalk in the circuit including toggle switch and crosstalk circumvention system(Fig. 1-1-5). | |
| - | <div align="right"><a href=" | + | <div align="right"><a href="https://2013.igem.org/Team:Tokyo_Tech/Modeling/Crosstalk_Circumvention#1._Introduction">(Go to Modeling page)</a></div> |
| - | + | ||
| - | + | ||
</p></h2> | </p></h2> | ||
</td> | </td> | ||
| - | <td | + | |
| - | < | + | </tr> |
| - | <h4>[ | + | <tr> |
| - | + | <td colspan=3> | |
| - | < | + | <div align="center"><a href="https://2013.igem.org/Team:Tokyo_Tech/Modeling/Crosstalk_Circumvention#3._Analytical_method_of_effectiveness_of_crosstalk_prevention_circuit"><img src="/wiki/images/thumb/b/b5/Titech2013_Ninja_State_Switching_2-1_4-3.jpg/700px-Titech2013_Ninja_State_Switching_2-1_4-3.jpg" width="700"></a></div><br> |
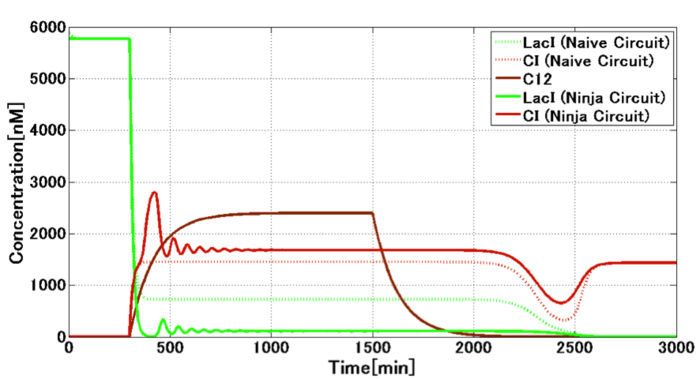
| - | + | <div align="center"><h4>[Fig. 1-1-5. Our mathematical model for the circuit of <i>E. ninja</i>]<br> | |
| - | <a href=""><img src="https://static.igem.org/mediawiki/2013/ | + | The solid/dotted lines stand for the case with/without the crosstalk circumvention. |
| - | <h4>[ | + | The expression of LacI is repressed through the crosstalk circumvention. |
| - | + | ||
| - | + | </h4></div> | |
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td> | ||
| + | <a href="https://2013.igem.org/Team:Tokyo_Tech/Experiment/pSB-M13_Plasmid_Assay#1._Introduction"><img src="https://static.igem.org/mediawiki/2013/5/57/Titech2013_Project_M13_shuriken_Fig_2-2-3.png" width="240"></a><br> | ||
| + | <h4>[Fig. 1-1-6. Our new part for inducible phage release]<br>We designed a new part for inducible phage release. Varoius promoters are allowed to be inserted upstream of <i>g2p</i> to regulate the phage release. | ||
</h4> | </h4> | ||
| + | </td> | ||
| + | <td> | ||
| + | <a href="https://2013.igem.org/Team:Tokyo_Tech/Experiment/Inducible_Plaque_Forming_Assay#1._Introduction"><img src="https://static.igem.org/mediawiki/2013/1/1e/Titech2013_M13_Plux_Fig_3-4-4-A.jpg" width="220"></a><br> | ||
| + | <h4>[Fig. 1-1-7. Distribution of plaques and analysis]<br> | ||
| + | We analyzed the distribution of the plaques by using ImageJ. | ||
| + | |||
| + | </td> | ||
| + | <td> | ||
| + | <h2><p> | ||
| + | In addition, <i>E. ninja</i> releases M13 phage, which corresponds to shuriken, when <i>E. ninja</i> receives <i>E. samurai</i> signal. The inducible phage release will open a new way in synthetic biology by achieving the programmed DNA messaging (Fig. 1-1-6). | ||
| + | <div align="right"><a href="https://2013.igem.org/Team:Tokyo_Tech/Project/M13_Shuriken#1._Abstract">(Go to Shuriken page)</a></div> | ||
| + | </p></h2> | ||
</td> | </td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
| - | <td colspan=2 | + | <td colspan=2> |
<h2><p> | <h2><p> | ||
| - | In the second-life story, E. ninja starts farming in a peaceful village. | + | In the second-life story, <i>E. ninja</i> starts farming in a peaceful village. He can increase plant growth by synthesizing several plant hormones depending on the soil environment. We constructed an improved phosphate sensor (<i>phoA</i> promoter, <a href=http://parts.igem.org/Part:BBa_K1139201>BBa_K1139201</a>). Also, we learned methods for quantitative analysis for cytokinin, one of the plant hormones, through a bioassay of cucumber seed sprouts (Fig. 1-1-8). Towards further consideration of farming with microbes, we also continued the human practice investigation through some interviews with science foundations and organizations (Fig. 1-1-8). |
| + | <div align="right"><a href="https://2013.igem.org/Team:Tokyo_Tech/Project/Farming">(Go to Farming page)</a></div> | ||
</p></h2> | </p></h2> | ||
| - | + | <br><br><br><br><br><br><br><h3><font size="6">F</font>uture Works</h3> | |
</td> | </td> | ||
| - | <td | + | <td> |
| - | <a href=""><img src="https://static.igem.org/mediawiki/2013/ | + | <a href="https://2013.igem.org/Team:Tokyo_Tech/Experiment/Quantitative_Analysis_of_Cytokinin#1._Quantitative_analysis_of_cytokinins_using_cucumber_cotyledons"><img src="https://static.igem.org/mediawiki/2013/8/8a/Titech2013_home_Fig_1-1-7_cucumber.png" width="300"></a><br> |
