Team:Goettingen/Team/Reporter
From 2013.igem.org
FMCommmichau (Talk | contribs) |
(→Discussion) |
||
| (4 intermediate revisions not shown) | |||
| Line 47: | Line 47: | ||
===Reporter Team=== | ===Reporter Team=== | ||
==Introduction== | ==Introduction== | ||
| - | In order to facilitate the development of new antibiotics, we want to build an “Antibiotic Detector” that can identify substances | + | In order to facilitate the development of new antibiotics, we want to build an “Antibiotic Detector” that can identify substances, affecting homeostasis of cyclic-di-AMP (c-di-AMP). c-di-AMP is a signal nucleotide that is essential for viability of many Gram-positive bacteria. It is not found in humans as well as in Gram-negative bacteria, including <i>E. coli</i>. For more information see [[Team:Goettingen/Project/OurProject|Our Project overview]]. |
| - | For constructing a c-di-AMP detector, we need | + | For constructing a c-di-AMP detector, we need elements that are capable of sensing c-di-AMP! Moreover, in order to visualize the output of the sensors, we need a reporter. |
==DarR reporter system ([http://parts.igem.org/Part:BBa_K1045017 part BBa_K1045017]):== | ==DarR reporter system ([http://parts.igem.org/Part:BBa_K1045017 part BBa_K1045017]):== | ||
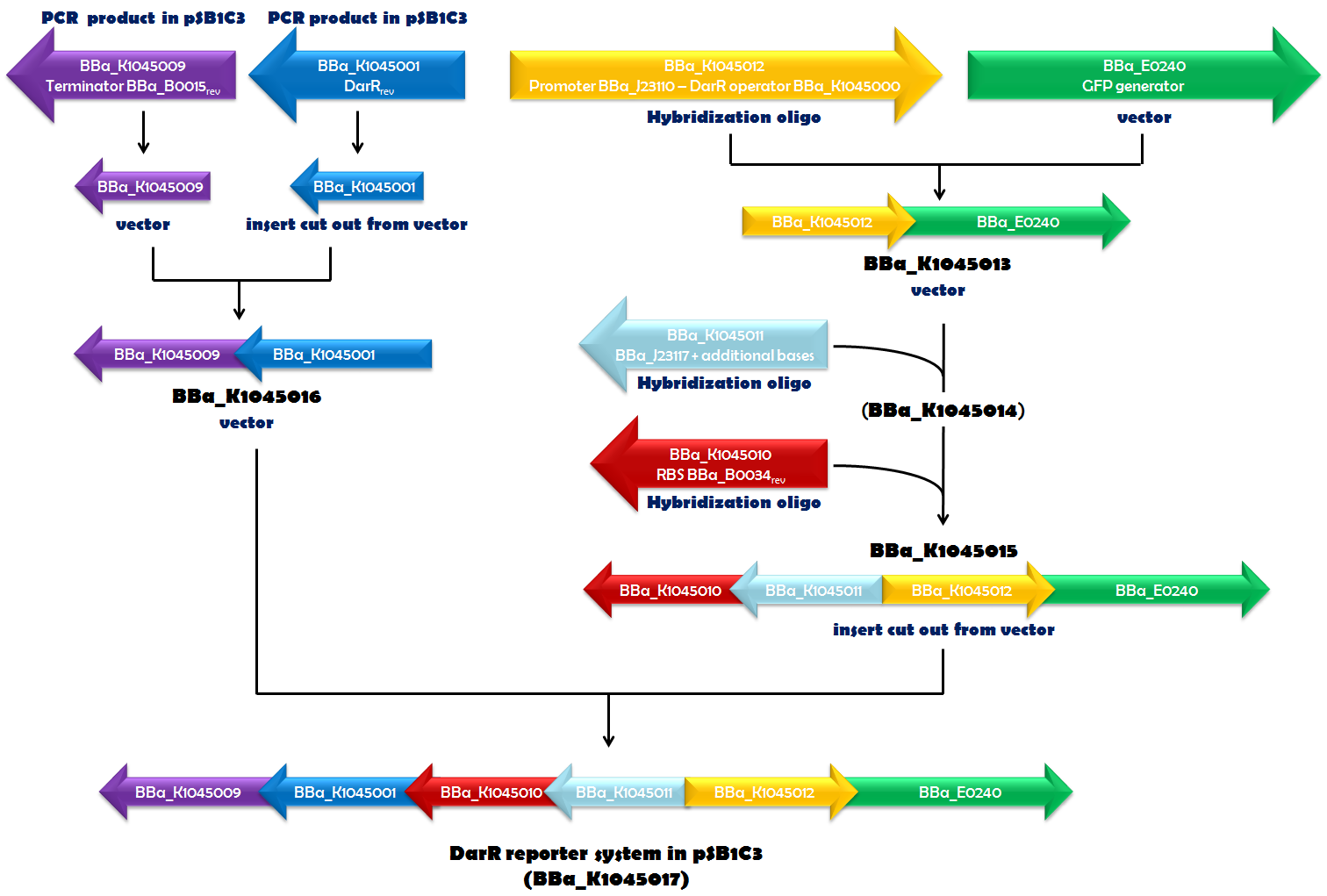
| - | Recently, in ''Mycobacterium smegmatis'', | + | Recently, in ''Mycobacterium smegmatis'', the transcriptional repressor DarR was identified and shown to respond to c-di-AMP. It can bind to a specific DNA sequence, called the DarR operator. DNA-binding activity of DarR is strongly enhanced by c-di-AMP. Since DarR is a c-di-AMP sensor, we intended to use it for our reporter system. We cloned the DNA sequence, encoding DarR into pSB1C3 ([http://parts.igem.org/Part:BBa_K1045001 BBa_K1045001]). In addition to this, we constructed the DarR operator as a Biobrick ([http://parts.igem.org/Part:BBa_K1045000 BBa_K1045000]). We placed the DarR operator between a strong promoter ([http://parts.igem.org/Part:BBa_J23110 BBa_J23110]) and the GFP generator [http://parts.igem.org/Part:BBa_E0240 BBa_E0240]. In the very same plasmid but oriented in the opposite direction, we assembled a <i>darR</i> expression unit. Synthesis of DarR is driven by a weak promoter based on [http://parts.igem.org/Part:BBa_J23117 BBa_J23117] and terminated by [http://parts.igem.org/Part:BBa_K1045009 BBa_K1045009]. This terminator is based on [http://parts.igem.org/Part:BBa_B0015 BBa_B0015], which is part of [http://parts.igem.org/Part:BBa_E0240 BBa_E0240], as