Team:Peking
From 2013.igem.org
XingjiePan (Talk | contribs) |
XingjiePan (Talk | contribs) |
||
| (108 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
<style type="text/css"> | <style type="text/css"> | ||
/* hiding the top section*/ | /* hiding the top section*/ | ||
| - | body{position:absolute; top:0px; width:100%; height: | + | body{position:absolute; top:0px; width:100%; height:1300px;} |
#top-section{ | #top-section{ | ||
height:0px; | height:0px; | ||
| Line 139: | Line 139: | ||
.Navbar_Item > ul > li {float:left; list-style:none; text-align:center; background-color:transparent; position:relative; top:10px; padding:0 10px;} | .Navbar_Item > ul > li {float:left; list-style:none; text-align:center; background-color:transparent; position:relative; top:10px; padding:0 10px;} | ||
.Navbar_Item:hover{ border-bottom:1px solid #D00000; color:#D00000; background-color:#fafaf8;} | .Navbar_Item:hover{ border-bottom:1px solid #D00000; color:#D00000; background-color:#fafaf8;} | ||
| - | .Navbar_Item:hover > ul{ display:block;} | + | .Navbar_Item:hover > ul{zoom:1; display:block;} |
.Navbar_Item:hover >a {color:#D00000; background-color:#fafaf8;} | .Navbar_Item:hover >a {color:#D00000; background-color:#fafaf8;} | ||
.Navbar_Item > ul > li:hover {border-bottom:1px solid #D00000; color:#D00000; background-color:#fafaf8;} | .Navbar_Item > ul > li:hover {border-bottom:1px solid #D00000; color:#D00000; background-color:#fafaf8;} | ||
.Navbar_Item > ul > li:hover >a {color:#D00000} | .Navbar_Item > ul > li:hover >a {color:#D00000} | ||
| + | .BackgroundofSublist{position:absolute; left:-1000px; width:2000px; height:80px; background-color:#ffffff; opacity:0;} | ||
| Line 151: | Line 152: | ||
#DataPage_Sublist{position:relative; top:0px; left:-60px;} | #DataPage_Sublist{position:relative; top:0px; left:-60px;} | ||
#Safety_Sublist{position:relative; top:0px; left:-180px;} | #Safety_Sublist{position:relative; top:0px; left:-180px;} | ||
| - | #HumanPractice_Sublist{position:relative; top:0px; left:- | + | #HumanPractice_Sublist{position:relative; top:0px; left:-470px;} |
#iGEM_logo{position:absolute; top:30px; left:1090px; height:80px;} | #iGEM_logo{position:absolute; top:30px; left:1090px; height:80px;} | ||
| Line 158: | Line 159: | ||
/*Major body*/ | /*Major body*/ | ||
#MajorBody{position:absolute; top:0px; left:0px; width:1200px; height:1380px; background-color:#f9f9f7;} | #MajorBody{position:absolute; top:0px; left:0px; width:1200px; height:1380px; background-color:#f9f9f7;} | ||
| - | #ProjectTitle{position:absolute; left:0px | + | #ProjectTitle{position:absolute; left:0px; width:1200px; height:184px; top:150px; z-index:600;} |
#TitleBackground{position:absolute; left:0px; top:0px; width:1200px;} | #TitleBackground{position:absolute; left:0px; top:0px; width:1200px;} | ||
| - | #LittleScout{position: | + | #LittleScout{position:relative; left:0px; top:-3px; width:70px;} |
| - | #ProjectName{position:absolute; left:405px; top: | + | #ProjectName{position:absolute; left:405px; top:20px; border-bottom:0px; color:#FFFFFF; font-size:48px; font-family:calibri,Arial, Helvetica, sans-serif;} |
| - | #ProjectSubname{position:absolute; left:0px; width:1200px; top: | + | #ProjectSubname{position:absolute; left:0px; width:1200px; top:105px; border-bottom:0px; color:#FFFFFF; font-size:18px; font-family: calibri, Arial, Helvetica, sans-serif; text-align:center;} |
#MajorBodyBackground{position:absolute; top:334px; left:0px; width:1200px;} | #MajorBodyBackground{position:absolute; top:334px; left:0px; width:1200px;} | ||
| + | #PreLocation{font-size:14px; font-family:arial,helvetica,sans-serif; color:#ca4321;position:absolute;top:190px;left:0px;text-align:center;width:1200px;line-height:20px; font-weight:bold;} | ||