| - | <h4>[ | + | <h4>[Fig. 1-1-8. Our bioassay of cucumber seed sprouts]<br> |
| - | + | We cultivated the sprouts in standard cytokinin sample solutions and then measured the weights of the sprouts and the concentrations of chlorophyll. | |
| - | + | ||
</h4> | </h4> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| + | <tr> | ||
| + | <td colspan=3> | ||
| + | <h2><p> | ||
| + | We believe that our project can contribute to various fields. First, our crosstalk circumvention system gives more flexibility to design genetic circuits. By adding only a few genes, you can circumvent crosstalk of AHL. Second, our inducible phage release system can make DNA messaging more complex and more diverse. Moreover, for bioremediation, we can search for new M13 phage hosts by using our designed M13 phage. Finally, our farming project is expected to act as a pioneering trail toward new approaches in farming. Especially, our strategy to produce plant hormones in temporal patterns in <i>E. coli</i> can be applied to studying the plants' response to external plant hormones. We hope to contribute to spreading the importance and the great possibilities of synthetic biology through the public. | ||
| + | </p></h2> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr height="0"><td width="250" height="0"> </td><td width="250" height="0"> </td><td width="250" height="0"> </td></tr> | ||
</table> | </table> | ||
| - | + | ||
</div> | </div> | ||
<div class="box" id="sponsor"> | <div class="box" id="sponsor"> | ||
<div align="center"> | <div align="center"> | ||
<a href="http://www.cosmobio.co.jp/index_e.asp"><img src="https://static.igem.org/mediawiki/2013/8/85/Titech2013_home_sponsor_Cosmobio.png" height="150"></a> | <a href="http://www.cosmobio.co.jp/index_e.asp"><img src="https://static.igem.org/mediawiki/2013/8/85/Titech2013_home_sponsor_Cosmobio.png" height="150"></a> | ||
| - | <a href="http://www.idtdna.com/site"><img src="https://static.igem.org/mediawiki/2013/c/c1/Titech2013_home_sponsor_IDT.png" height="100"></a> | + | <a href="http://worldwide.promega.com/country.aspx?returnUrl=/"><img src="https://static.igem.org/mediawiki/2013/7/72/Titech2013_home_sponsor_Promega.png" height="100"></a> |
| + | <a href="http://www.idtdna.com/site"><img src="https://static.igem.org/mediawiki/2013/c/c1/Titech2013_home_sponsor_IDT.png" height="100"></a> | ||
</div><div align="center"> | </div><div align="center"> | ||
<a href="http://lne.st/en/%E6%9C%80%E5%85%88%E7%AB%AF%E7%A7%91%E5%AD%A6%E3%81%AE%E3%83%AA%E3%83%90%E3%83%8D%E3%82%B9/"><img src="https://static.igem.org/mediawiki/2013/2/2c/Titech2013_home_sponsor_LeaveaNest.png" height="60"></a> | <a href="http://lne.st/en/%E6%9C%80%E5%85%88%E7%AB%AF%E7%A7%91%E5%AD%A6%E3%81%AE%E3%83%AA%E3%83%90%E3%83%8D%E3%82%B9/"><img src="https://static.igem.org/mediawiki/2013/2/2c/Titech2013_home_sponsor_LeaveaNest.png" height="60"></a> | ||
| - | <a href="http:// | + | <a href="http://www.ikedarika.co.jp/english/"><img src="https://static.igem.org/mediawiki/2013/0/0c/Titech2013_home_sponsor_ikedarika.png" height="60"></a> |
| - | </div><br | + | </div><br><div align="center"> |
| - | Tokyo | + | <font size="6" style="white-space:nowrap;">Tokyo Institute of Technology Found project</font><br><br><br> |
| - | Aizawa Foundation<br> | + | <font size="6" style="white-space:nowrap;">Aizawa Foundation</font><br><br><br> |
| - | Mr. Isao Ono | + | <font size="6" style="white-space:nowrap;">Mr. Isao Ono Mr. Fumio Hombo</font></div> |
| - | + | ||
<br> | <br> | ||
</div><br> | </div><br> | ||
| + | <div align="center"><a href="https://2013.igem.org/Team:Tokyo_Tech#top"><img src="https://static.igem.org/mediawiki/2013/f/f0/Titeh2013_backtotop.png" width="200px"></a></div> | ||
</div> | </div> | ||
| - | |||
| - | |||
Latest revision as of 03:43, 29 October 2013