well, and should ensure that transcription of <i>darR</i> and <i>gfp</i> does not influence each other. The ribosome-binding site (RBS) [http://parts.igem.org/Part:BBa_K1045010 BBa_K1045010] (derived from [http://parts.igem.org/Part:BBa_B0034 BBa_B0034]) was used as an RBS for translation of the <i>darR</i> mRNA. Assembly of all those parts (Fig. 1.1) required a complex cloning process which finally resulted in part [http://parts.igem.org/Part:BBa_K1045017 BBa_K1045017], the DarR reporter system (Fig. 1.2). ''E. coli'' was then transformed with this construct to obtain an ''in vivo'' screening system that interfere with c-di-AMP-binding to a c-di-AMP target. |
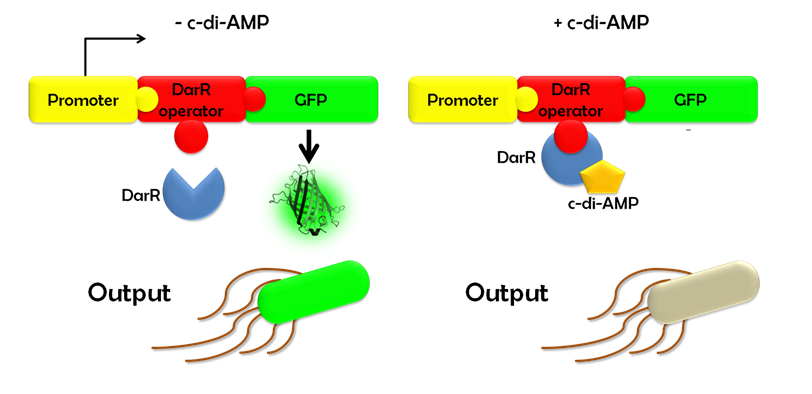
Ideally, this c-di-AMP-sensing system works in the following way (Fig. 1.3): Without c-di-AMP, DarR is not bound to the DarR operator, so GFP is expressed resulting in green fluorescing cells. With c-di-AMP, DarR can bind to its binding sequence, repressing <i>gfp</i> transcription, leading to non-fluorescent cells. In the same way, the system might react to compounds similar to c-di-AMP. It can therefore be used to screen for compounds similar to c-di-AMP. It can be used in a high-throughput scale, as well. This way, the DarR reporter system will make it possible to find inhibitors and competitors which might be used as antibiotics. | Ideally, this c-di-AMP-sensing system works in the following way (Fig. 1.3): Without c-di-AMP, DarR is not bound to the DarR operator, so GFP is expressed resulting in green fluorescing cells. With c-di-AMP, DarR can bind to its binding sequence, repressing <i>gfp</i> transcription, leading to non-fluorescent cells. In the same way, the system might react to compounds similar to c-di-AMP. It can therefore be used to screen for compounds similar to c-di-AMP. It can be used in a high-throughput scale, as well. This way, the DarR reporter system will make it possible to find inhibitors and competitors which might be used as antibiotics. | ||
| Line 71: | Line 71: | ||
<html> | <html> | ||
| - | <p><b>Reference | + | <p><b>Reference</b></p> |
| - | + | <p>Zhang <i>et al.</i> (2013) DarR, a TetR-like transcriptional factor, Is a cyclic di-AMP-responsive repressor in <i>Mycobacterium smegmatis. J Biol Chem</i> 288:3085–3096.</p> | |
| + | |||
<br /> | <br /> | ||
| Line 94: | Line 95: | ||
<p><b>References</b></p> | <p><b>References</b></p> | ||
| - | <p>Watson, P. Y. & Fedor, M. J. (2012) The ''ydaO'' motif is an ATP-sensing riboswitch in ''Bacillus subtilis'', '' | + | <p>Watson, P. Y. & Fedor, M. J. (2012) The ''ydaO'' motif is an ATP-sensing riboswitch in ''Bacillus subtilis'', ''Nat Chem Biol'' 8: 963-96.</p> |