#AbstractBox{position:absolute; left:430px; top:390px; width:710px; height:540px; background-color:transparent;} | #AbstractBox{position:absolute; left:430px; top:390px; width:710px; height:540px; background-color:transparent;} | ||
#AbstractBackground{width:100%; height:100%; background-color:#FFFFFF; opacity:0.8;} | #AbstractBackground{width:100%; height:100%; background-color:#FFFFFF; opacity:0.8;} | ||
| - | #AbstractContent{position:absolute; left: | + | #AbstractContent{position:absolute; left:40px; top:25px; width:630px; font-size:18px; line-height:25px ; font-family: calibri, Arial, sans-serif; text-align:justify;} |
#AbstractContent:first-letter{font-size:30px;} | #AbstractContent:first-letter{font-size:30px;} | ||
.SmallBoxes{width:340px; height:250px;background-color:#FFFFFF; overflow:hidden;font-family: calibri, Arial, Helvetica, sans-serif; } | .SmallBoxes{width:340px; height:250px;background-color:#FFFFFF; overflow:hidden;font-family: calibri, Arial, Helvetica, sans-serif; } | ||
.SmallBoxes > img{position:absolute; left:0px; top:0px;} | .SmallBoxes > img{position:absolute; left:0px; top:0px;} | ||
| - | .SmallBoxes p{position:absolute; left:20px; top: | + | .SmallBoxes p{position:absolute; left:20px; top:75px; width:300px; font-size:14px; color:#ffffff; text-align:justify;} |
| - | .SmallBoxes a{position:absolute; left: | + | .SmallBoxes a{position:absolute; left:240px; top:220px; width:300px; font-size:12px; color:#ffffff; text-align:justify;} |
| Line 198: | Line 200: | ||
/*end of Major body*/ | /*end of Major body*/ | ||
| - | #AcknowledgementBox{ position:absolute; left:0px; top: | + | #AcknowledgementBox{ position:absolute; left:0px; top:1280px; width:1200px;} |
| Line 211: | Line 213: | ||
<ul id="navigationbar"> | <ul id="navigationbar"> | ||
<li id="PKU_navbar_Home" class="Navbar_Item"> | <li id="PKU_navbar_Home" class="Navbar_Item"> | ||
| + | |||
<a href="https://2013.igem.org/Team:Peking">Home</a> | <a href="https://2013.igem.org/Team:Peking">Home</a> | ||
<ul id="Home_Sublist" > | <ul id="Home_Sublist" > | ||
| Line 216: | Line 219: | ||
</li> | </li> | ||
<li id="PKU_navbar_Team" class="Navbar_Item"> | <li id="PKU_navbar_Team" class="Navbar_Item"> | ||
| - | <a >Team</a> | + | <a href="">Team</a> |
<ul id="Team_Sublist"> | <ul id="Team_Sublist"> | ||
| + | <div class="BackgroundofSublist"></div> | ||
<li><a href="https://2013.igem.org/Team:Peking/Team/Members">Members</a></li> | <li><a href="https://2013.igem.org/Team:Peking/Team/Members">Members</a></li> | ||
<li><a href="https://2013.igem.org/Team:Peking/Team/Notebook">Notebook</a></li> | <li><a href="https://2013.igem.org/Team:Peking/Team/Notebook">Notebook</a></li> | ||
| Line 226: | Line 230: | ||
<a href="https://2013.igem.org/Team:Peking/Project">Project</a> | <a href="https://2013.igem.org/Team:Peking/Project">Project</a> | ||
<ul id="Project_Sublist"> | <ul id="Project_Sublist"> | ||
| - | <li><a href="https://2013.igem.org/Team:Peking/Project/ | + | <div class="BackgroundofSublist"></div> |
| + | <li><a href="https://2013.igem.org/Team:Peking/Project/SensorMining">Biosensor Mining</a></li> | ||
<li><a href="https://2013.igem.org/Team:Peking/Project/BioSensors">Biosensors</a></li> | <li><a href="https://2013.igem.org/Team:Peking/Project/BioSensors">Biosensors</a></li> | ||
| - | <li><a href="https://2013.igem.org/Team:Peking/Project/Plugins"> | + | <li><a href="https://2013.igem.org/Team:Peking/Project/Plugins">Adaptors</a></li> |
<li><a href="https://2013.igem.org/Team:Peking/Project/BandpassFilter">Band-pass Filter</a></li> | <li><a href="https://2013.igem.org/Team:Peking/Project/BandpassFilter">Band-pass Filter</a></li> | ||
| + | <li><a href="https://2013.igem.org/Team:Peking/Project/Devices">Devices</a></li> | ||