Project BackgroundIn this iGEM Competition, we intended to tell the public about the development of synthetic biology, especially about the network programming, as well as we enjoyed our activity for iGEM. Tokyo Tech 2013 assisted with an experiment workshop for high school students, participated in a poster session and collected feedback from public people as human practice (Fig. 1-1-1). We learned that an interesting story is an easier way to make general people understand the importance of genetic programming in synthetic biology. To respond further to the feedback that we received, we also address a farming issue. Thus, we designed a story which contains a farming issue and a state switching circuit in E. coli, the life of ninja: battle and farming. |
 [Fig. 1-1-1. Poster session]
|
|||||||
[Fig. 1-1-2. Ninja vs. Samurai in Tokyo Tech]
|
StoryNinja is a Japan's ancient spy-warrior. Ninja usually mimics an ordinary civilian. Once he detects samurai who is the target of assassination, he immediately gets ready for battle. He attacks samurai by throwing so-called "ninja stars". These weapons are called "shuriken" in Japanese (Fig. 1-1-2). |
|||||||
Click a step shown below to know what happens in our circuit!
|
||||||||
Project OverviewIn our programming of artificial genetic circuit, E. ninja heads casts. In response to E. civilian signal or E. samurai signal, E. ninja changes its state: "Mimic state" or "Attack state". The circuit of E. ninja contains a bi-stable switch part and a signal dependent switching part. We decided to use 3OC6HSL and 3OC12HSL as the signals. The crosstalk between these two signals is well known as a significant problem in synthetic biology. To realize an accurate switching, we designed a circuit which achieved the circumvention of crosstalk that occurs in bacterial cell-cell communication system (Fig. 1-1-4). |
[Fig. 1-1-3. The result of our wet experiment for the circumvention of the crosstalk] |
|||||||
 [Fig. 1-1-4. Our designed circuit for the circumvention of crosstalk]
|
||||||||
|
Our wet experiment results showed that the combination of lux/tet hybrid promoter and TetR protein circumvented the crosstalk by preventing the LasR protein from acting on LuxR-binding sequences (Fig. 1-1-3). Our mathematical model based on these results showed the circumvention of crosstalk in the circuit including toggle switch and crosstalk circumvention system(Fig. 1-1-5). |
||||||||
[Fig. 1-1-5. Our mathematical model for the circuit of E. ninja] |
||||||||
 [Fig. 1-1-6. Our new part for inducible phage release]
|
 [Fig. 1-1-7. Distribution of plaques and analysis] |
In addition, E. ninja releases M13 phage, which corresponds to shuriken, when E. ninja receives E. samurai signal. The inducible phage release will open a new way in synthetic biology by achieving the programmed DNA messaging (Fig. 1-1-6). |
||||||
|
In the second-life story, E. ninja starts farming in a peaceful village. He can increase plant growth by synthesizing several plant hormones depending on the soil environment. We constructed an improved phosphate sensor (phoA promoter, BBa_K1139201). Also, we learned methods for quantitative analysis for cytokinin, one of the plant hormones, through a bioassay of cucumber seed sprouts (Fig. 1-1-8). Towards further consideration of farming with microbes, we also continued the human practice investigation through some interviews with science foundations and organizations (Fig. 1-1-8). Future Works |
 [Fig. 1-1-8. Our bioassay of cucumber seed sprouts]
|
|||||||
|
We believe that our project can contribute to various fields. First, our crosstalk circumvention system gives more flexibility to design genetic circuits. By adding only a few genes, you can circumvent crosstalk of AHL. Second, our inducible phage release system can make DNA messaging more complex and more diverse. Moreover, for bioremediation, we can search for new M13 phage hosts by using our designed M13 phage. Finally, our farming project is expected to act as a pioneering trail toward new approaches in farming. Especially, our strategy to produce plant hormones in temporal patterns in E. coli can be applied to studying the plants' response to external plant hormones. We hope to contribute to spreading the importance and the great possibilities of synthetic biology through the public. |
||||||||
 "
"