| - | <p>Nelson ''et al.'' (2013) Riboswitches in eubacteria sense the second messenger c-di-AMP. '' | + | <p>Nelson ''et al.'' (2013) Riboswitches in eubacteria sense the second messenger c-di-AMP. ''Nat Chem Biol'' 8: 963-96. doi: 10.1038/nchembio.1363.</p> |
| Line 123: | Line 124: | ||
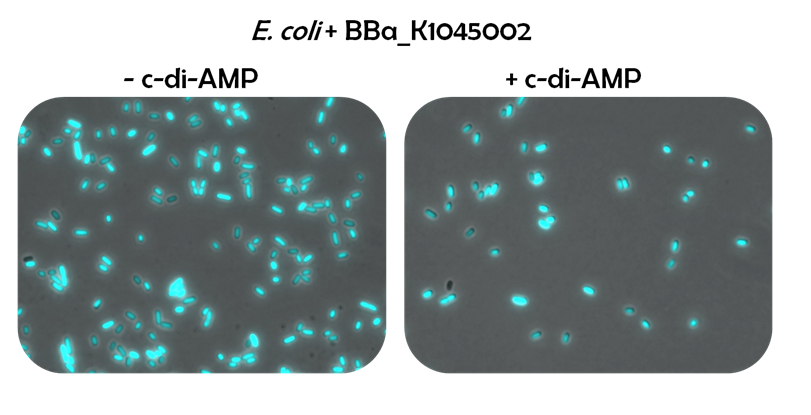
Regarding the alternative reporter system involving the c-di-AMP-responsive ''ydaO'' riboswitch, optimization is needed, as well. The results obtained from fluorescence microscopy showed that the riboswitch reporter construct is expressed in ''E. coli'' though the native Bacillus promoter and Bacillus RBS were employed. Hence, ''E. coli'' seems to be able to use these elements. The strong CFP fluorescence of the ''E. coli'' cells even suggests a strong promoter activity. The high transcript levels that might be caused by this strong promoter could account for the observed ineffectiveness of exogenously applied c-di-AMP: There might be more riboswitch than c-di-AMP to bind to. Consequently, one could try to increase the c-di-AMP amounts. Alternatively, one could fuse the riboswitch biobrick [http://parts.igem.org/Part:BBa_K1045005 BBa_K1045005] to a weaker promoter to reduce the mRNA levels. Another possibility might be to switch to low-copy plasmids for expression of the reporter system. If despite reduced transcript levels, c-di-AMP has no effect on the riboswitch reporter system, ''E. coli'' might be unable to take up c-di-AMP. Yet, this would not put an end to the construction of a screening system for antibiotics targeting c-di-AMP. It has been reported, that B. subtilis is able to take up c-di-AMP (Oppenheimer-Shaaman ''et al.'', 2011). Thus, one could put more effort in unraveling the ways by which ''B. subtilis'' and other bacteria take up c-di-AMP. This knowledge might allow the engineering of ''E. coli'' regarding c-di-AMP uptake. In parallel to this time-consuming approach, one could try to synthesize c-di-AMP ''in vivo'' by expressing diadenylate cyclases in ''E. coli'' cells containing an optimized c-di-AMP reporter system. Using DACs of different activity, the output of the reporter system could be characterized. Finally, having a well characterized c-di-AMP reporter system combined with a DAC, it is possible to screen ''in vivo'' for antibiotics interfering not only with the function of the essential signaling nucleotide c-di-AMP, but also