</ul> | </ul> | ||
</li> | </li> | ||
<li id="PKU_navbar_Model" class="Navbar_Item"> | <li id="PKU_navbar_Model" class="Navbar_Item"> | ||
| - | <a href=" | + | <a href="">Model</a> |
<ul id="Model_Sublist"> | <ul id="Model_Sublist"> | ||
| + | <div class="BackgroundofSublist"></div> | ||
| + | <li><a href="https://2013.igem.org/Team:Peking/Model">Band-pass Filter</a></li> | ||
| + | <li><a href="https://2013.igem.org/Team:Peking/ModelforFinetuning">Biosensor Fine-tuning</a></li> | ||
</ul> | </ul> | ||
</li> | </li> | ||
<li id="PKU_navbar_HumanPractice" class="Navbar_Item" style="width:90px"> | <li id="PKU_navbar_HumanPractice" class="Navbar_Item" style="width:90px"> | ||
| - | <a href=" | + | <a href="">Data page</a> |
| - | <ul id="DataPage_Sublist"> | + | <ul id="DataPage_Sublist"> |
| + | <div class="BackgroundofSublist"></div> | ||
<li><a href="https://2013.igem.org/Team:Peking/DataPage/Parts">Parts</a></li> | <li><a href="https://2013.igem.org/Team:Peking/DataPage/Parts">Parts</a></li> | ||
<li><a href="https://2013.igem.org/Team:Peking/DataPage/JudgingCriteria">Judging Criteria</a></li> | <li><a href="https://2013.igem.org/Team:Peking/DataPage/JudgingCriteria">Judging Criteria</a></li> | ||
| Line 252: | Line 262: | ||
<li id="PKU_navbar_HumanPractice" class="Navbar_Item" style="width:120px"> | <li id="PKU_navbar_HumanPractice" class="Navbar_Item" style="width:120px"> | ||
<a href="https://2013.igem.org/Team:Peking/HumanPractice">Human Practice</a> | <a href="https://2013.igem.org/Team:Peking/HumanPractice">Human Practice</a> | ||
| - | <ul id="HumanPractice_Sublist"> | + | <ul id="HumanPractice_Sublist"> |
| - | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/Questionnaire">Questionnaire</a></li> | + | <div class="BackgroundofSublist"></div> |
| - | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/FactoryVisit"> | + | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/Questionnaire">Questionnaire Survey</a></li> |
| - | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/ | + | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/FactoryVisit">Visit and Interview</a></li> |
| - | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/ | + | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/ModeliGEM">Practical Analysis</a></li> |
| + | <li><a href="https://2013.igem.org/Team:Peking/HumanPractice/iGEMWorkshop">Team Communication</a></li> | ||
| + | |||
</ul> | </ul> | ||
</li> | </li> | ||
</ul> | </ul> | ||
| - | <a href="https://igem.org/ | + | <a href="https://2013.igem.org/Main_Page"><img id="iGEM_logo" src="https://static.igem.org/mediawiki/igem.org/4/48/Peking_igemlogo.jpg"/></a> |
</div> | </div> | ||
<!--end navigationbar--> | <!--end navigationbar--> | ||
| Line 269: | Line 281: | ||
<div id="ProjectTitle"> | <div id="ProjectTitle"> | ||
<img id="TitleBackground" src="https://static.igem.org/mediawiki/igem.org/c/c5/Peking2013_home_title.jpg" /> | <img id="TitleBackground" src="https://static.igem.org/mediawiki/igem.org/c/c5/Peking2013_home_title.jpg" /> | ||
| - | <img id="LittleScout" src="https://static.igem.org/mediawiki/igem.org/8/8a/Peking2013_home_Telescope.png" /> | + | |
| - | + | <h1 id="ProjectName">Aromatics Scouts<img id="LittleScout" src="https://static.igem.org/mediawiki/igem.org/8/8a/Peking2013_home_Telescope.png" /></h1> | |
| - | <h1 id="ProjectSubname">A Comprehensive Biosensor Toolkit | + | <h1 id="ProjectSubname">A Comprehensive Biosensor Toolkit for Profiling Aromatic Compounds in the Environment</h1> |
| + | <p id="PreLocation"> | ||
| + | <B>★Presentation:</B> Room 34-101, Saturday 12:30 PM, Session 2 | ||
| + |     <B>★Poster:</B> Sunday, #13 (Stata) | ||
| + | </p> | ||
| + | |||
| + | |||
| + | |||
</div> | </div> | ||
<img id="MajorBodyBackground" src="https://static.igem.org/mediawiki/igem.org/9/97/Peking2013_Home_background.jpg" /> | <img id="MajorBodyBackground" src="https://static.igem.org/mediawiki/igem.org/9/97/Peking2013_Home_background.jpg" /> | ||
| Line 279: | Line 298: | ||
<p id="AbstractContent">Monitoring aromatic compounds in the environment remains a substantial challenge today. Noting the power of biosensors for quick and convenient testing, Peking iGEM has developed a functionally comprehensive biosensor toolkit to profile aromatics in the environment. | <p id="AbstractContent">Monitoring aromatic compounds in the environment remains a substantial challenge today. Noting the power of biosensors for quick and convenient testing, Peking iGEM has developed a functionally comprehensive biosensor toolkit to profile aromatics in the environment. | ||
<br/><br/> | <br/><br/> | ||
| - | Transcriptional regulators that each | + | Transcriptional regulators that each detect a specific class of aromatics were first bioinformatically determined; and then utilized to build a comprehensive set of biosensor circuits. Characterization on the detection profiles of individual biosensors and the orthogonality/crosstalk between them proved that these biosensors are very capable at profiling aromatics present in water. |
<br/><br/> | <br/><br/> | ||
| - | Moreover, for the ease of practical applications, two types of genetic devices were also developed as plug-ins for biosensors: "Adaptors", a set of conceptually novel devices to convert undetectable | + | Moreover, for the ease of practical applications, two types of genetic devices were also developed as plug-ins for biosensors: "Adaptors", a set of conceptually novel devices to convert undetectable compounds into detectable compounds, and "Band-pass Filter", a "concentration filter" that allows the detection of analyte concentration within a specific range. |
<br/><br/> | <br/><br/> | ||
We expect that these novel biosensors, together with the plug-in devices, will serve as intriguing synthetic biological tools for diverse practical applications. | We expect that these novel biosensors, together with the plug-in devices, will serve as intriguing synthetic biological tools for diverse practical applications. | ||
| Line 290: | Line 309: | ||
<div id="Appendix1Cover" > | <div id="Appendix1Cover" > | ||
<img id="AppendixImg1" src="https://static.igem.org/mediawiki/igem.org/e/ec/Peking2013_homepage_appendix1.jpg" /> | <img id="AppendixImg1" src="https://static.igem.org/mediawiki/igem.org/e/ec/Peking2013_homepage_appendix1.jpg" /> | ||
| - | <h1 class="AppendixHeadLine"> | + | <h1 class="AppendixHeadLine">Biosensor Mining</h1> |
<img id="Appendix1Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/1/13/Peking2013_home_appendix1icon.png" /> | <img id="Appendix1Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/1/13/Peking2013_home_appendix1icon.png" /> | ||
| - | <p> | + | <p>The core component of our biosensor toolkit is the transcriptional regulators that sense aromatics. For the comprehensiveness of aromatics-sensing, a data-mining process was conducted to mine transcriptional regulators for each typical class of aromatics from the database Uniprot. </p> |
| - | <a href="">LEARN MORE</a> | + | <a href="https://2013.igem.org/Team:Peking/Project/SensorMining">LEARN MORE</a> |
</div> | </div> | ||
</div> | </div> | ||
| Line 302: | Line 321: | ||
<h1 class="AppendixHeadLine">Biosensors</h1> | <h1 class="AppendixHeadLine">Biosensors</h1> | ||
<img id="Appendix2Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/b/b2/Peking2013_home_appendix2icon.png" /> | <img id="Appendix2Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/b/b2/Peking2013_home_appendix2icon.png" /> | ||