for its essential biosynthesis enzyme, the DAC. | Regarding the alternative reporter system involving the c-di-AMP-responsive ''ydaO'' riboswitch, optimization is needed, as well. The results obtained from fluorescence microscopy showed that the riboswitch reporter construct is expressed in ''E. coli'' though the native Bacillus promoter and Bacillus RBS were employed. Hence, ''E. coli'' seems to be able to use these elements. The strong CFP fluorescence of the ''E. coli'' cells even suggests a strong promoter activity. The high transcript levels that might be caused by this strong promoter could account for the observed ineffectiveness of exogenously applied c-di-AMP: There might be more riboswitch than c-di-AMP to bind to. Consequently, one could try to increase the c-di-AMP amounts. Alternatively, one could fuse the riboswitch biobrick [http://parts.igem.org/Part:BBa_K1045005 BBa_K1045005] to a weaker promoter to reduce the mRNA levels. Another possibility might be to switch to low-copy plasmids for expression of the reporter system. If despite reduced transcript levels, c-di-AMP has no effect on the riboswitch reporter system, ''E. coli'' might be unable to take up c-di-AMP. Yet, this would not put an end to the construction of a screening system for antibiotics targeting c-di-AMP. It has been reported, that B. subtilis is able to take up c-di-AMP (Oppenheimer-Shaaman ''et al.'', 2011). Thus, one could put more effort in unraveling the ways by which ''B. subtilis'' and other bacteria take up c-di-AMP. This knowledge might allow the engineering of ''E. coli'' regarding c-di-AMP uptake. In parallel to this time-consuming approach, one could try to synthesize c-di-AMP ''in vivo'' by expressing diadenylate cyclases in ''E. coli'' cells containing an optimized c-di-AMP reporter system. Using DACs of different activity, the output of the reporter system could be characterized. Finally, having a well characterized c-di-AMP reporter system combined with a DAC, it is possible to screen ''in vivo'' for antibiotics interfering not only with the function of the essential signaling nucleotide c-di-AMP, but also for its essential biosynthesis enzyme, the DAC. | ||
| + | |||
| + | '''References:''' | ||
| + | |||
| + | Yaara Oppenheimer-Shaanan, Ezequiel Wexselblatt, Jehoshua Katzhendler, Eylon Yavin & Sigal Ben-Yehuda (2011) “c-di-AMP reports DNA integrity during sporulation in ''Bacillus subtilis''”, EMBO reports Vol. 12, No. 6, pp. 594-601 | ||
=== === | === === | ||
Latest revision as of 09:30, 28 October 2013
 "
"