| + | <p>A comprehensive set of biosensor circuits have been implemented using the aromatics-sensing transcriptional regulators. Each biosensor has a specific aromatics-sensing profile. Furthermore, the orthogonality of their sensing profiles was carefully assessed for practical applications.</p> | ||
| + | <a href="https://2013.igem.org/Team:Peking/Project/BioSensors">LEARN MORE</a> | ||
</div> | </div> | ||
</div> | </div> | ||
<div id="AppendixBox3" class="SmallBoxes"> | <div id="AppendixBox3" class="SmallBoxes"> | ||
| - | <h1 id="Appendix3Title"><i> | + | <h1 id="Appendix3Title"><i>Converting the undetectable into<br/>the detectable</i></h1> |
<div id="Appendix3Cover" > | <div id="Appendix3Cover" > | ||
<img id="AppendixImg3" src="https://static.igem.org/mediawiki/igem.org/1/1b/Peking2013_home_appendix3.jpg" /> | <img id="AppendixImg3" src="https://static.igem.org/mediawiki/igem.org/1/1b/Peking2013_home_appendix3.jpg" /> | ||
<h1 class="AppendixHeadLine">Adaptors</h1> | <h1 class="AppendixHeadLine">Adaptors</h1> | ||
<img id="Appendix3Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/c/cd/Peking2013_home_appendix3icon.png" /> | <img id="Appendix3Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/c/cd/Peking2013_home_appendix3icon.png" /> | ||
| + | <p> To expand the detection profiles of some biosensors, aromatics-metabolizing enzymes were taken from natural metabolic pathways, working as Adaptors to convert undetectable chemicals into detectable aromatics when coupled with biosensor circuits.</p> | ||
| + | <a href="https://2013.igem.org/Team:Peking/Project/Plugins">LEARN MORE</a> | ||
</div> | </div> | ||
</div> | </div> | ||
<div id="AppendixBox4" class="SmallBoxes"> | <div id="AppendixBox4" class="SmallBoxes"> | ||
| - | <h1 id="Appendix4Title"><i>Rapidly | + | <h1 id="Appendix4Title"><i>Rapidly determining <br/> the analyte concentration</i></h1> |
<div id="Appendix4Cover" > | <div id="Appendix4Cover" > | ||
<img id="AppendixImg4" src="https://static.igem.org/mediawiki/igem.org/e/e6/Peking2013_home_appendix4.jpg" /> | <img id="AppendixImg4" src="https://static.igem.org/mediawiki/igem.org/e/e6/Peking2013_home_appendix4.jpg" /> | ||
<h1 class="AppendixHeadLine">Band-pass Filter</h1> | <h1 class="AppendixHeadLine">Band-pass Filter</h1> | ||
<img id="Appendix4Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/4/42/Peking2013_home_appendix4icon.png" /> | <img id="Appendix4Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/4/42/Peking2013_home_appendix4icon.png" /> | ||
| + | <p>For the ease of practical analysis, a genetic device called "Band-pass Filter" has been constructed to allow the detection of analyte concentration within a specific range. Biosensors equipped with the Band-pass Filter can robustly quantify the aromatics in environmental samples. </p> | ||
| + | <a href="https://2013.igem.org/Team:Peking/Project/BandpassFilter">LEARN MORE</a> | ||
</div> | </div> | ||
</div> | </div> | ||
<div id="AppendixBox5" class="SmallBoxes"> | <div id="AppendixBox5" class="SmallBoxes"> | ||
| - | <h1 id="Appendix5Title"><i> | + | <h1 id="Appendix5Title"><i>Communications, Questionnaire survey, <br/>Factory Visit</i> and interview</h1> |
<div id="Appendix5Cover" > | <div id="Appendix5Cover" > | ||
<img id="AppendixImg5" src="https://static.igem.org/mediawiki/igem.org/d/da/Peking2013_home_appendix5.jpg" /> | <img id="AppendixImg5" src="https://static.igem.org/mediawiki/igem.org/d/da/Peking2013_home_appendix5.jpg" /> | ||
<h1 class="AppendixHeadLine">Human Practice</h1> | <h1 class="AppendixHeadLine">Human Practice</h1> | ||
<img id="Appendix5Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/8/85/Peking2013_home_appendix5icon.png" /> | <img id="Appendix5Icon" class="AppendixIcon" src="https://static.igem.org/mediawiki/igem.org/8/85/Peking2013_home_appendix5icon.png" /> | ||
| + | <p>To obtain the information about public awareness and the situation of aromatics pollution, a survey including factory visit and questionnaires has been conducted. We also guided an iGEM HS team and held "Model iGEM" as a competition without competitiveness.</p> | ||
| + | <a href="https://2013.igem.org/Team:Peking/HumanPractice">LEARN MORE</a> | ||
</div> | </div> | ||
</div> | </div> | ||
Latest revision as of 18:10, 28 October 2013

Aromatics Scouts
A Comprehensive Biosensor Toolkit for Profiling Aromatic Compounds in the Environment
★Presentation: Room 34-101, Saturday 12:30 PM, Session 2 ★Poster: Sunday, #13 (Stata)

Monitoring aromatic compounds in the environment remains a substantial challenge today. Noting the power of biosensors for quick and convenient testing, Peking iGEM has developed a functionally comprehensive biosensor toolkit to profile aromatics in the environment.
Transcriptional regulators that each detect a specific class of aromatics were first bioinformatically determined; and then utilized to build a comprehensive set of biosensor circuits. Characterization on the detection profiles of individual biosensors and the orthogonality/crosstalk between them proved that these biosensors are very capable at profiling aromatics present in water.
Moreover, for the ease of practical applications, two types of genetic devices were also developed as plug-ins for biosensors: "Adaptors", a set of conceptually novel devices to convert undetectable compounds into detectable compounds, and "Band-pass Filter", a "concentration filter" that allows the detection of analyte concentration within a specific range.
We expect that these novel biosensors, together with the plug-in devices, will serve as intriguing synthetic biological tools for diverse practical applications.
Mining aromatics-sensing Biobricks
from the genomic database

Biosensor Mining
The core component of our biosensor toolkit is the transcriptional regulators that sense aromatics. For the comprehensiveness of aromatics-sensing, a data-mining process was conducted to mine transcriptional regulators for each typical class of aromatics from the database Uniprot.
LEARN MOREHigh-performance, profile-
specific biosensors

Biosensors
A comprehensive set of biosensor circuits have been implemented using the aromatics-sensing transcriptional regulators. Each biosensor has a specific aromatics-sensing profile. Furthermore, the orthogonality of their sensing profiles was carefully assessed for practical applications.
LEARN MOREConverting the undetectable into
the detectable

Adaptors
To expand the detection profiles of some biosensors, aromatics-metabolizing enzymes were taken from natural metabolic pathways, working as Adaptors to convert undetectable chemicals into detectable aromatics when coupled with biosensor circuits.
LEARN MORERapidly determining
the analyte concentration
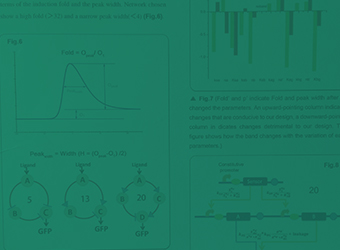
Band-pass Filter
For the ease of practical analysis, a genetic device called "Band-pass Filter" has been constructed to allow the detection of analyte concentration within a specific range. Biosensors equipped with the Band-pass Filter can robustly quantify the aromatics in environmental samples.
LEARN MORECommunications, Questionnaire survey,
Factory Visit and interview

Human Practice
To obtain the information about public awareness and the situation of aromatics pollution, a survey including factory visit and questionnaires has been conducted. We also guided an iGEM HS team and held "Model iGEM" as a competition without competitiveness.
LEARN MORE
 "
"

